Team:Paris Liliane Bettencourt/Project/Population counter/Results
From 2010.igem.org
| Line 12: | Line 12: | ||
</a> | </a> | ||
<a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/SIP"> | <a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/SIP"> | ||
| - | <img src="https://static.igem.org/mediawiki/2010/4/4c/SIP.png" width=" | + | <img src="https://static.igem.org/mediawiki/2010/4/4c/SIP.png" width="132" height="107" align=right title="SIP"> |
</a> | </a> | ||
<a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Memo-cell"> | <a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Memo-cell"> | ||
| - | <img src="https://static.igem.org/mediawiki/2010/a/aa/Memo_cell-01.jpg" width=" | + | <img src="https://static.igem.org/mediawiki/2010/a/aa/Memo_cell-01.jpg" width="129" height="107" align=right title="Memo-Cell"> |
</a> | </a> | ||
<br /> | <br /> | ||
Revision as of 23:14, 27 October 2010
Contents |
Construction of our counters
First of all, we spent a significant part of the summer to do cloning using the standard biobrick assembly protocol. For this, we have used a lot of biobricks submitted to the parts registry by previous teams, thank you! Using these parts we have created plenty of new composite parts that are highly modular (different kind of RBS and promoters) and could serve next iGEM teams. We have also created some new biobricks that are at the core of our project. We hope other iGEM teams will work with integrons and might need these parts. We have constructed both the basic counter (with RFP and Tetracycline resistance gene) and the full counter + timer designs.
Moreover, each of our constructs was verified by sequencing. For more information you can take a look at the parts that we sent to the registry.
Proof that IntI1 integrase works !
Then we wanted to see if the IntI1 integrase actually works. We have done a double transformation "RFP Counter" (on pSB1A2 which is a high copy plasmid) + "pBAD-IntI1" (on pSulib which is a low copy plasmid with kanamycin resistance) to obtain a working counter. The protocol is quite simple : we dilute an overnight culture, wait until OD 0,2 and add arabinose to begin the test (t=0h). At regular time intervals, we plated on plates containing ampicillin (this is not important if the cells loose the "pBad-Int" plasmid because we don't want recombinations to happen on the plates).
This experiment shows us that IntI1 is effective ! The double terminator flanked by attC sites has been excised and the constitutive promoter upstream permits the expression of RFP. However, we have 2 important notes :
- At the beginning of the test, there already colonies that are fluorescent... We can explain this fact by the leaking of pBad that control the expression of the integrase. Thereby the integrase is expressed at a low level and catalyzes the excision of the terminator.
- The expression of RFP is inhomogeneous because the counter is on a high copy plasmid. So some cells have 2 recombined plasmids while others have 20 recombined plasmids !
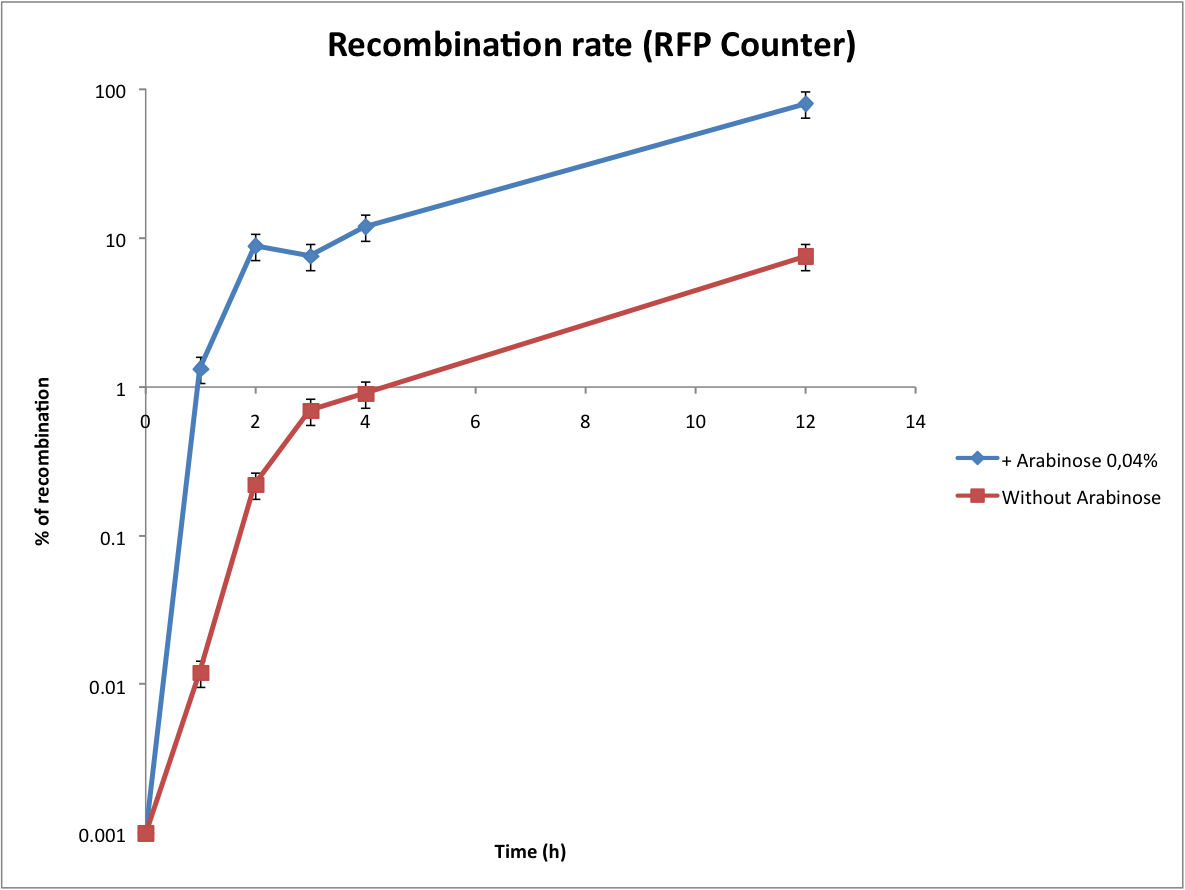
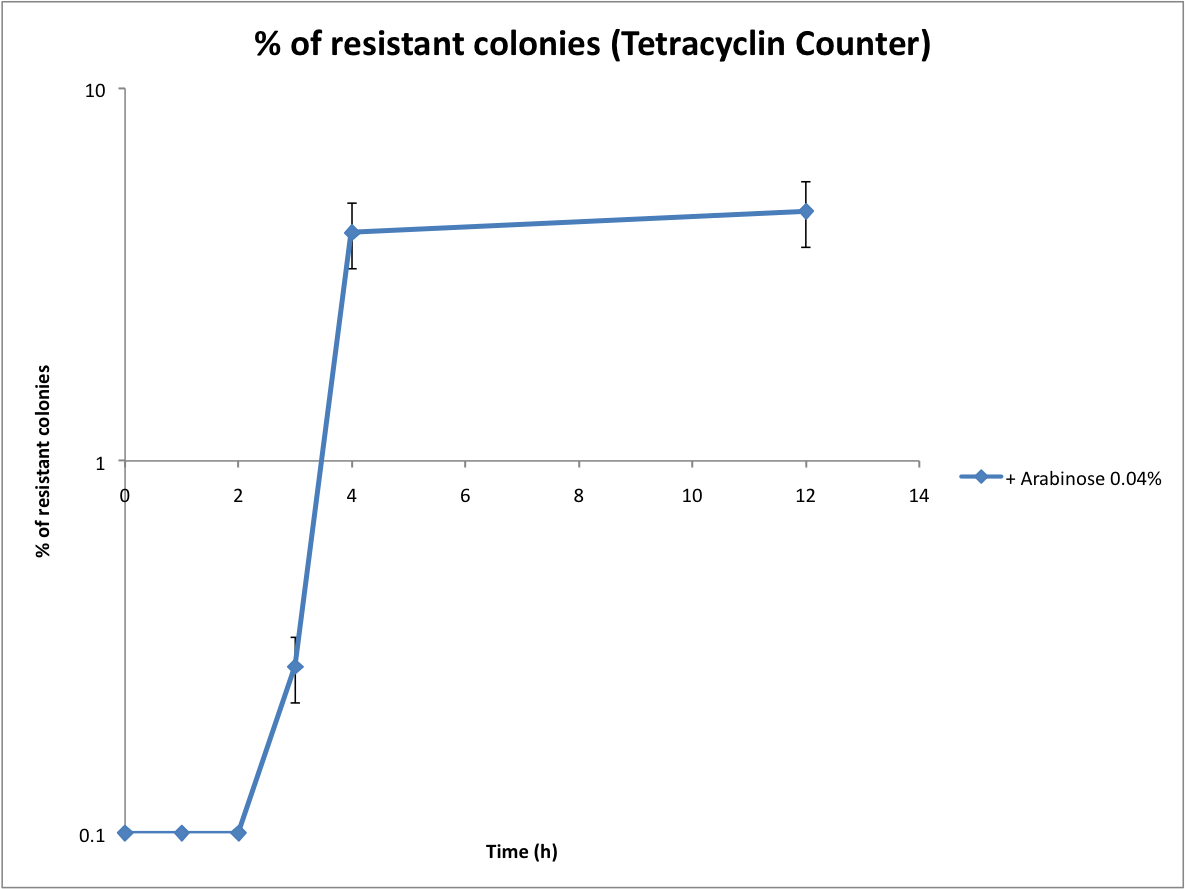
Characterization of the recombination rate
on a plate containing ampicillin, kanamycin and 1% glucose (which blocks the pBAD promoter and avoid recombination events to occur before the test of our counter). and 1% glucose (without glucose otherwise pBad cannot be activated)
- Check the excision of the terminator by sequencing


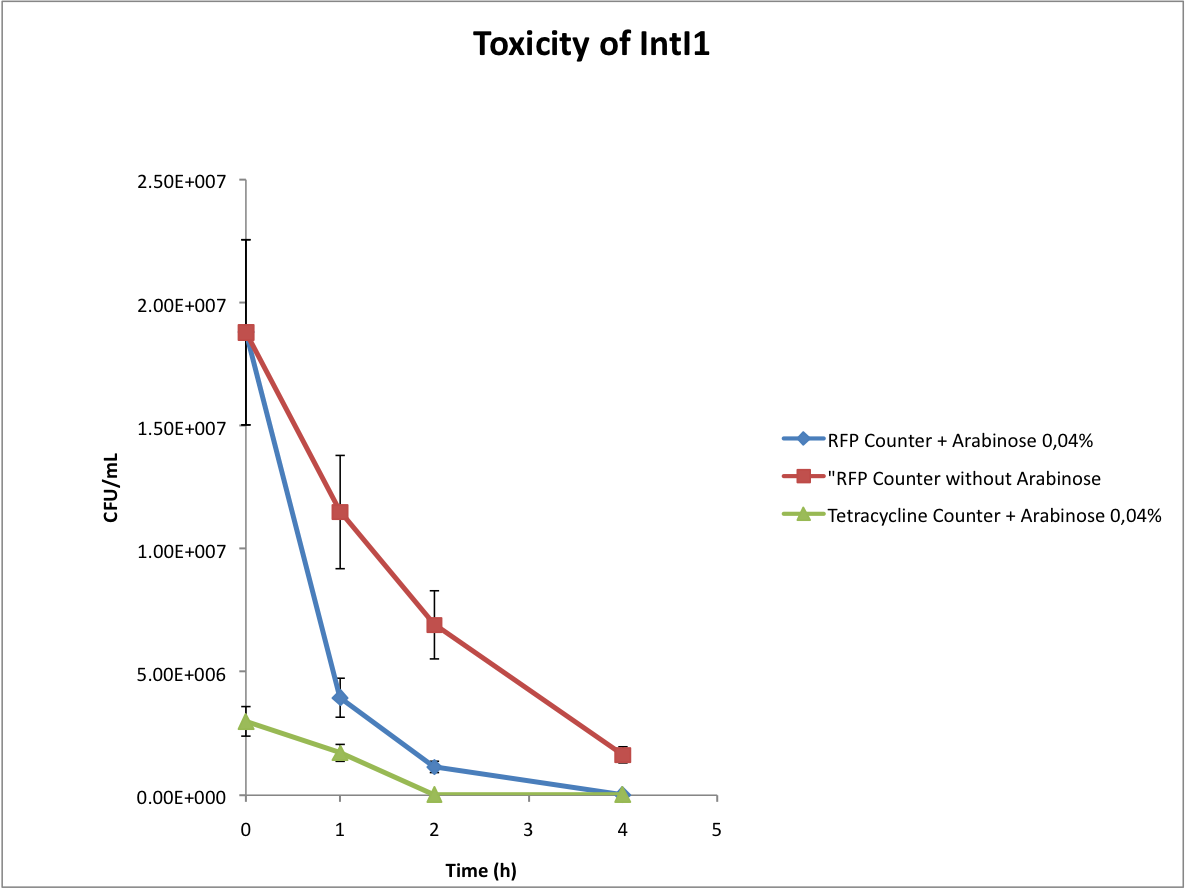
Verify the toxicity of the integrase
 "
"