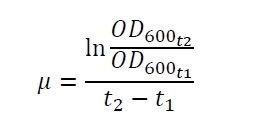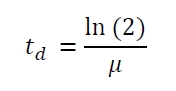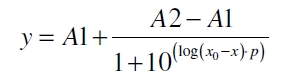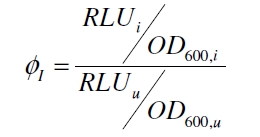Team:Bielefeld-Germany/Results/Characterization/K389015
From 2010.igem.org
Contents |
Characterization of <partinfo>K389015</partinfo>
On this page the experiments and results that lead to the <partinfo>K389015</partinfo> characterization data presented on our characterization page are shown in detail.
Growth functions and Luciferase expression for <partinfo>K389015</partinfo>
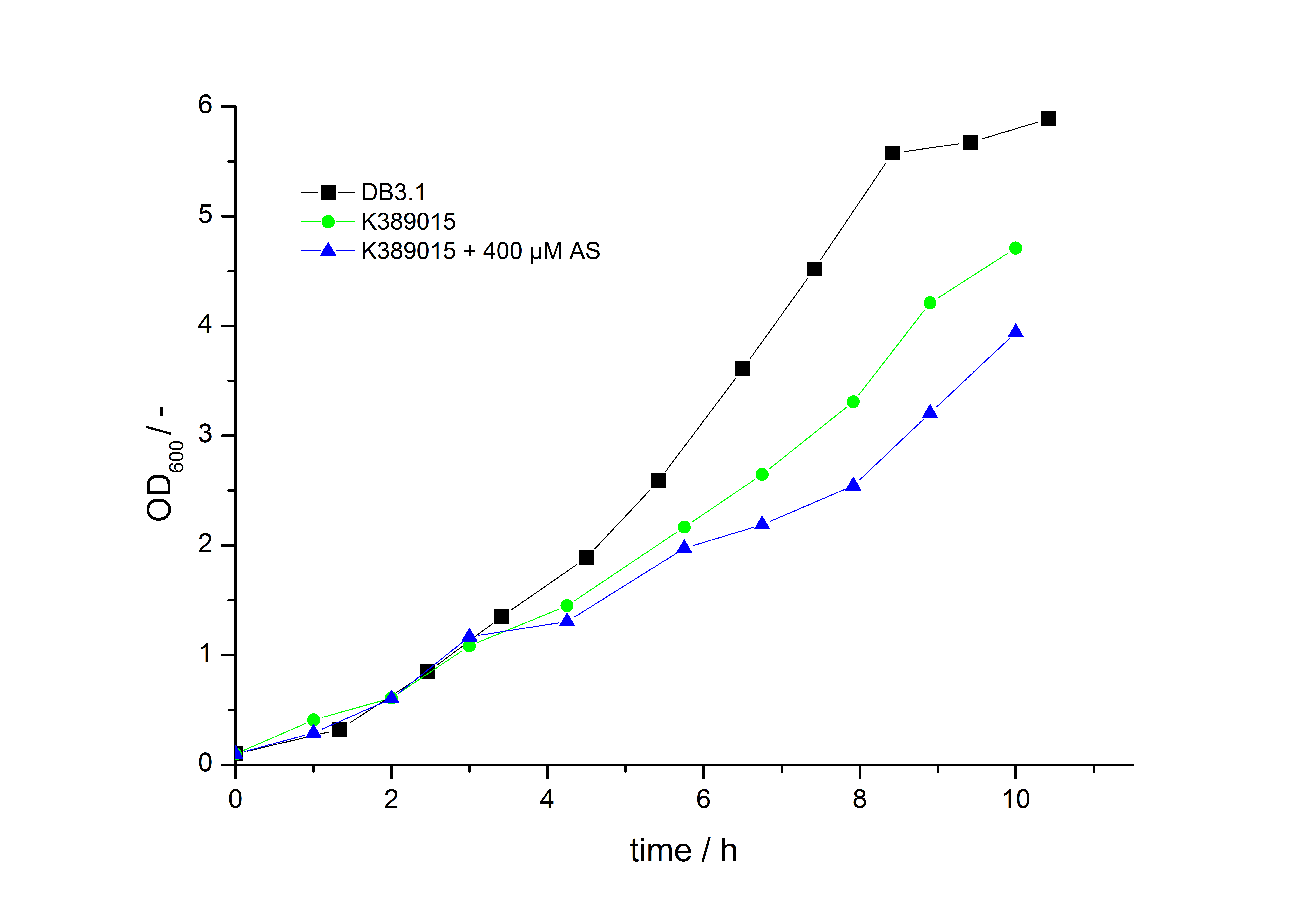
To characterize this part we performed several cultivations with different concentrations of [http://www.chemblink.com/products/2478-38-8.htm acetosyringone] as inducer and measured the luminescence emitted by the luciferasereaction with Luciferin (Protocol). We used Escherichia coli DB3.1 carrying the pSB1C3::K389015 plasmid. Even without inducer the bacteria carrying the plasmid showed decelerated growth. In addition acetosyringone affected the growth rates (we used a stocksolution of 20 mM acetosyringone solved in 10 % (v/v) DMSO). Growth curves, averaged specific growth rates and doubling times are shown below. It can be observed, that E. coli carrying the pSB1C3::K389015 plasmid growths nearly linear.
The specific growth rates µ and doubling times td are calculated with the OD600 and following formulas:
| E. coli DB3.1 | µ / h-1 | td / h |
|---|---|---|
| without plasmid | 0.35 | 1.98 |
| carrying K389015 | 0.31 | 2.24 |
| carrying K389015 with 400 µM acetosyringone | 0.26 | 2.67 |
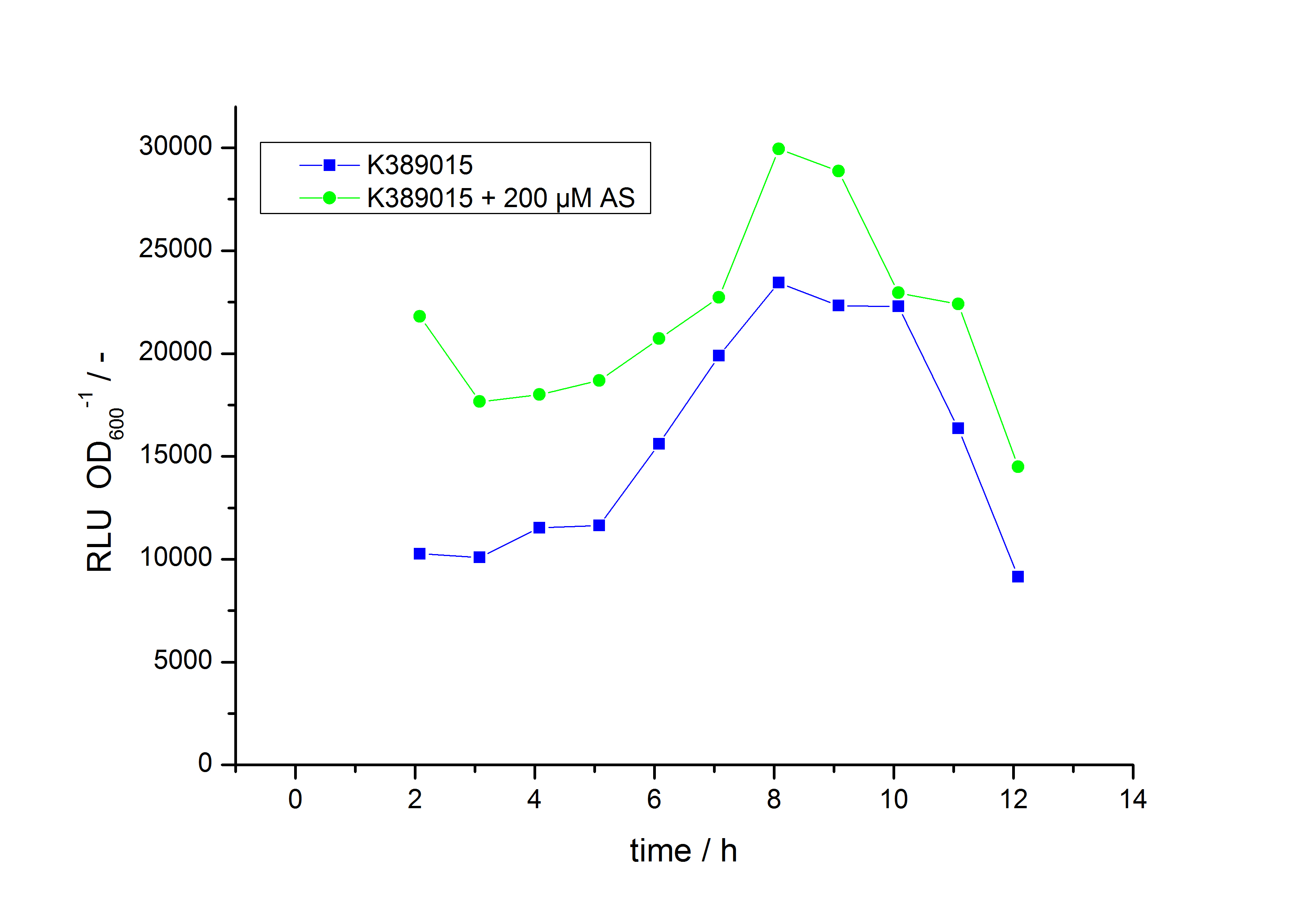
Exemplary induction curves with the luminescence normalized to OD600 are shown in Fig. 2. We observed a basal transcription, but the induction with acetosyringone is undoubtedly. The detailed data analysis and transfer function is described below.
Transfer function
The data for the transfer function was measured and analyzed as described below. Nelson et al. suggests using a dose response function and fitting it with a logistical equation for the data analysis of receptor systems ([http://www.nature.com/nature/journal/v416/n6877/abs/nature726.html Nelson et al., 2002]). The data was fitted with a function of the form
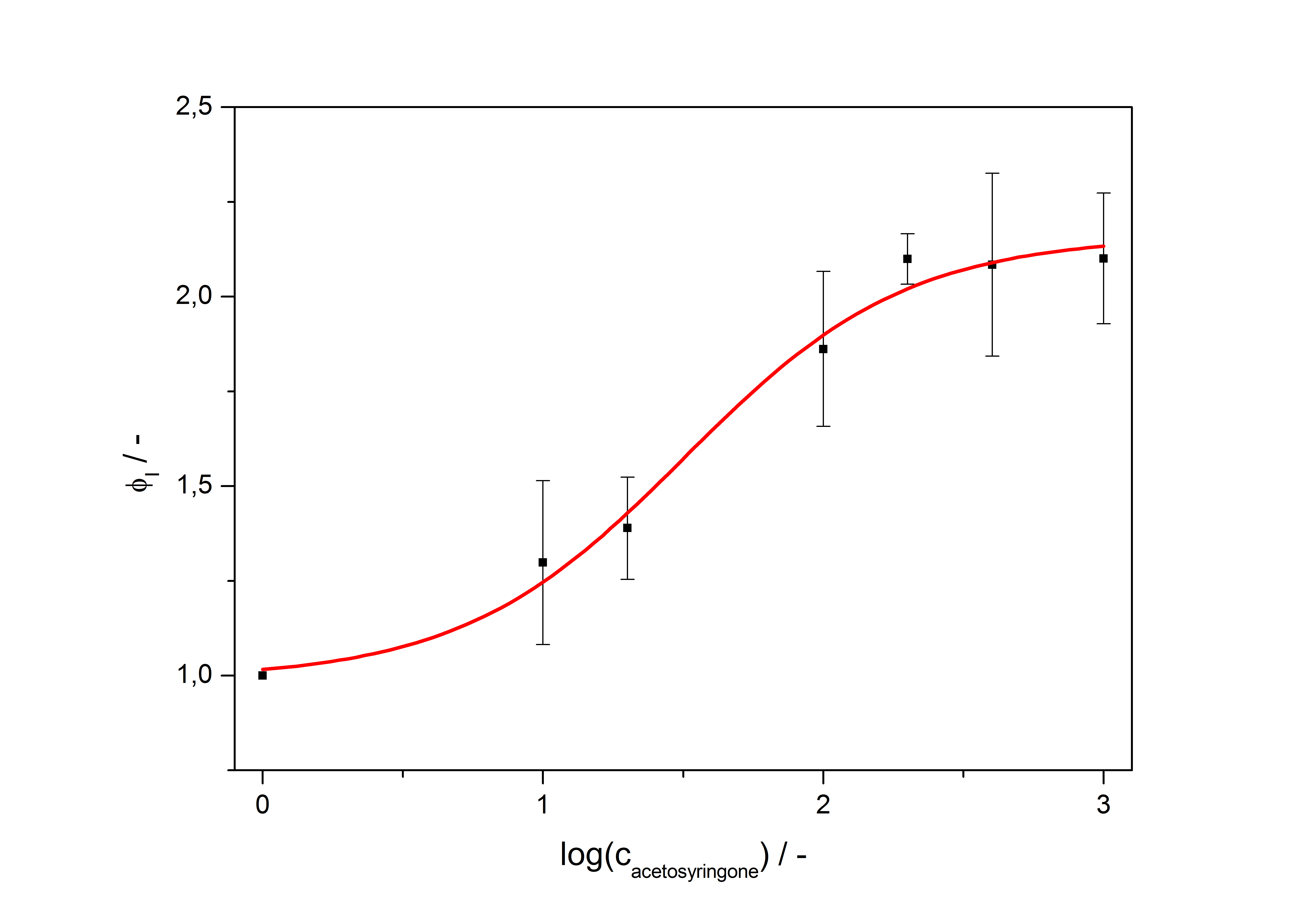
with the Hill coefficient p, the bottom asymptote A1, the top asymptote A2 and the switch point log(x0). Figure 3 shows the measured ratio ɸI between induced (i) and uninduced (u) relative luminescence units (RLU) per OD600 plotted against the logarithm of the concentration of the inductor [http://www.chemblink.com/products/2478-38-8.htm acetosyringone] in µM. The fit has an R2 = 0.98.
The important data from the transfer function is summarized in table 1:
| Parameter | Value |
|---|---|
| Hill coefficient | 1.092 |
| [http://partsregistry.org/Switch_Point Switch point] | 31.6 µM |
| Top asymptote | 2.16 |
The fully induced VirA/G signaling system with luciferase readout has a 2.2 fold increased expression compared to the uninduced system. The Hill coefficient is > 1, so a positive cooperation can be observed ([http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WMD-4V42JG5-1&_user=10&_rdoc=1&_fmt=&_orig=search&_sort=d&_docanchor=&view=c&_acct=C000050221&_version=1&_urlVersion=0&_userid=10&md5=b6431553217aca1129c5b441f4b78425 D Chu et al., 2009]). The [http://partsregistry.org/Switch_Point switch point] of the system is at about 32 µM, so this is the concentration at which the device output is 50% of the maximum output.
Response time
The system needs at least one hour to show a measurable reaction to an induction with acetosyringone. In the following illustration the reaction of the system to induction with 200 µM acetosyringone in the exponential growth phase is shown. For a good separation of the induced system from the uninduced system at least two hours are needed.
Data Analysis
Because the luciferase accumulation is very different in different cultivations, the uninduced negative control was used as internal standard. To show the behaviour of the VirA/G signaling system when induced, the ratio ɸI between induced (i) and uninduced (u) relative luminescence units (RLU) per OD600 is calculated:
As seen above, at least one hour is needed to separate the induced luminescence signal from the uninduced, so ɸI > 1. Within a cultivation ɸI is rising during the first hours and is decreasing after it reached a maximum at OD600 ~ 1. This is shown in figure 3:
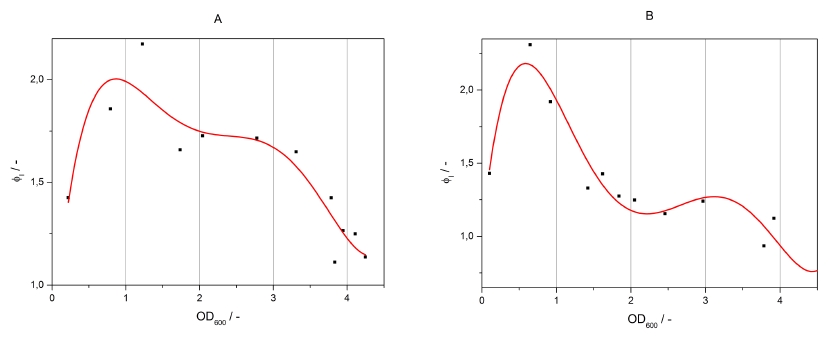
To measure the ratio of increasing promoter activity by inducing the system ɸI samples for analyzation should be taken at OD600 = 1 +/- 0.5. The highest ɸI in this range of the cultivation is taken for the calculation of the transfer function.
Plasmid conformation analysis
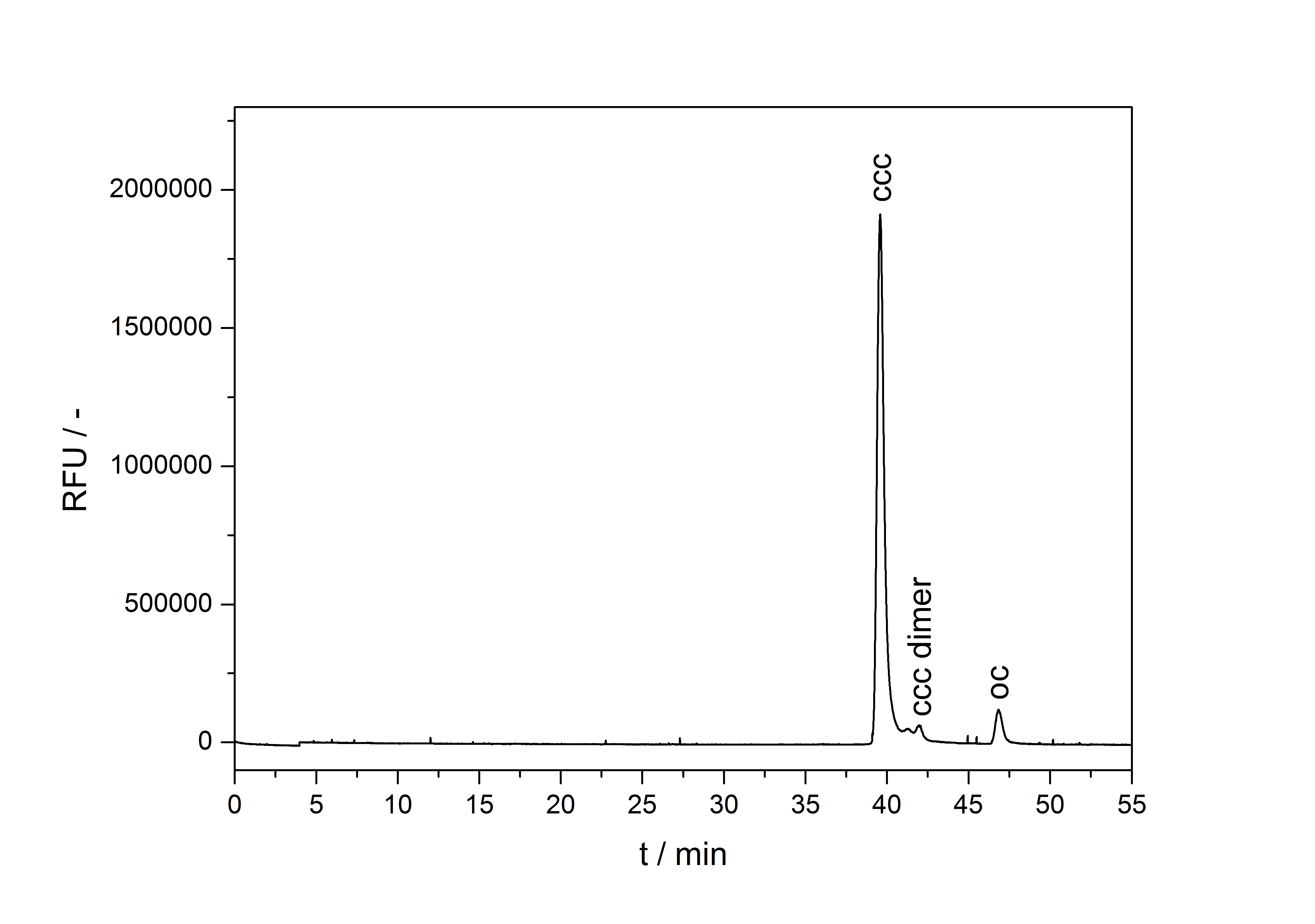
A plasmid conformation analysis for the BioBrick <partinfo>K389015</partinfo> in <partinfo>pSB1C3</partinfo> was performed by the [http://web.plasmidfactory.com/de/ PlasmidFactory] by Capillary Gel Electrophoresis (CGE). The chromatogram is shown in fig. 6 and the results in tab. 3. The data shows a high percentage of covalently closed circular (ccc) plasmid DNA. This is the biological active shape of plasmids so a high percentage of ccc plasmid DNA indicates a high quality of plasmid DNA ([http://web.plasmidfactory.com/en/service_CGE.html PlasmidFactory]).
| Conformation | Ratio / % |
|---|---|
| ccc monomer | 91 |
| ccc dimer | 3.7 |
| oc | 5.3 |
References
- Chu D, Zabet NR, Mitavskiy B (2009) [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WMD-4V42JG5-1&_user=10&_rdoc=1&_fmt=&_orig=search&_sort=d&_docanchor=&view=c&_acct=C000050221&_version=1&_urlVersion=0&_userid=10&md5=b6431553217aca1129c5b441f4b78425 Models of transcription factor binding: Sensitivity of activation functions to model assumptions], J Theor Biol 257(3):419-429.
- Greg Nelson, Jayaram Chandrashekar, Mark A. Hoon, Luxin Feng, Grace Zhao, Nicholas J. P. Ryba & Charles S. Zuker (2002) [http://www.nature.com/nature/journal/v416/n6877/abs/nature726.html An amino-acid taste receptor ], Nature 416: 199-202.
- [http://www.promega.com/tbs/tb281/tb281.pdf Promega Luciferase Assay System]
- [http://web.plasmidfactory.com/en/service_CGE.html PlasmidFactory Homepage]
 "
"