Team:Tokyo Tech/Project/Artificial Cooperation System
From 2010.igem.org
(→Works) |
|||
| (162 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
<div id="tf_menu"> | <div id="tf_menu"> | ||
| - | menu | + | <font size="5" color="#eb8300"><b><center>Project menu</center></b></font> |
| + | |||
| + | <center> | ||
| + | <table id="table-01"> | ||
| + | <tr> | ||
| + | <td>[[Team:Tokyo_Tech|1 Graphic abstract]]<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <td>2 Apple reporter<br> | ||
| + | :[[Team:Tokyo_Tech/Project/Apple_Reporter|2-1 Color]] | ||
| + | :[[Team:Tokyo_Tech/Project/Apple_Reporter2|2-2 Fragrance]] | ||
| + | </td> | ||
| + | <tr> | ||
| + | <th>3 Artificial Cooperation System -YOU ARE HERE!-<br> | ||
| + | :[[Team:Tokyo_Tech/Project/Artificial_Cooperation_System/lux_act_rep|3-1 ''lux'' activation/repression promoter]] | ||
| + | :[[Team:Tokyo_Tech/Project/Artificial_Cooperation_System/Cm_assay|3-2 resistance gene activation device]] | ||
| + | :[[Team:Tokyo_Tech/Project/Artificial_Cooperation_System/luxI_assay|3-3 ''lux''I Assay]] | ||
| + | :[[Team:Tokyo_Tech/Project/Artificial_Cooperation_System/modeling|3-4 modeling]] | ||
| + | </th> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>[[Team:Tokyo_Tech/Project/wolf_coli|4 Wolf coli overview]]<br> | ||
| + | :[[Team:Tokyo_Tech/Project/wolf_coli/New_Series_of_PompC|4-1 New seriesof P''ompC'']] | ||
| + | :[[Team:Tokyo_Tech/Project/wolf_coli/lacIM1|4-2 lacIM1 for band-detect network ]] | ||
| + | :[[Team:Tokyo_Tech/Project/wolf_coli/System|4-3 Wolf coli system]] | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </center> | ||
| + | |||
| + | |||
</div> <!-- tf_menu --> | </div> <!-- tf_menu --> | ||
| - | <div id="tf_SubWrapper"> | + | <div id="tf_SubWrapper"> |
| - | ==Artificial Cooperation System | + | <font size="5"><b>3 Artificial Cooperation System</b></font> |
| - | = | + | |
| - | + | =Artificial Cooperation System= | |
| - | + | =Abstract= | |
| - | + | We built Artificial Cooperation System to endow E.coli with humanity. In this system quorum sensing system and chloramphenicol resistance protein generator activated by LuxR are very important. First, we constructed and characterized several promoters regulated by LuxR and 3OC6HSL. We designed and constructed a new LuxR repression promoter. Also, we characterized the functions of a LuxR repression promoter and a LuxR activation promoter in the registry. Second, we constructed a new chloramphenicol resistance protein generator which is regulated by LuxR activation promoter. We confirmed that the cell with the chloramphenicol resistance protein generator can survive under high concentration of chloramphenicol when 3OC6HSL exists. Third, we confirmed that LuxI can produce sufficient 3OC6HSL to activate chloramphenicol resistance gene. Forth, we confirmed the feasibility of Artificial Cooperation System from simulation and experimental data. | |
| - | + | ||
| + | [[IMAGE:Tokyotech_plux_act_final.jpg|300px|left|thumb|fig.3-1-1 luxR activation promoter assay (worked by Kitano Shohei & Eriko Uchikoshi)]] | ||
| + | |||
| + | [[IMAGE:Tokyotech_plux_rep_final.jpg|300px|right|thumb|fig.3-1-3 luxR repression promoter assay (worked by Shohei Kitano & Eriko Uchikoshi)]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | [[Image:tokyotech_Cm-survival.jpg|300px|left|thumb|Fig3-2-1 Chloramphenicol resistance device. This work done by Yusuke Kaneta]] | ||
| + | [[IMAGE:Tokyotech Fig.K395146-1.jpg|300px|left|thumb|Fig3-3-1 LuxI and LuxR activation promoter assay. This work done by Eriko Uchikoshi and Shohei Kitano]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | =Introduction= | ||
| + | ==What’s Artificial Cooperation System?== | ||
| + | Artificial Cooperation System was built to engineer ''E. coli'' that behaves kindly and have sympathy. In our idea of ''E. coli'' with humanity, there are two types of cells. In normal conditions, they are competitors for survival. While in critical conditions, such that one is dying, another cell would notice the crisis of dying cell and help her. For realization of our idea, we constructed a system that named “Artificial Cooperation System”. | ||
| + | |||
| + | [[Image:Tokyotech_top10.jpg|center|640px|thumb|Fig 3-0-1.They are competitors. 2.One is dying, another cell notices it. 3.The healthy cell rescues the dying cell. 4. The dying cell recovers and she gives "Apples" as expression of appreciation]] | ||
| + | |||
| + | ==Genetic Circuit== | ||
| + | We designed the following circuit. In Artificial Cooperation System, there are two types of ''E. coli'', Cell A and Cell B. Cell A has resistance to kanamycin and Cell B has resistance to chloramphenicol originally. In cell A (Fig 3-0-2), production of LasI is repressed by LuxR/3OC6HSL complex and chloramphenicol resistance gene is activated by LuxR/3OC6HSL complex. In cell B (Fig3-0-3), production of LuxI is repressed by LasR/3OC12HSL complex and kanamycin resistance gene is activated by LasR/3OC12HSL complex. This means that 3OC12HSL produced by cell A and 3OC6HSL produced by cell B indirectly repress each other’s production. On the other hand, 3OC12HSL activates resistance gene of cell B and 3OC6HSL activates resistance gene of cell A. Apple gene is composed of crtEBIYZW and MpAAT1. This expresses only when the cell is rescued and recovered. | ||
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <embed | ||
| + | src="https://static.igem.org/mediawiki/2010/5/53/Tokyotech_ACS.swf" | ||
| + | width="500" | ||
| + | height="375" | ||
| + | /> | ||
| + | </center> | ||
| + | </html> | ||
| + | |||
| + | |||
| + | |||
| + | [[IMAGE:Tokyotech cellA circuit.png|400px|left|thumb|Fig3-0-2.Genetic circuit in cell A]] | ||
| + | [[IMAGE:Tokyotech cellB corcuit.png|400px|left|thumb|Fig3-0-3.Genetic circuit in cell B ]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
Pcon: constitutive promoter.<br> | Pcon: constitutive promoter.<br> | ||
| Line 32: | Line 158: | ||
Plas act: promoter activated by LasR and 3OC12HSL. We call this promoter lasR activation promoter.<br> | Plas act: promoter activated by LasR and 3OC12HSL. We call this promoter lasR activation promoter.<br> | ||
Plas rep: promoter repressed by LasR and 3OC12HSL. We call this promoter lasR repression promoter.<br> | Plas rep: promoter repressed by LasR and 3OC12HSL. We call this promoter lasR repression promoter.<br> | ||
| + | lasI: enzyme repressed by LuxR and 3OC6HSL. We call this promoter LuxR repression promoter.<br> | ||
| + | luxI: enzyme repressed by LasR and 3OC12HSL. We call this promoter LasR repression promoter.<br> | ||
| + | |||
| + | |||
| + | |||
| + | ---- | ||
| + | <FONT SIZE="4">Normal</FONT> | ||
| + | |||
| + | [[IMAGE:tokyotech_normal.jpg|400px|left|thumb|Fig3-0-4. Normal condition]] | ||
| + | [[IMAGE: Tokyotech ppt normal.jpg|200px|thumb|Fig3-0-5. Normal condition]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | In normal situation, Cell A and Cell B are competitors and recognize each other by using quorum sensing. First, LasI protein in Cell A produces 3OC12HSL. Second, these signal molecules diffuse through cell membranes and bind to LasR protein in Cell B. Third, LasR/3OC12HSL complex represses the production of LuxI protein. Similarly in cell B, LuxI protein produces 3OC6HSL. These signal molecules diffuse through cell membranes and bind to LuxR protein in Cell A. LuxR/3OC6HSL complex represses the production of LasI protein. Although Cell A and Cell B have chloramphenicol and kanamycin resistance gene respectively, they don’t express in this situation. This is because the concentration of 3OC12HSL and 3OC6HSL is too low to activate the expressions of kanamycin resistance gene and chloramphenicol resistance gene, respectively.<br> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <br> | ||
| + | ---- | ||
| + | <FONT SIZE="4"> when Cell A is dying </FONT> | ||
| + | |||
| + | [[IMAGE:tokyotech_cell A is dyng.jpg|400px|left|thumb|Fig 3-0-6. Cell A is dying.]] | ||
| + | [[IMAGE: Tokyotech ppt cellA is dying.jpg|200px|thumb|Fig 3-0-7. Cell A is dying.]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | As we mentioned, originally Cell A has a resistance gene against kanamycin and does not have that against chloramphenicol. On the other hand, Cell B has resistance to chloramphenicol and doesn’t have resistance to kanamycin. Therefore, the addition of chloramphenicol only decreases the number of Cell A. It means that 3OC12HSL decreases. It also results in the decrease of LasR/3OC12HSL complex that repress the expression of LasI in Cell B. By this process, Cell B notices that Cell A is dying and tries to rescue it.<br> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ---- | ||
| + | <FONT SIZE="4"> when Cell B tries to rescue Cell A </FONT> | ||
| + | |||
| + | [[IMAGE:tokyotech_rescue.png|400px||left|thumb|Fig 3-0-8. Cell B is trying to rescue cell A]] | ||
| + | [[IMAGE: Tokyotech ppt cellB try.jpg|200px|thumb|Fig 3-0-9. Cell B is trying to resucue cell A]] | ||
| + | |||
| + | |||
| + | |||
| + | [[IMAGE:tokyotech_apple_vol2.jpg|400px|left|thumb|Fig 3-0-10. Cell A is rescued by cell B.]] | ||
| + | [[IMAGE: Tokyotech ppt cellA is helped.jpg|200px|thumb|Fig 3-0-11. Cell A is rescued by cell B.]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | As we mentioned, the expression of ''luxI'' is repressed by LasR/3OC12HSL complex. Therefore, the decrease of 3OC12HSL/LasR complex causes the overexpression of ''luxI'' and the increase of 3OC6HSL. This results in the expression of chloramphenicol resistance gene in the Cell A. Consequently, the Cell A is recovered and begins to produce the apple reporters in return! | ||
| + | |||
| + | =Result= | ||
| + | In order to make the Artificial Cooperation System, we constructed and characterized several important parts and devices. First we characterized LuxR repression promoter R0061 and activation promoter R0062, both of which are existing parts. Second, we designed and characterized a new LuxR repression promoter [http://partsregistry.org/Part:BBa_K395008 K395008]. Third, we designed and characterized a NEW BioBrick device [http://partsregistry.org/Part:BBa_K395162 K395162], a chloramphenicol resistance generator, which is activated by a LuxR activation promoter. Forth, we confirmed that LuxI can produce sufficient AHL to help dying cell. Finally, we simulated our Artificial Cooperation System and confirmed the feasibility of our system. | ||
| + | |||
| + | == LuxR repression promoter and activation promoter== | ||
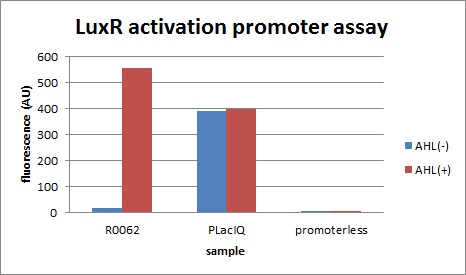
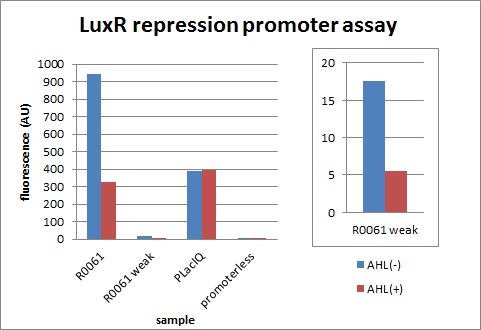
| + | We characterized LuxR repression promoters and activation promoters, which play important roles in the Artificial Cooperation System. The following figures are the result of experiments. Fig3-1-1 shows that under the regulations by R0062 (LuxR activation promoter), GFP expression was activated in the presence of 3OC6HSL. Fig3-1-3 shows that under the regulations by R0061 and K395008, GFP expressions were repressed in the presence of 3OC6HSL. Also, for realization of the Artificial Cooperative System work, the strength and the threshold of these promoters are so important. Thus we tried tuning promoters by changing sequence of consensus -35/-10 and lux box. [[Team:Tokyo_Tech/Project/Artificial_Cooperation_System/lux_act_rep|(See more…)]] | ||
| + | |||
| + | [[IMAGE:Tokyotech_plux_act_final.jpg|300px|left|thumb|fig.3-1-1 luxR activation promoter assay (worked by Kitano Shohei & Eriko Uchikoshi)]] | ||
| + | |||
| + | [[IMAGE:Tokyotech_plux_rep_final.jpg|300px|left|thumb|fig.3-1-3 luxR repression promoter assay (worked by Shohei Kitano & Eriko Uchikoshi)]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ==Chloramphenicol resistance geneator activated by LuxR== | ||
| + | We succeeded in construction and characterization of a NEW Biobrick device of a chloramphenicol resistance (''CmR'') generator | ||
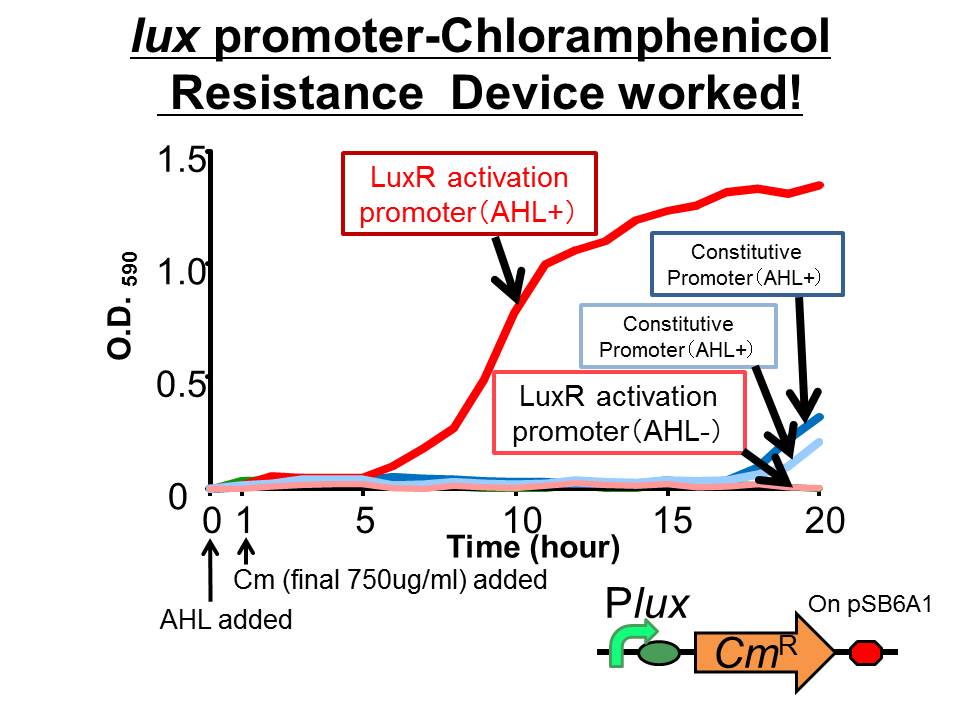
| + | ([http://partsregistry.org/Part:BBa_K395162 BBa_K395162]) which is activated by LuxR activation promoter. Fig 3-2-1 shows that in the presence of 3OC6HSL, the device is activated and as a result the cell was able to survive under high chloramphenicol concentration (750 ug/ml). [[Team:Tokyo_Tech/Project/Artificial_Cooperation_System/Cm_assay|(See more…)]] | ||
| + | |||
| + | [[Image:tokyotech_Cm-survival.jpg|400px|left|thumb|Fig3-2-1 Chloramphenicol resistance device. This work done by Yusuke Kaneta]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ==LuxI can produce sufficient 3OC6HSL to activate LuxR activation promoter== | ||
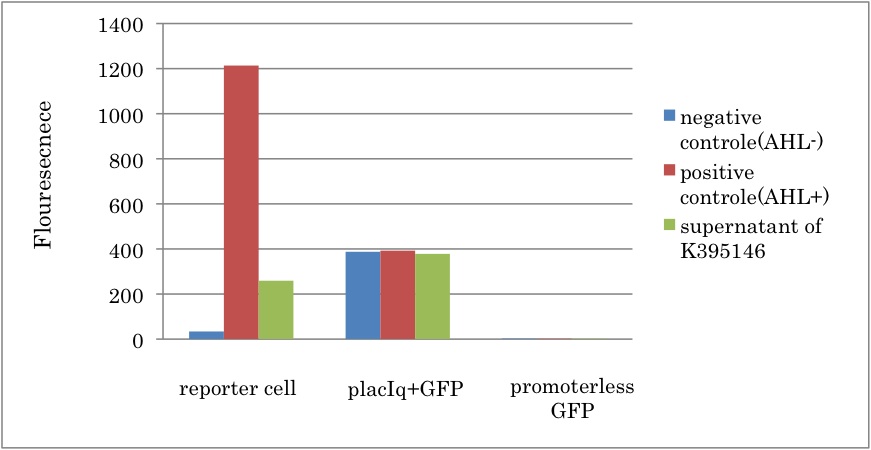
| + | We constructed a new LuxI generator ([http://partsregistry.org/Part:BBa_K395146 BBa_K395146]) and characterized its function. By combining I14032 (PlacI<sup>q</sup>) and K092400 (rbs-luxI-ter), we constructed K395146. Then we confirmed that the cell with K395146 produced the sufficient amount of 3OC6HSL to activate LuxR activation promoter (R0062) [[Team:Tokyo_Tech/Project/Artificial_Cooperation_System/luxI_assay|(See more…)]] | ||
| + | |||
| + | [[IMAGE:Tokyotech Fig.K395146-1.jpg|400px|left|thumb|Fig3-3-1 LuxI and LuxR activation promoter assay. This work done by Eriko Uchikoshi and Shohei Kitano]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | + | ==Mathematical modeling== | |
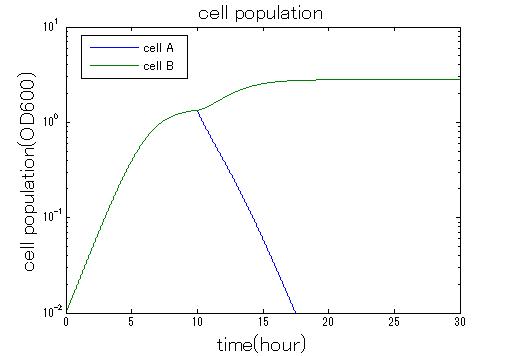
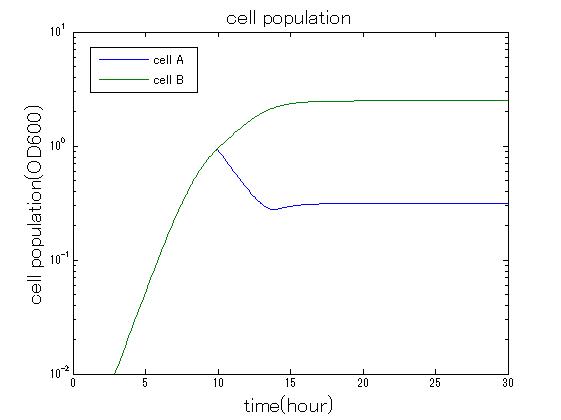
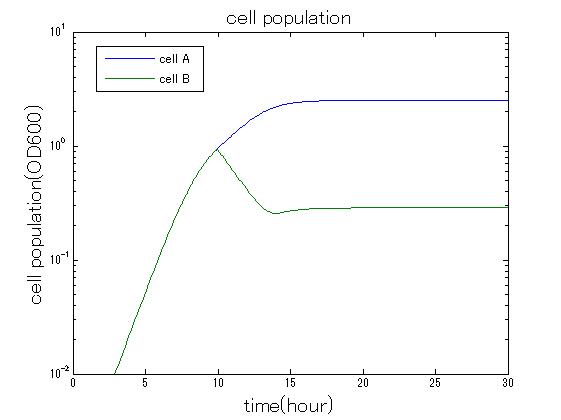
| + | In order to confirm the feasibility of the Artificial Cooperation System, we simulated the system under typical four experimental conditions. First, when the system worked and any antibiotics did not exist, cell A and cell B in the system showed no difference of growth (Fig.3-0-2). Second, when the system did not work and chloramphenicol existed, the number of cell A decreased while the number of cell B increased (Fig.3-0-3). Third, when the system worked and chloramphenicol existed, the number of cell A increased after temporal decline (Fig.3-0-4), which means it was rescued by Artificial Cooperation System. Fourth, in the contrast, when the system worked and kanamycin existed, the number of cell B increased after temporal decline (Fig.3-0-5). From the above results, the feasibility of the Artificial Cooperation System was confirmed.[[Team:Tokyo_Tech/Project/Artificial_Cooperation_System/modeling|(See more…)]] | ||
| + | [[IMAGE:tokyotech_no_anti_sim_result_cell.jpg|300px|left|thumb|Fig3-0-2 Simulation result in absence of antibiotics. This work done by Shohei Kitano]] | ||
| + | [[IMAGE:Tokyptech no system sim result.jpg|300px|left|thumb|Fig3-0-3 Simulation result without Artificial Cooperation System. This work done by Shohei Kitano]] | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | + | [[IMAGE:tokyotech_cm_sim_result_cell.jpg|300px|left|thumb|Fig3-0-4 Simulation result in the presence of chloramphenicol. This work done by Shohei Kitano]] | |
| + | [[IMAGE:tokyotech_kan_sim_result_cell.jpg|300px|left|thumb|Fog3-0-5 Simulation result in the presence of kanamycin. This work done by Shohei Kitano]] | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| Line 135: | Line 436: | ||
| + | <!--このしたから--> | ||
| - | + | =Conclusion= | |
| + | We succeeded in constructing two new BioBrick, LuxR repression promoter (K395008) and chloramphenicol resistance gene generator activated by LuxR and 3OC6HSL. We characterized existing LuxR repression promoter and LuxR activation promoter in the registry. From the experimental data and simulation result, we confirmed the feasibility of the Artificial Cooperation System, ''E. coli'' with humanity. | ||
| - | + | <!-- --> | |
Latest revision as of 03:59, 28 October 2010
3 Artificial Cooperation System
Contents |
Artificial Cooperation System
Abstract
We built Artificial Cooperation System to endow E.coli with humanity. In this system quorum sensing system and chloramphenicol resistance protein generator activated by LuxR are very important. First, we constructed and characterized several promoters regulated by LuxR and 3OC6HSL. We designed and constructed a new LuxR repression promoter. Also, we characterized the functions of a LuxR repression promoter and a LuxR activation promoter in the registry. Second, we constructed a new chloramphenicol resistance protein generator which is regulated by LuxR activation promoter. We confirmed that the cell with the chloramphenicol resistance protein generator can survive under high concentration of chloramphenicol when 3OC6HSL exists. Third, we confirmed that LuxI can produce sufficient 3OC6HSL to activate chloramphenicol resistance gene. Forth, we confirmed the feasibility of Artificial Cooperation System from simulation and experimental data.
Introduction
What’s Artificial Cooperation System?
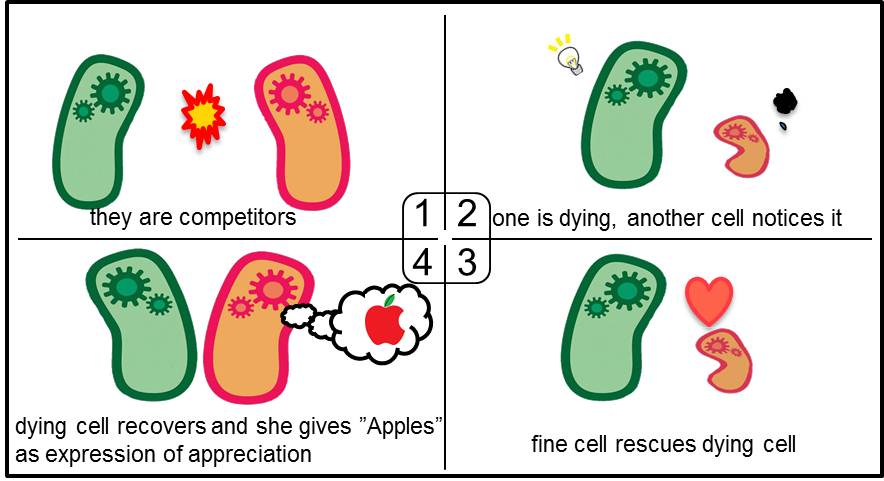
Artificial Cooperation System was built to engineer E. coli that behaves kindly and have sympathy. In our idea of E. coli with humanity, there are two types of cells. In normal conditions, they are competitors for survival. While in critical conditions, such that one is dying, another cell would notice the crisis of dying cell and help her. For realization of our idea, we constructed a system that named “Artificial Cooperation System”.
Genetic Circuit
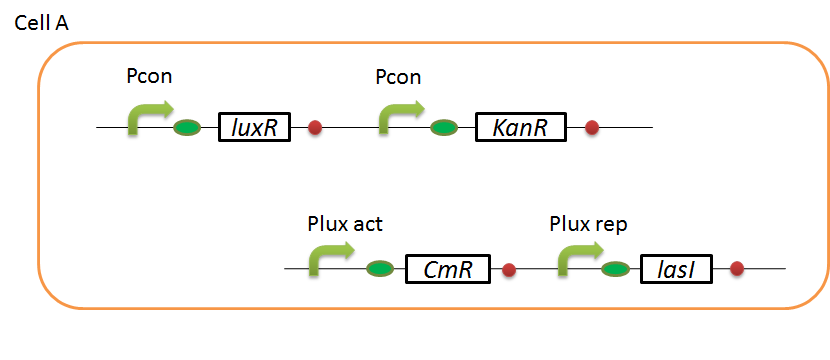
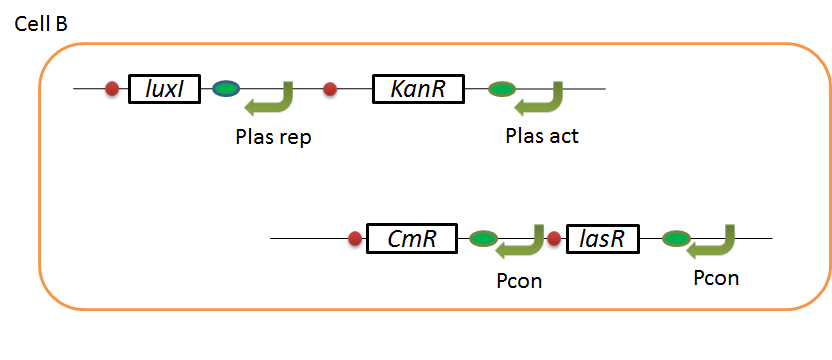
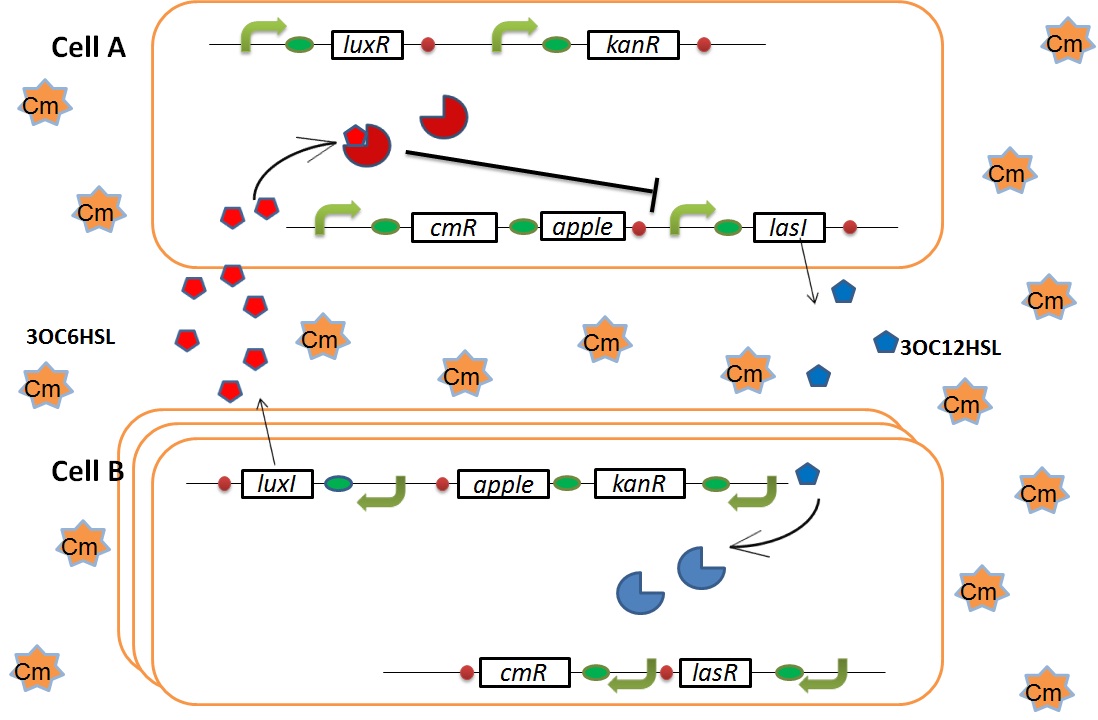
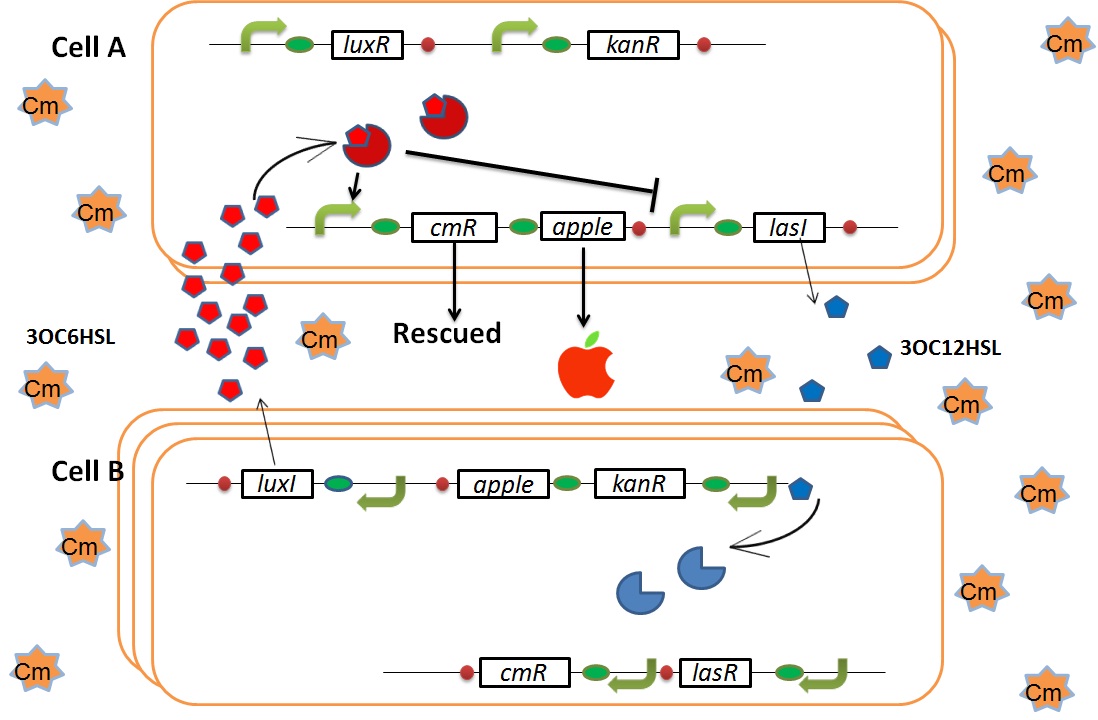
We designed the following circuit. In Artificial Cooperation System, there are two types of E. coli, Cell A and Cell B. Cell A has resistance to kanamycin and Cell B has resistance to chloramphenicol originally. In cell A (Fig 3-0-2), production of LasI is repressed by LuxR/3OC6HSL complex and chloramphenicol resistance gene is activated by LuxR/3OC6HSL complex. In cell B (Fig3-0-3), production of LuxI is repressed by LasR/3OC12HSL complex and kanamycin resistance gene is activated by LasR/3OC12HSL complex. This means that 3OC12HSL produced by cell A and 3OC6HSL produced by cell B indirectly repress each other’s production. On the other hand, 3OC12HSL activates resistance gene of cell B and 3OC6HSL activates resistance gene of cell A. Apple gene is composed of crtEBIYZW and MpAAT1. This expresses only when the cell is rescued and recovered.
Pcon: constitutive promoter.
Plux act: promoter activated by LuxR and 3OC6HSL. We call this promoter luxR activation promoter.
Plux rep: promoter repressed by LuxR and 3OC6HSL. We call this promoter luxR repression promoter.
Plas act: promoter activated by LasR and 3OC12HSL. We call this promoter lasR activation promoter.
Plas rep: promoter repressed by LasR and 3OC12HSL. We call this promoter lasR repression promoter.
lasI: enzyme repressed by LuxR and 3OC6HSL. We call this promoter LuxR repression promoter.
luxI: enzyme repressed by LasR and 3OC12HSL. We call this promoter LasR repression promoter.
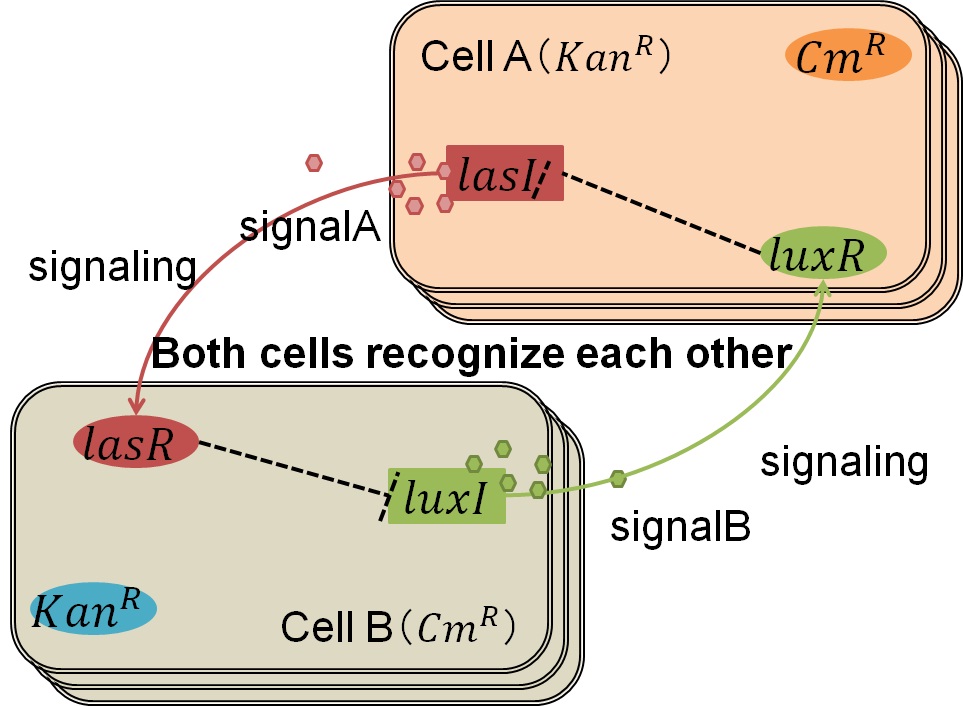
Normal
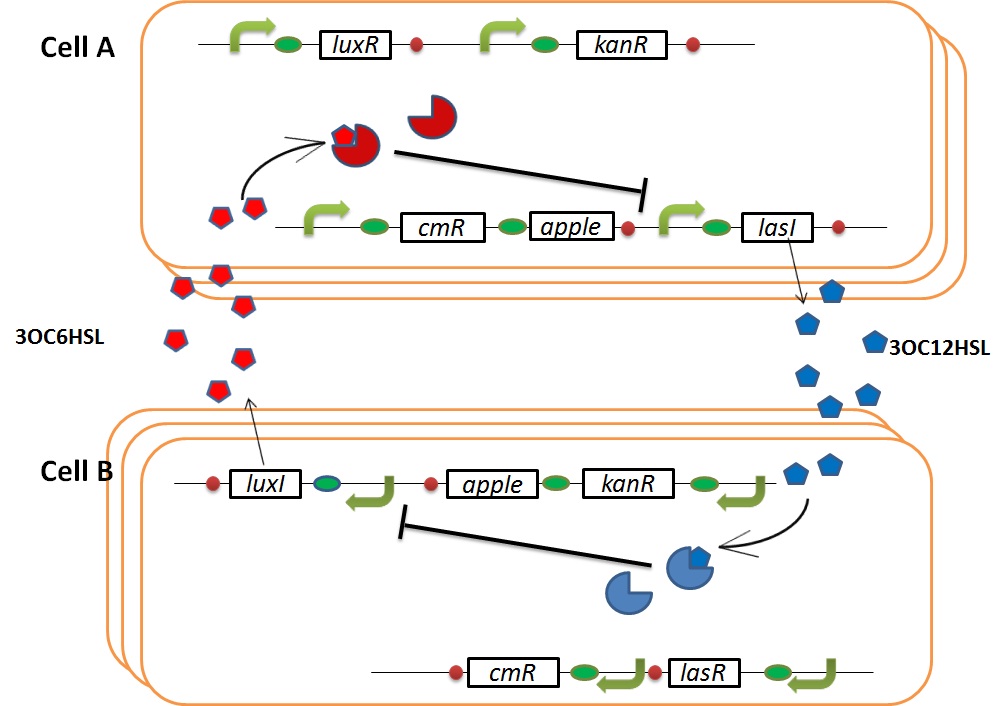
In normal situation, Cell A and Cell B are competitors and recognize each other by using quorum sensing. First, LasI protein in Cell A produces 3OC12HSL. Second, these signal molecules diffuse through cell membranes and bind to LasR protein in Cell B. Third, LasR/3OC12HSL complex represses the production of LuxI protein. Similarly in cell B, LuxI protein produces 3OC6HSL. These signal molecules diffuse through cell membranes and bind to LuxR protein in Cell A. LuxR/3OC6HSL complex represses the production of LasI protein. Although Cell A and Cell B have chloramphenicol and kanamycin resistance gene respectively, they don’t express in this situation. This is because the concentration of 3OC12HSL and 3OC6HSL is too low to activate the expressions of kanamycin resistance gene and chloramphenicol resistance gene, respectively.
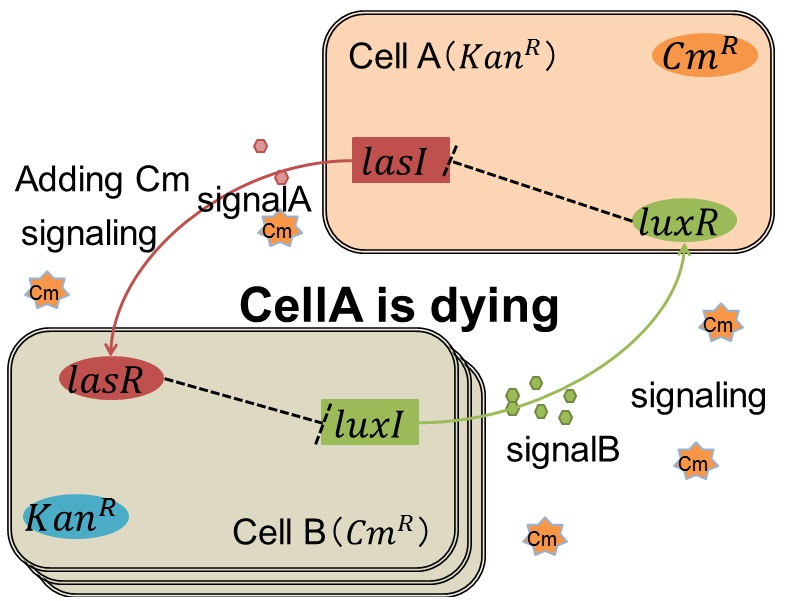
when Cell A is dying
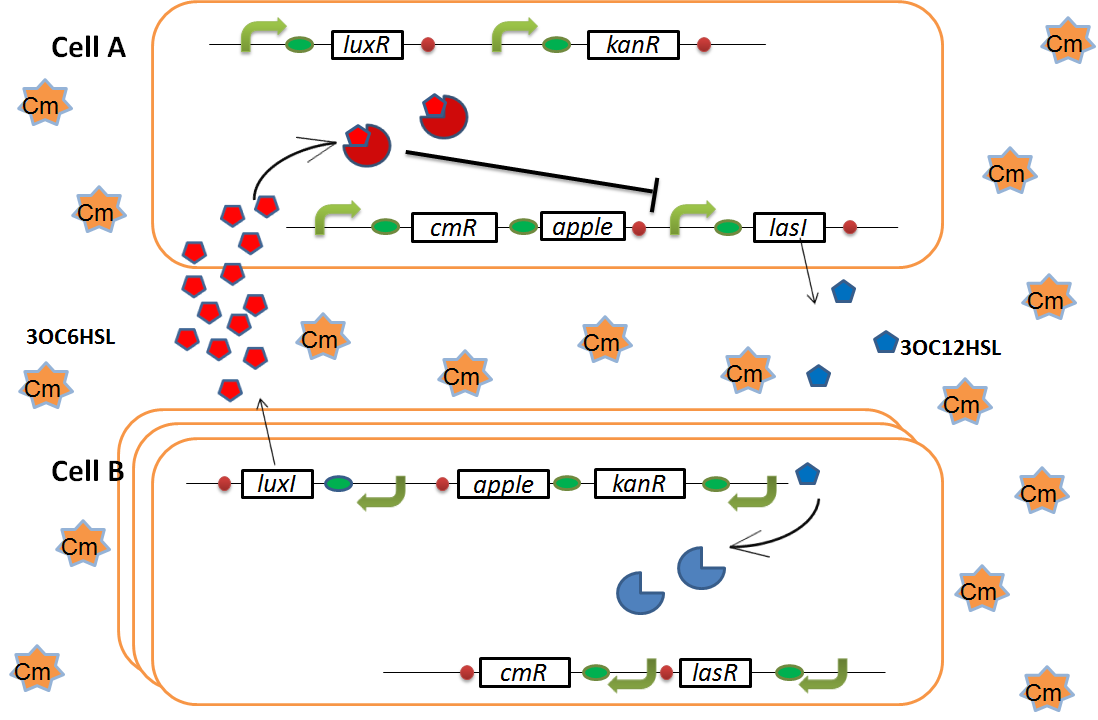
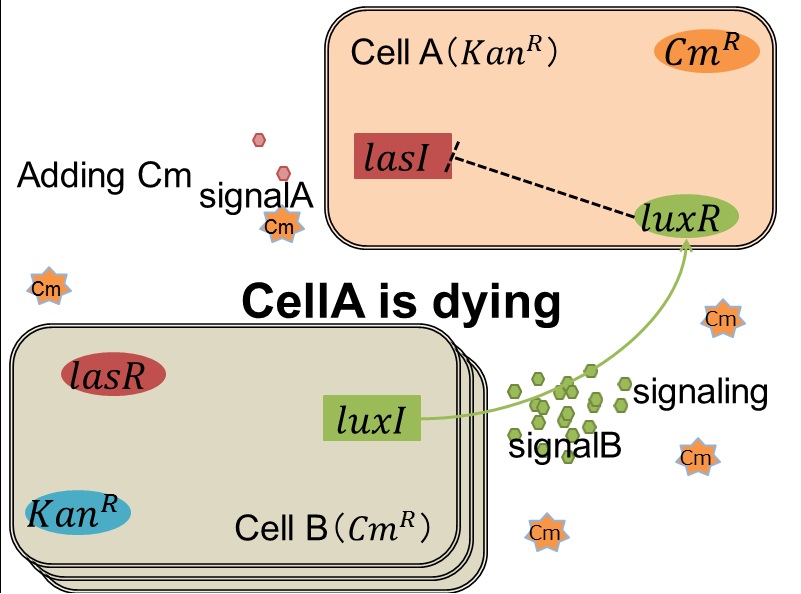
As we mentioned, originally Cell A has a resistance gene against kanamycin and does not have that against chloramphenicol. On the other hand, Cell B has resistance to chloramphenicol and doesn’t have resistance to kanamycin. Therefore, the addition of chloramphenicol only decreases the number of Cell A. It means that 3OC12HSL decreases. It also results in the decrease of LasR/3OC12HSL complex that repress the expression of LasI in Cell B. By this process, Cell B notices that Cell A is dying and tries to rescue it.
when Cell B tries to rescue Cell A
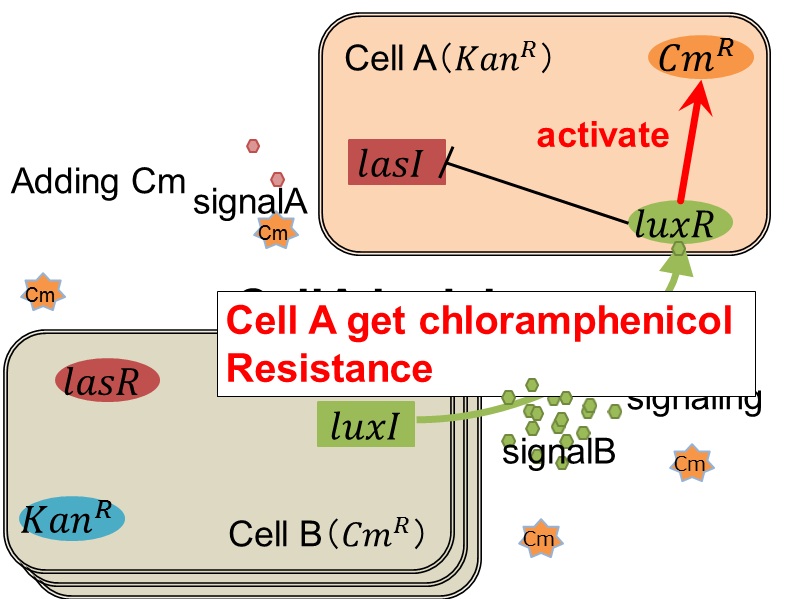
As we mentioned, the expression of luxI is repressed by LasR/3OC12HSL complex. Therefore, the decrease of 3OC12HSL/LasR complex causes the overexpression of luxI and the increase of 3OC6HSL. This results in the expression of chloramphenicol resistance gene in the Cell A. Consequently, the Cell A is recovered and begins to produce the apple reporters in return!
Result
In order to make the Artificial Cooperation System, we constructed and characterized several important parts and devices. First we characterized LuxR repression promoter R0061 and activation promoter R0062, both of which are existing parts. Second, we designed and characterized a new LuxR repression promoter [http://partsregistry.org/Part:BBa_K395008 K395008]. Third, we designed and characterized a NEW BioBrick device [http://partsregistry.org/Part:BBa_K395162 K395162], a chloramphenicol resistance generator, which is activated by a LuxR activation promoter. Forth, we confirmed that LuxI can produce sufficient AHL to help dying cell. Finally, we simulated our Artificial Cooperation System and confirmed the feasibility of our system.
LuxR repression promoter and activation promoter
We characterized LuxR repression promoters and activation promoters, which play important roles in the Artificial Cooperation System. The following figures are the result of experiments. Fig3-1-1 shows that under the regulations by R0062 (LuxR activation promoter), GFP expression was activated in the presence of 3OC6HSL. Fig3-1-3 shows that under the regulations by R0061 and K395008, GFP expressions were repressed in the presence of 3OC6HSL. Also, for realization of the Artificial Cooperative System work, the strength and the threshold of these promoters are so important. Thus we tried tuning promoters by changing sequence of consensus -35/-10 and lux box. (See more…)
Chloramphenicol resistance geneator activated by LuxR
We succeeded in construction and characterization of a NEW Biobrick device of a chloramphenicol resistance (CmR) generator ([http://partsregistry.org/Part:BBa_K395162 BBa_K395162]) which is activated by LuxR activation promoter. Fig 3-2-1 shows that in the presence of 3OC6HSL, the device is activated and as a result the cell was able to survive under high chloramphenicol concentration (750 ug/ml). (See more…)
LuxI can produce sufficient 3OC6HSL to activate LuxR activation promoter
We constructed a new LuxI generator ([http://partsregistry.org/Part:BBa_K395146 BBa_K395146]) and characterized its function. By combining I14032 (PlacIq) and K092400 (rbs-luxI-ter), we constructed K395146. Then we confirmed that the cell with K395146 produced the sufficient amount of 3OC6HSL to activate LuxR activation promoter (R0062) (See more…)
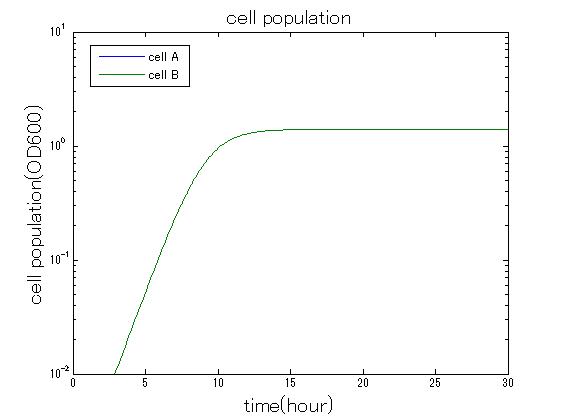
Mathematical modeling
In order to confirm the feasibility of the Artificial Cooperation System, we simulated the system under typical four experimental conditions. First, when the system worked and any antibiotics did not exist, cell A and cell B in the system showed no difference of growth (Fig.3-0-2). Second, when the system did not work and chloramphenicol existed, the number of cell A decreased while the number of cell B increased (Fig.3-0-3). Third, when the system worked and chloramphenicol existed, the number of cell A increased after temporal decline (Fig.3-0-4), which means it was rescued by Artificial Cooperation System. Fourth, in the contrast, when the system worked and kanamycin existed, the number of cell B increased after temporal decline (Fig.3-0-5). From the above results, the feasibility of the Artificial Cooperation System was confirmed.(See more…)
Conclusion
We succeeded in constructing two new BioBrick, LuxR repression promoter (K395008) and chloramphenicol resistance gene generator activated by LuxR and 3OC6HSL. We characterized existing LuxR repression promoter and LuxR activation promoter in the registry. From the experimental data and simulation result, we confirmed the feasibility of the Artificial Cooperation System, E. coli with humanity.
 "
"