Team:Freiburg Bioware/NoteBook/Labjournal/October2
From 2010.igem.org
(→Mini-prep of mutual pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI) |
(→Biobrick assembly of Rep78 and Rep52) |
||
| Line 241: | Line 241: | ||
====<p style="font-size:15px; background-color:#66bbff;">Biobrick assembly of Rep78 and Rep52</p>==== | ====<p style="font-size:15px; background-color:#66bbff;">Biobrick assembly of Rep78 and Rep52</p>==== | ||
| - | Investigator: Kira <br /> | + | '''Investigator: Kira <br /> |
'''Comment:''' After replacing the mutated Rep parts by the ordered Rep parts, PCR amplification has to be done in order to produce a biobrick. | '''Comment:''' After replacing the mutated Rep parts by the ordered Rep parts, PCR amplification has to be done in order to produce a biobrick. | ||
PCR program: | PCR program: | ||
Revision as of 16:43, 19 October 2010
- March (labday 1)
- April (labday 2 - 5)
- May (labday 6 - 17)
- June (labday 18 - 45)
- July (labday 46 - 75)
- August part 1 (labday 76 - 92)
- August part 2 (labday 93 - 106)
- September part 1 (labday 107 - 123)
- September part 2 (labday 124 - 135)
- October part 1 (labday 136 - 145 )
- October part 2 (labday 146 - 155 )
- October part 3 (labday 156 - 166 )
- November (labday 167 - 170 )
- Cellculture
150. labday 15.10.2010
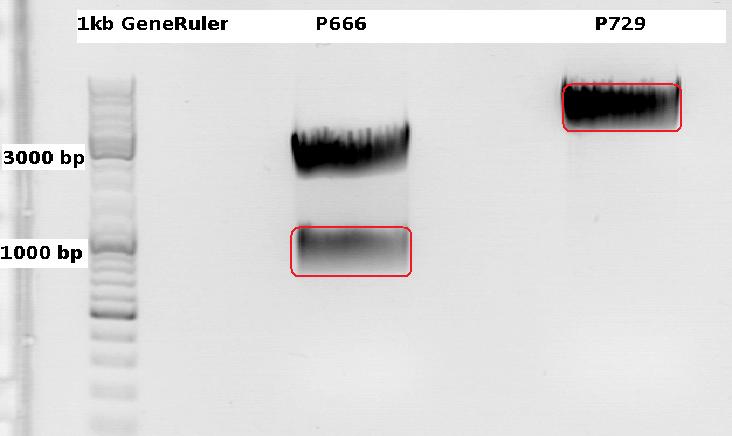
Cloning CFP (from P666: PSB1C3_CFP) into pSB1C3_leftITR_CMV_beta-globin (P729)
Investigator Patrick
Digestions, 2 h 10 minutes, 37 °C:
- P666: 5 µl DNA, 2 µl BSA, 2 µl Buffer 4 (10x), 1 µl Xba, 1 µl PstI, 9 µl H2O
- P729: 4 µl DNA, 2 µl BSA, 2 µl Buffer 4 (10x), 1 µl SpeI, 1 µl PstI, 10 µl H2O
Expected results for the 1% agarose gel:
- P666: about 2100 and 750 bp
- P729: about 3300 and 20 bp
The gelextraction ...
P728: 11,8 ng/µl
P822: 34,6 ng/µl
... and following ligation (2,5 µl Insert, 5,5 µl vector, 1 µl T4 DNA Ligase, 1 µl T4 DNA Ligase Buffer (10x), 40 minutes, RT) were performed according to the standard protocol. After the transformation (with XL1B) the cells were plated and put into the 37°C room.
Two additional transformations were performed with ligations from Volker labeled: "ligation viral brick 453 empty" & "viral brick 587 empty".
The following day the plates were checked for clones. Unfortunately there grew no clones on these two plates contrary to my plate with a a lot of clones.
Midi-Prep
Investigator: Chris W.
Midi-Prep of:
pSB1C3_001_RC_IRCK_P5tataless clone 1 =P866 =B516
pSB1C3_001_CMV_VP123_587-KO_Z34C_spacer clone2 =P867 =B526
pSB1C3_001_CMV_VP123_587-KO_Z34C clone2 =P868 =B529
pSB1C3_CMV_Zegfr:1907_MiddleLinker_VP2/3_587-KO_BAP clone 1 =P869 =B680
pSB1C3_CMV_Zegfr:1907_MiddleLinker_VP2/3_587-KO_6xHis clone 1 =P870 =B200
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | P866 | P867 | P868 | P869 | P870 |
| concentration (ng/µl) | 899,80 | 954,46 | 406,97 | 1642,76 | 1585,12 |
mini prep of several constructs
Investigator: Kira
c(rep52_1)=299,04 ng/ul
c(rep52_2)=290,07 ng/ul
c(rep78_1)=142,32 ng/ul
c(rep78_2)=175,36 ng/ul
Cell culture
Investigator: Kira
The cells are still alive. Medium was exchanged.--> RNA will be harvested tomorrow
Mini-Prep and test digestion of pSB1C3_CD_SDM-PstI_hGH_rITR
Investigator: Stefan
Glycerol stocks were prepared:
- B694 = pSB1C3_CD_SDM-PstI_hGH_rITR clone 1
- B695 = pSB1C3_CD_SDM-PstI_hGH_rITR clone 2
Mini-Prep was performed according to standard protocol:
- P875 = pSB1C3_CD_SDM-PstI_hGH_rITR clone 1 c = 73,1 ng/µl
- P876 = pSB1C3_CD_SDM-PstI_hGH_rITR clone 2 c = 78,3 ng/µl
Test digestion:
| Components | P875 + P876 / µl |
| DNA | 4 |
| Buffer 4 | 1 |
| BSA (10x) | 1 |
| XbaI | 0,3 |
| AgeI | 0,3 |
| H2O | 3,4 |
| Total volume | 10 |
Gel:
0,5g agarose, 50 ml TAE (1%), 3 µl GELRED, 115 Volt, running time ~50 minutes
Comment: Test digestion looks allright, cloning will be continued using P876.
151. labday 16.10.2010
Biobrick assembly: pSB1C3_lITR_CMV_ß-globin_CD_hGH_rITR and pSB1C3_lITR_phTERT_ß-globin_CD_hGH_rITR
Investigator: Achim
Plasmids:
- P729: pSB1C3_lITR_CMV_ß-Globin
- c= 243.4 ng/µl
- P730: pSB1C3_lITR_pHTERT_ß-Globin
- c= 81.1 ng/µl
- P876: pSB1C3_CD_SDM-PstI_hGH_rITR
- c= 78.3 ng/µl
Digestion:
| components | I1 (P729) | I2 (P730) | V (P876) |
| DNA | 6 | 14 | 14 |
| BSA (10x) | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 |
| Enzyme EcoI | 1 | 1 | 1 |
| Enzyme XbaI | - | - | 1 |
| Enzyme SpeI | 1 | 1 | - |
| H2O | 8 | - | - |
| Total | 20 | 20 | 20 |
Digestion: 2h, 37°C
Prep. gel:
- 0,8%, run for 45 min
- Corresponding bands were cut out
Gel ex.
- Nanodrop concentrations:
- I1: 37.54 ng/µl
- I2: 25.38 ng/µl
- V: 26.47 ng/µl
Ligation:
| ligation name | I1 + V | I2 + V |
| volume of vector | 4.69 | 4.23 |
| volume of insert | 3.31 | 3.77 |
| T4 ligase buffer (10x) | 1 | 1 |
| T4 ligase | 1 | 1 |
- Ligation @ RT for 40 min
Trafo:
- Done by Kira
Mini-prep of mutual pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI
Investigator Patrick
Yielded concentrations & given numbers:
- pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI clone 1: 208,4 ng/µl , P877 / B696
- pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI clone 2: 251,6 ng/µl , P878 / B696
- pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI clone 3: 212,6 ng/µl , P889 / B696
Test-digestion: 0,5 µl SpeI, 0,5 µl PstI, 3 µl DNA, 1 µl Buffer 4, 1 µl BSA, 4 µl H2O, 40 minutes, 37°C
Expected results: fragments with about 650 and 4900 bp
Obviously, this test digestion has to be repeated.
Preparations for tomorrow:
- Mini-prep of 3 mutual pSB1C3_leftITR_CMV_beta-globin_CFP clones (have to be picked from the plate)
- Midi-prep of pHelper
- Midi-prep of B689:pSB1C3_lITR_CMV_betaglobin_mGMK_TK30_SDM-PstI_hGH_rITR clone 2
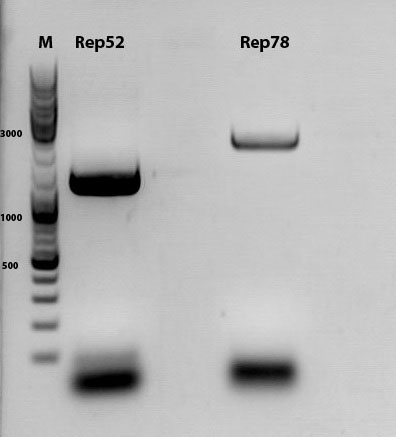
Biobrick assembly of Rep78 and Rep52
Investigator: Kira
Comment: After replacing the mutated Rep parts by the ordered Rep parts, PCR amplification has to be done in order to produce a biobrick.
PCR program:
c(Rep52)=299 ng/ul
c(Rep78)=175 ng/ul
Rep52: praefix 094 & suffix 097
Rep78: praefix 093 & suffix 097
| components | volume in µl |
| 5x Phusion HF buffer | 10 |
| 10 mM dNTP mix | 1 |
| primer_for (1:10 dilution) | 2,5 |
| primer_rev (1:10 dilution) | 2,5 |
| DNA template (1:100) | 0,5 |
| DMSO | 0,5 |
| Phusion polymerase | 0,5 |
| H2O | 32,5 |
| Total volume (e.g. 50 µl) | 50 |
| Cycles | Temperature | Time |
| 98°C | 30 sec | |
| 10x | 98°C | 15 sec |
| 63°C | 25 sec | |
| 72°C | 32 sec | |
| 20x | 98°C | 15 sec |
| 66°C | 25 sec | |
| 72°C | 32 sec | |
| 1x | 72°C | 5 min |
| Hold 4°C |
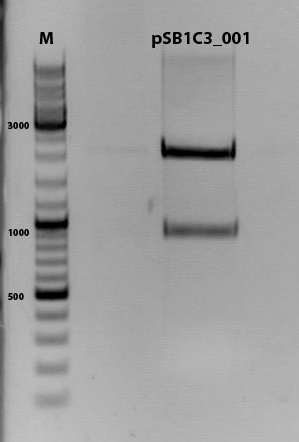
Digestion of plasmid backbone:
pSB1C3_001 is used as backbone
| Components | <b>vector Volume/µL |
| DNA | 3,5 µl |
| BSA (10x) | 2 µl |
| Buffer no. 4 (10x) | 2,0 µl |
| Enzyme 1 EcoRI-HF | 0,5 µl |
| Enzyme 2 SpeI | 1,0 µl |
| H2O | 15 µl |
| Total volume | 25 |
incubation @ 37 C for approx. 2 h
1% agarose gel
Digestion of PCR product:
| Components | PCR product Volume/µL |
| DNA | 35,0 µl |
| BSA (100x) | 0,45 µl |
| Buffer no. 4 | 4,5 µl |
| Enzyme 1 EcoRI-HF | 1,5 µl |
| Enzyme 2 SpeI | 2,0 µl |
| H2O | 1,5 µl |
| Total volume | 45 |
incubation @ 37 C for approx. 2 h
T4 ligation for 40 min
Transformation according to the standard protocol
RNA harvesting
Investigator: Kira
After transfection, the cells were incubated for 48 hours. Today, the cells will be harvested and RNA extracted, in order to perform RT-PCR and an additional PCR for evaluation of promoter activity.
The transfected cells were trypsinised and centrifuged for 2 min. The supernatant was discarded and pellet washed 2x with PBS. RNeasy Kit [Qiagen] was used for RNA extraction according to the manufacturer protocol.
c(CMV)= 335,69 ng/ul
c(P40)= 857,92 ng/ul
c(AAV_RC)= 760,21 ng/ul
152. labday 17.10.2010
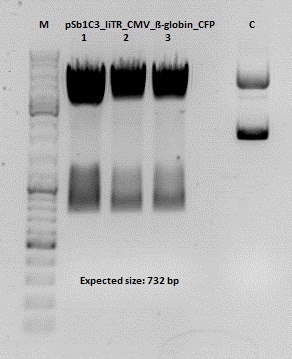
Test digestion of pSB1C3_lITR_CMV_ß-globin_CFP
Investigator: Anna
Vector name:
pSB1C3_lITR_CMV_betaglobin_CFP_cl1 (P880): c = 452,67 ng/µl
pSB1C3_lITR_CMV_betaglobin_CFP_cl2 (P881): c = 288,88 ng/µl
pSB1C3_lITR_CMV_betaglobin_CFP_cl3 (P882): c = 288,36 ng/µl
Test Digestion:
| components | volume P880 - P882 /µl | volume P434 /µl |
| DNA | 2 | 2 |
| BSA (10x) | 1 | 1 |
| Buffer 4 (10x) | 1 | 1 |
| Enzyme NgoMIV | 0,3 | 0,3 |
| Enzyme AgeI | 0,3 | 0,3 |
| H2O | 5,4 | 5,4 |
| Total volume | 10 | 10 |
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
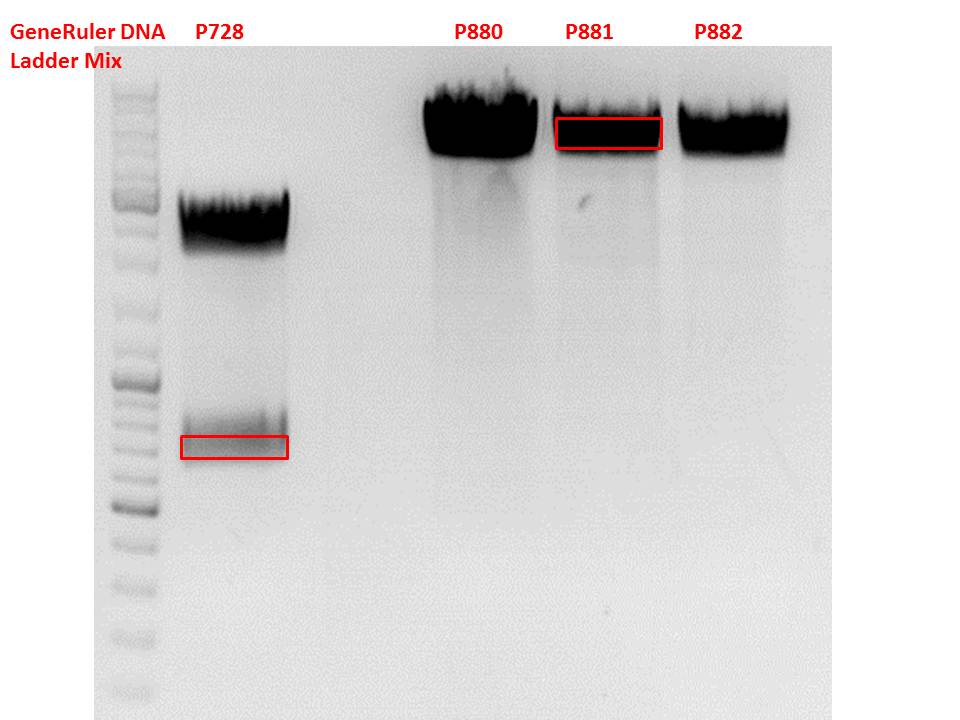
Cloning of hGH_rITR into pSB1C3_lITR_CMV_betaglobin_CFP
Investigator: Stefan
Cloning of our last GOI!
Vector name:
pSB1C3_lITR_CMV_betaglobin_CFP cl 1-3 (P880-P882)
Insert name:
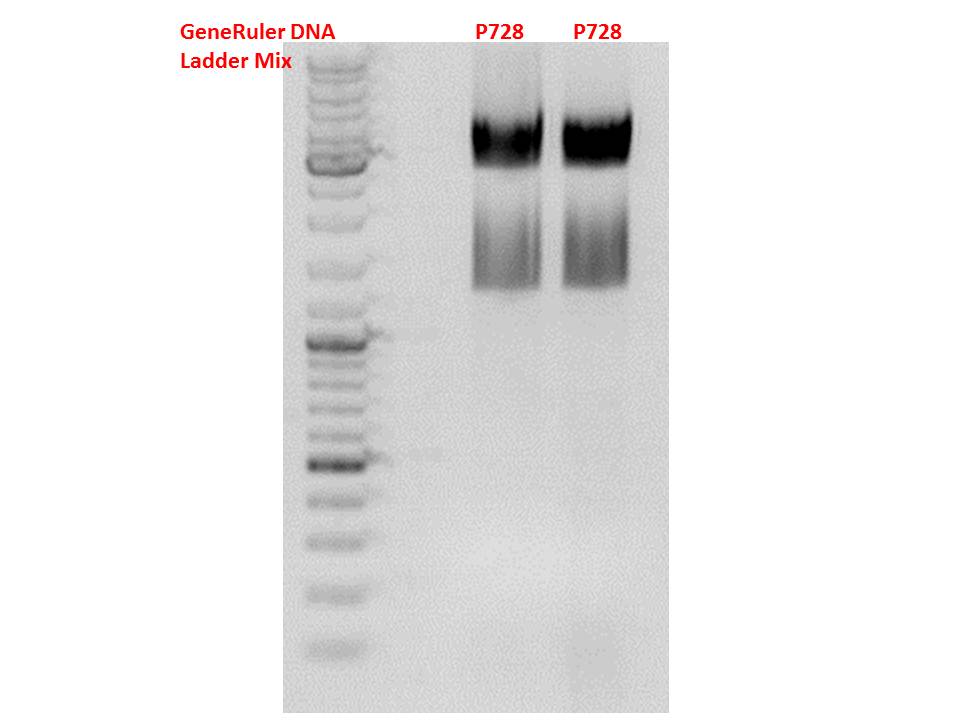
pSB1C3_hGH_rITR (P728)
Digestion:
| components | volume P880 - P882 /µl | volume P728 /µl |
| DNA | 3 | 8 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme PstI | 1 | 1 |
| Enzyme XbaI | - | 1 |
| Enzyme SpeI | 1 | - |
| H2O | 11 | 4 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 |
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
Test digestion of all constructs looked alright, therefore, cloning was continued using P881 only.
Gel extraction:
Was performed according to protocol.
T4 Ligation:
| ligation name | 728 + 881 |
| volume of vector | 2,68 |
| volume of insert | 5,32 |
| T4 ligase buffer (10x) | 1 |
| T4 ligase | 1 |
Transformation:
Transformation was performed according to standard protocol using BL21 cells.
RT-PCR
Investigator: Kira
For further experiments, RNA has to be translated into cDNA. The PCR was performed according to the manufacturer protocol.
153. labday 18.10.2010
quantitative real-time PCR for detection of virus titer
Investigator: Achim
- qPCR of harvested virus particles to determine the virus titers of our different constructs
- Total number of samples: 58
Test-digestion of pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI clone 1, 2 and 3 (P877, 878 and 879)
Investigator Patrick
Check the plasmid for leftITR:
0,5 µl EcoRI, 1 µl BstEII, 7 µl DNA, 2 µl Buffer 4, 2 µl BSA, 7,5 µl H2O, 45 minutes 37°C, 45 minutes 60°C
Check the plasmid for hGH_rITR and ... :
0,5 µl AgeI, 0,5 µl PstI, 3 µl DNA, 1 µl BSA, 1 µl Buffer 4, 4 µl H2O, 70 minutes 37°C
Expected results:
- leftITR: about 190 bp
- hGH_rITR: about 670 bp
Unfortunately the digestions had to be reapeated because i didnt switch on the current so the samples and especially the 1kb GeneRuler marker diffused.
The second run: see above
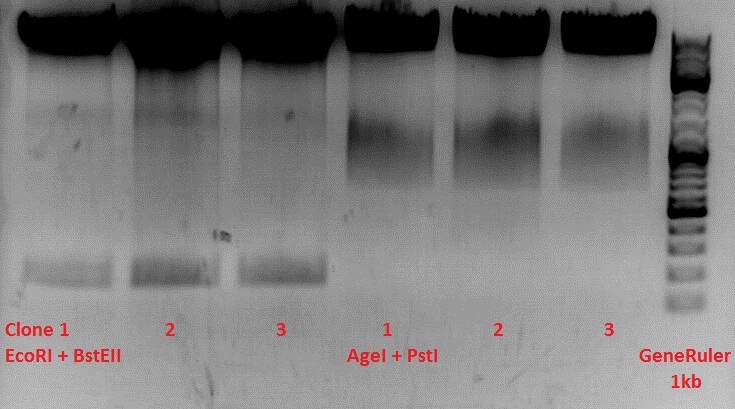
PstI or BstEII seems to work not properly
Mini-Prep and test digestion of several constructs
Investigator: Jessica
Glycerol stocks were prepared:
- B702 = pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR clone 1
- B703 = pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR clone 2
- B704 = pSB1C3_001_VP3 clone 1
- B705 = pSB1C3_001_VP3 clone 2
- B706 = pSB1C3_001_VP2 clone 1
- B707 = pSB1C3_001_VP2 clone 2
- B708 = pSB1C3_001_Rep78 clone 1
- B709 = pSB1C3_001_Rep78 clone 2
- B710 = pSB1C3_001_Rep52 clone 1
- B711 = pSB1C3_001_Rep52 clone 2
- B712 = pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR clone 1
- B713 = pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR clone 2
- B714 = pSB1C3_001_VP1 clone 1
- B715 = pSB1C3_001_VP1 clone 2
Mini-Prep was performed according to standard protocol:
- P886 = pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR clone 1 c= 232,2ng/µl
- P887 = pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR clone 2 c= 186,2ng/µl
- P888 = pSB1C3_001_VP3 clone 1 c= 300,8ng/µl
- P889 = pSB1C3_001_VP3 clone 2 c= 284,4ng/µl
- P890 = pSB1C3_001_VP2 clone 1 c= 298,3ng/µl
- P891 = pSB1C3_001_VP2 clone 2 c= 299,9ng/µl
- P892 = pSB1C3_001_Rep78 clone 1 c= 143,9ng/µl
- P893 = pSB1C3_001_Rep78 clone 2 c= 163,4ng/µl
- P894 = pSB1C3_001_Rep52 clone 1 c= 166,6ng/µl
- P895 = pSB1C3_001_Rep52 clone 2 c= 181,6ng/µl
- P896 = pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR clone 1 c= 250,5ng/µl
- P897 = pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR clone 2 c= 173,4ng/µl
- P898 = pSB1C3_001_VP1 clone 1 c= 272,7ng/µl
- P899 = pSB1C3_001_VP1 clone 2 c= 294,8ng/µl
- P900 = pSB1C3_hGH_rITR (from B160) c= 136,7ng/µl
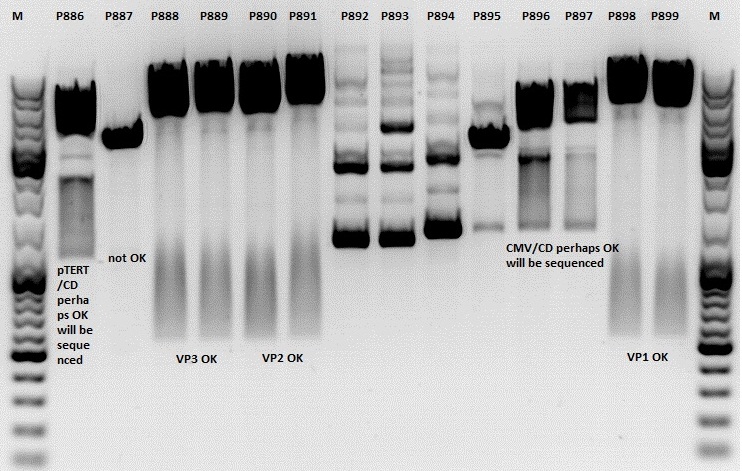
Test digestion:
| Components | P886,887,892,893,894,895,896,897 / µl | P888,889,890,891898,899 / µl |
| DNA | 1,5 | 1,5 |
| Buffer | (4) 1 | (2) 1 |
| BSA (10x) | 1 | 1 |
| NgoMIV | 0,4 | - |
| XbaI | 0,4 | - |
| PstI | - | 0,6 |
| XcmI | - | 0,4 |
| H2O | 4,5 | 4,5 |
| Total volume | 10 | 10 |
Gel:
1,0g agarose, 100 ml TAE (1%), 6 µl GELRED, Volt, running time minutes
Comment: Rep 52/78 will be checked,pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR and pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR will be sequenced
PCR of mGMK and SR39
Investigator: Anna
Plasmids:
pSB1C3_mGMK_TK30_SDM-PstI clone 2(P804)
pSB1C3_mGMK_sr39 clone 1(P860)
Oligos:
O193: pTK30_for
O81: pmgmk_tk30_suffix_RFC25_rev
PCR Mix:
| Components | Volume /µl |
| Phusion Buffer | 10 |
| dNTP | 1 |
| Primer_for | 2,5 |
| Primer_rev | 2,5 |
| DNA template | 1 |
| H2O | 32,5 |
| Total volume | 50 |
PCR Program:
| Cycles | Temperature | Time |
| 98°C | 60 sec | |
| 98°C | 15 sec | |
| 8x | 52°C | 25 sec |
| 72°C | 25 sec | |
| 98°C | 15 sec | |
| 17x | 67°C | 25 sec |
| 72°C | 25 sec | |
| 1x | 72°C | 5 min |
| Hold 4°C |
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
[[Image:|550px|]]
154. labday 19.10.2010
Midi-Prep
Investigator: Chris W.
Midi-Prep of:
pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone1 =P901 =B200
pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI clone 2 =P902 =B697
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | P901 | P902 |
| concentration (ng/µl) | 1563,63 | 1348,26 |
Continuation of PCR of mGMK and SR39
Investigator: Jessica
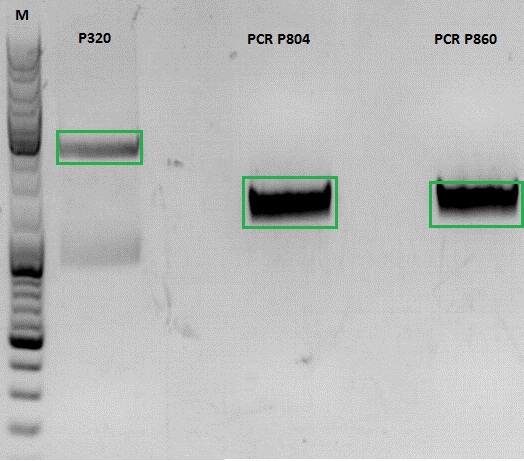
- vector (P320) and PCR product was digested
| Components | P320 / µl | PCR product P804 and P860 / µl |
| DNA | 1,5 | 20 |
| Buffer | 2 | 3 |
| BSA (10x) | 2 | 3 |
| AgeI | 1 | 1,5 |
| XbaI | 1 | 1,5 |
| H2O | 14,5 | 1 |
| Total volume | 20 | 30 |
Ligation
- P320 c= 5,08 ng/µl
- P804 c= 11,73 ng/µl
- P860 c= 26,57 ng/µl
- P320 + P804: 4,93µl : 3,07µl
- P320 + P860: 6,28µl : 1,72µl
Transformation with BL21 and Cm
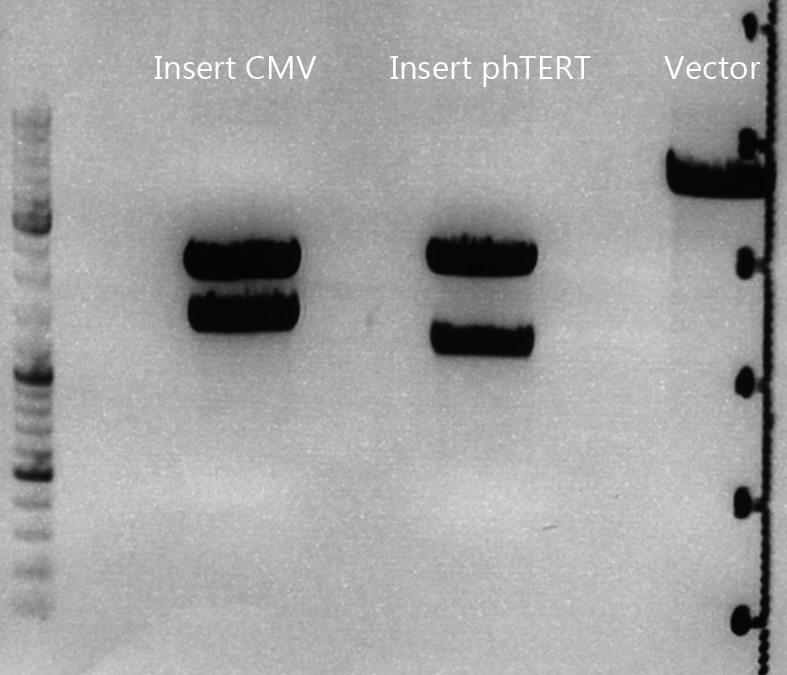
Cloning of CMV into pSB1C3_001_VP1, pSB1C3_001_VP2 and pSB1C3_001_VP3
Investigator: Anna
Plasmids:
- P888: pSB1C3_001_VP3
- c = ng/µl
- P890: pSB1C3_001_VP2
- c = ng/µl
- P898: pSB1C3_001_VP1
- c = ng/µl
Digestion:
| components | I1 (P729) | I2 (P730) | V (P876) |
| DNA | 6 | 14 | 14 |
| BSA (10x) | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 |
| Enzyme EcoI | 1 | 1 | 1 |
| Enzyme XbaI | - | - | 1 |
| Enzyme SpeI | 1 | 1 | - |
| H2O | 8 | - | - |
| Total | 20 | 20 | 20 |
Digestion: 2h, 37°C
Prep. gel:
- 0,8%, run for 45 min
- Corresponding bands were cut out
Gel ex.
- Nanodrop concentrations:
- I1: 37.54 ng/µl
- I2: 25.38 ng/µl
- V: 26.47 ng/µl
Ligation:
| ligation name | I1 + V | I2 + V |
| volume of vector | 4.69 | 4.23 |
| volume of insert | 3.31 | 3.77 |
| T4 ligase buffer (10x) | 1 | 1 |
| T4 ligase | 1 | 1 |
- Ligation @ RT for 40 min
Trafo:
- Done by Kira
155. labday 20.10.2010
156. labday 21.10.2010
157. labday 22.10.2010
158. labday 23.10.2010
159. labday 24.10.2010
160. labday 25.10.2010
161. labday 26.10.2010
162. labday 27.10.2010
163. labday 28.10.2010
164. labday 29.10.2010
165. labday 30.10.2010
166. labday 31.10.2010
 "
"