Team:Paris Liliane Bettencourt/Project/Population counter/Results
From 2010.igem.org
| (28 intermediate revisions not shown) | |||
| Line 5: | Line 5: | ||
<p style="display:block;"> | <p style="display:block;"> | ||
<a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Population_counter"> | <a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Population_counter"> | ||
| - | <img src="https://static.igem.org/mediawiki/2010/ | + | <img src="https://static.igem.org/mediawiki/2010/9/93/Pop_counter_logo-01.jpg" width="151" height="125" title="Population Counter"> |
</a> | </a> | ||
| - | <font size=4>Population counter | + | <font size=4>Population counter</font> |
<a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/SIP"> | <a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/SIP"> | ||
| - | <img src="https://static.igem.org/mediawiki/2010/4/4c/SIP.png" width=" | + | <img src="https://static.igem.org/mediawiki/2010/4/4c/SIP.png" width="89" height="89" align=right title="SIP"> |
</a> | </a> | ||
<a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Memo-cell"> | <a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Memo-cell"> | ||
| - | <img src="https://static.igem.org/mediawiki/2010/ | + | <img src="https://static.igem.org/mediawiki/2010/a/aa/Memo_cell-01.jpg" width="108" height="89" align=right title="Memo-Cell"> |
</a> | </a> | ||
<br /> | <br /> | ||
| Line 29: | Line 29: | ||
<li><a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Population_counter/Results" target="_self">Results</a></li> | <li><a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Population_counter/Results" target="_self">Results</a></li> | ||
<li><a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Parts" target="_self">Parts</a></li> | <li><a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Parts" target="_self">Parts</a></li> | ||
| + | <li><a href="https://2010.igem.org/Team:Paris_Liliane_Bettencourt/Project/Population_counter/References" target="_self">References</a></li> | ||
</ul> | </ul> | ||
</div> | </div> | ||
<br /><br /> | <br /><br /> | ||
| + | </p> | ||
| - | </ | + | </html><p style="display:block;"> |
| - | < | + | <br> |
| - | < | + | ==Construction of our counters== |
| + | <html><p style="display:block;"> | ||
| + | <br>First of all, we have spent a significant part of the summer to do cloning using the standard biobrick assembly protocol. For this, we have used a lot of biobricks submitted to the parts registry by previous teams, thank you ! Using these parts we have created plenty of new composite parts that are highly modular (different kind of RBS and promoters) and could serve next iGEM teams. We have also created some new biobricks that are at the core of our project. We hope that other iGEM teams will work with integrons and might need these parts. We have constructed both the basic counter (with RFP and Tetracycline resistance gene) and the full counter + timer designs. | ||
| + | <br>Moreover, each of our constructs was verified by sequencing. For more information you can take a look at the parts that we sent to the registry. | ||
| + | <br></html> | ||
<br> | <br> | ||
| - | <br> | + | ==Proof that IntI1 integrase works !== |
| + | <html><p style="display:block;"> | ||
| + | <br> Then we wanted to see if the IntI1 integrase actually works. We have done a double transformation "RFP Counter" (on pSB1A2 which is a high copy plasmid) + "pBAD-IntI1" (on pSulib which is a low copy plasmid with kanamycin resistance) to obtain a working counter. The protocol of the counter testing is quite simple : we dilute an overnight culture 1/100, wait until OD 0.2 and add arabinose to begin the test (t=0h). At regular time intervals, we plated on plates containing ampicillin and glucose. We didn't have to put kanamycin into the plates because we don't want recombinations to happen on the plates. So we don't need the cells to keep their pBAD/Int plasmid. Moreover, to make sure that to recombinations happen after the cells have been plated, we have added glucose to the plates because glucose tightly represses the pBAD promoter, so that we can be sure that the integrase isn't expressed. | ||
| + | <br><br><a href="https://static.igem.org/mediawiki/2010/1/11/Ara-01.png" rel="zoombox"><img style="border:5px solid white" src="https://static.igem.org/mediawiki/2010/1/11/Ara-01.png" width="600px"></a> | ||
| - | |||
| - | <br> | + | <br> This experiment shows us that IntI1 is working! The double terminator flanked by attC sites has been excised and the constitutive promoter upstream permits the expression of RFP. However, we have 2 important notes : |
| - | + | </html> | |
| + | * At the beginning of the test, there are already some fluorescent colonies... We can explain this fact by the leaking of pBad that controls the expression of the integrase. Thereby the integrase is expressed at a low level and excises the terminator while the cells are growing in an overnight culture, even before we start our test. To overcome this problem we had to add glucose into the overnight culture media. Thus, glucose blocks the pBAD promoter and avoids recombination events to occur before our test begins. | ||
| + | *The expression of RFP is inhomogeneous because the counter is on a high copy plasmid. So some cells have 2 recombined plasmids while others have 20 recombined plasmids ! To obtain a more linear response we have integrated our counter in a low copy plasmid (pSB4A5). In this way, there is more chance that only one event will happen in each cell. | ||
<br><br> | <br><br> | ||
| - | + | ||
| - | < | + | ==Proof that the Tetracyline Counter works and first evaluation of the recombination rate== |
| - | <br> | + | <html><p style="display:block;"> |
| - | + | <br>Following the same protocol, we tested the Tetracycline Counter. Only one copy of TetA(C) expressed is sufficient to induce tetracycline resistance. Therefore the number of resistant colonies is a good approximation of the recombination rate.</html> | |
| - | + | ||
| - | + | ||
| - | + | ||
<br>[[Image:Tetracyclin resistance.jpg|centre|600px]] | <br>[[Image:Tetracyclin resistance.jpg|centre|600px]] | ||
| + | |||
<br><br> | <br><br> | ||
| - | + | ||
| + | ==Confirmation of the specific excision of the terminator by sequencing== | ||
| + | <html><p style="display:block;"> | ||
| + | <br> We had to be sure that the clones that have become resistant to tetracycline have changed their phenotype because the terminator got excised and not simply due to leakage. That's why we miniprepped our resistant clones and sent these plasmids to sequencing. Here you can see the nucleotide alignments which prove that the IntI1 integrase has specifically excised the terminator flanked by two attC sites. | ||
<br><br> | <br><br> | ||
| - | + | ||
| + | <a href="https://static.igem.org/mediawiki/2010/5/59/Unexcised_terminator-01.png" rel="zoombox"><img style="border:2px solid black" src="https://static.igem.org/mediawiki/2010/5/59/Unexcised_terminator-01.png" width="900px"> | ||
| + | |||
| + | <a href="https://static.igem.org/mediawiki/2010/3/3f/Excised_terminator-01.png" rel="zoombox"><img style="border:2px solid black" src="https://static.igem.org/mediawiki/2010/3/3f/Excised_terminator-01.png" width="900px"> | ||
| + | <br><br></html> | ||
| + | <br><br> | ||
| + | |||
| + | ==Characterization of the recombination rate== | ||
| + | <html> | ||
| + | <p style="display:block;"> | ||
| + | <br>At this point, it is important to determine the recombination rate of IntI1. To do this we have done our double transformations again on plates containing 1% glucose to inhibit the expression of IntI1 before the beginning of the test. We did our overnight cultures with glucose for the same reason, then we diluted, waited until OD 0.2 and added arabinose 0,04%. The counters are on a high copy number plasmid, so as we have explained before it is difficult to estimate the number of recombinants per cell. But we found a trick to calculate the recombination rate ! | ||
| + | <br>In every cell there is a lot of plasmids, some of them are recombined, others not. To have a good idea of this proportion, we did minipreps at regular time intervals in parallel with direct plating of the cultures. Then we transformed the minipreps that contain the whole population of plasmids. Statistically, only one plasmid can be integrated per competent cell. After the plasmid is integrated into the cell, it is multiplied to reach its copy-number. This can be regarded as an <b>amplification of the recombination result</b>. This way, some cells are red (because they were transformed with an excised plasmid and then copied it) while the others are not fluorescent (because they were transformed with the wild-type plasmid). This protocol is more complicated but it permits to significantly increase the sensitivity of the test. | ||
| + | <br>However, the experiment with the Tetracycline Counter failed. The direct plating gives positive results but the transformation of the minipreps gives no colonies on tetracycline plates. After some researches we noted that the <b>TetA(C) gene is toxic for the cells if over-expressed</b> <a href="http://partsregistry.org/wiki/index.php/Part:BBa_J31007">(see description of the biobrick)</a>. The principle of this experiment is to obtain only recombined plasmids in the cells, so we assume that it is lethal for the cells when expressed on a high copy number plasmid. | ||
| + | </html> | ||
| + | <br>[[Image:Recombination rate.jpg|centre|600px]] | ||
| + | |||
| + | <br>This graph shows the evolution of the recombination rate deduced from the number of recombined plasmids. In order to assure efficient counting, the recombination rate has to be low (under 5%). From these results we can see that arabinose induction triggers too much recombinations. Actually the absence of arabinose is a perfect condition to test the counter (leaking of pBad). | ||
| + | |||
| + | <br><br> | ||
| + | ==Characterisation of the integrase toxicity== | ||
| + | <html> | ||
| + | <p style="display:block;"><br> When doing the experiments, we noted that the number of colonies on our plates abnormally decreased with time. We deduced that the expression of the integrase is toxic for the cells, as you can see below.</html> | ||
<br>[[Image:IntI1 toxicity.jpg|centre|600px]] | <br>[[Image:IntI1 toxicity.jpg|centre|600px]] | ||
| + | |||
| + | |||
<br><br> | <br><br> | ||
| - | + | ==Test of the microfluidic device== | |
| + | <html> | ||
| + | <p style="display:block;"> | ||
| + | We have replicated the original pattern from the mask and made PDMS chips. We have managed to test the microfluidic chip. It works : the cells can get trapped inside the chambers, and then grow inside the chambers filling them up. The fact that the cells grow to a high density is important because we need the culture to be dense enough for the quorum sensing to work. | ||
| + | <br>Here are the videos of our microfluidic device we have taken : | ||
| + | <br><br><center><object width="480" height="385"><param name="movie" value="http://www.youtube.com/v/TK-hAuKtmTw?fs=1&hl=en_US"></param><param name="allowFullScreen" value="true"></param><param name="allowscriptaccess" value="always"></param><embed src="http://www.youtube.com/v/TK-hAuKtmTw?fs=1&hl=en_US" type="application/x-shockwave-flash" allowscriptaccess="always" allowfullscreen="true" width="480" height="385"></embed></object> | ||
| + | <br><br><object width="480" height="385"><param name="movie" value="http://www.youtube.com/v/2TvztGO4two?fs=1&hl=en_US"></param><param name="allowFullScreen" value="true"></param><param name="allowscriptaccess" value="always"></param><embed src="http://www.youtube.com/v/2TvztGO4two?fs=1&hl=en_US" type="application/x-shockwave-flash" allowscriptaccess="always" allowfullscreen="true" width="480" height="385"></embed></object> | ||
| + | </center> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | </html> | ||
Latest revision as of 03:55, 28 October 2010
Construction of our counters
First of all, we have spent a significant part of the summer to do cloning using the standard biobrick assembly protocol. For this, we have used a lot of biobricks submitted to the parts registry by previous teams, thank you ! Using these parts we have created plenty of new composite parts that are highly modular (different kind of RBS and promoters) and could serve next iGEM teams. We have also created some new biobricks that are at the core of our project. We hope that other iGEM teams will work with integrons and might need these parts. We have constructed both the basic counter (with RFP and Tetracycline resistance gene) and the full counter + timer designs.
Moreover, each of our constructs was verified by sequencing. For more information you can take a look at the parts that we sent to the registry.
Proof that IntI1 integrase works !
Then we wanted to see if the IntI1 integrase actually works. We have done a double transformation "RFP Counter" (on pSB1A2 which is a high copy plasmid) + "pBAD-IntI1" (on pSulib which is a low copy plasmid with kanamycin resistance) to obtain a working counter. The protocol of the counter testing is quite simple : we dilute an overnight culture 1/100, wait until OD 0.2 and add arabinose to begin the test (t=0h). At regular time intervals, we plated on plates containing ampicillin and glucose. We didn't have to put kanamycin into the plates because we don't want recombinations to happen on the plates. So we don't need the cells to keep their pBAD/Int plasmid. Moreover, to make sure that to recombinations happen after the cells have been plated, we have added glucose to the plates because glucose tightly represses the pBAD promoter, so that we can be sure that the integrase isn't expressed.

This experiment shows us that IntI1 is working! The double terminator flanked by attC sites has been excised and the constitutive promoter upstream permits the expression of RFP. However, we have 2 important notes :
- At the beginning of the test, there are already some fluorescent colonies... We can explain this fact by the leaking of pBad that controls the expression of the integrase. Thereby the integrase is expressed at a low level and excises the terminator while the cells are growing in an overnight culture, even before we start our test. To overcome this problem we had to add glucose into the overnight culture media. Thus, glucose blocks the pBAD promoter and avoids recombination events to occur before our test begins.
- The expression of RFP is inhomogeneous because the counter is on a high copy plasmid. So some cells have 2 recombined plasmids while others have 20 recombined plasmids ! To obtain a more linear response we have integrated our counter in a low copy plasmid (pSB4A5). In this way, there is more chance that only one event will happen in each cell.
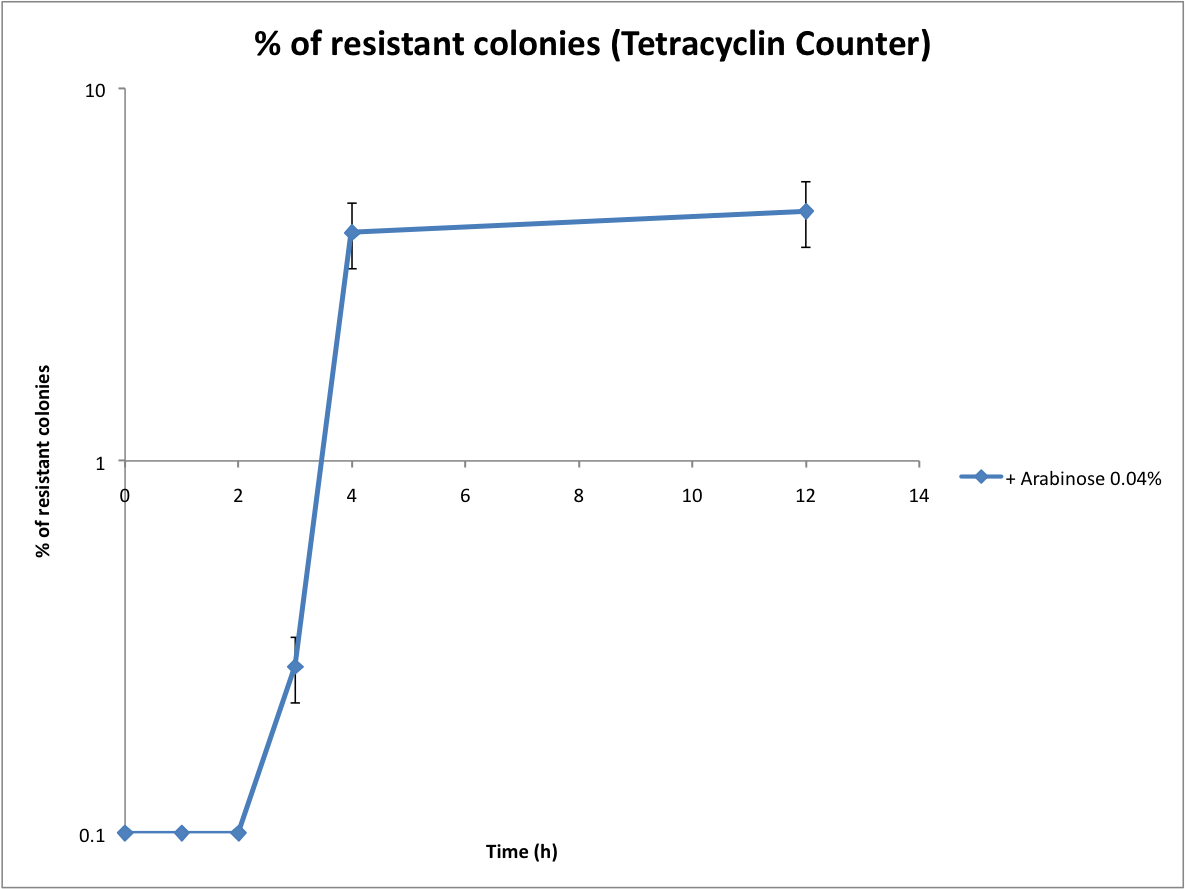
Proof that the Tetracyline Counter works and first evaluation of the recombination rate
Following the same protocol, we tested the Tetracycline Counter. Only one copy of TetA(C) expressed is sufficient to induce tetracycline resistance. Therefore the number of resistant colonies is a good approximation of the recombination rate.
Confirmation of the specific excision of the terminator by sequencing
We had to be sure that the clones that have become resistant to tetracycline have changed their phenotype because the terminator got excised and not simply due to leakage. That's why we miniprepped our resistant clones and sent these plasmids to sequencing. Here you can see the nucleotide alignments which prove that the IntI1 integrase has specifically excised the terminator flanked by two attC sites.


Characterization of the recombination rate
At this point, it is important to determine the recombination rate of IntI1. To do this we have done our double transformations again on plates containing 1% glucose to inhibit the expression of IntI1 before the beginning of the test. We did our overnight cultures with glucose for the same reason, then we diluted, waited until OD 0.2 and added arabinose 0,04%. The counters are on a high copy number plasmid, so as we have explained before it is difficult to estimate the number of recombinants per cell. But we found a trick to calculate the recombination rate !
In every cell there is a lot of plasmids, some of them are recombined, others not. To have a good idea of this proportion, we did minipreps at regular time intervals in parallel with direct plating of the cultures. Then we transformed the minipreps that contain the whole population of plasmids. Statistically, only one plasmid can be integrated per competent cell. After the plasmid is integrated into the cell, it is multiplied to reach its copy-number. This can be regarded as an amplification of the recombination result. This way, some cells are red (because they were transformed with an excised plasmid and then copied it) while the others are not fluorescent (because they were transformed with the wild-type plasmid). This protocol is more complicated but it permits to significantly increase the sensitivity of the test.
However, the experiment with the Tetracycline Counter failed. The direct plating gives positive results but the transformation of the minipreps gives no colonies on tetracycline plates. After some researches we noted that the TetA(C) gene is toxic for the cells if over-expressed (see description of the biobrick). The principle of this experiment is to obtain only recombined plasmids in the cells, so we assume that it is lethal for the cells when expressed on a high copy number plasmid.
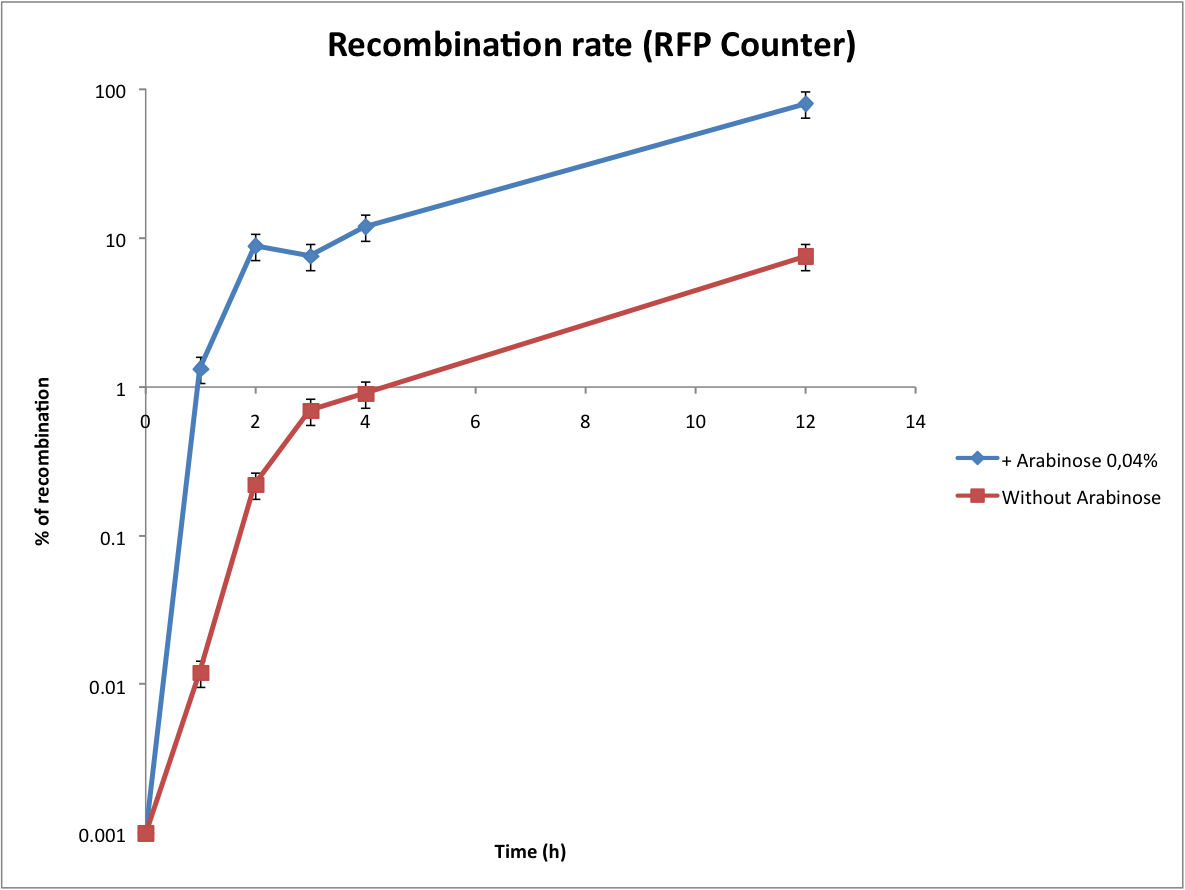
This graph shows the evolution of the recombination rate deduced from the number of recombined plasmids. In order to assure efficient counting, the recombination rate has to be low (under 5%). From these results we can see that arabinose induction triggers too much recombinations. Actually the absence of arabinose is a perfect condition to test the counter (leaking of pBad).
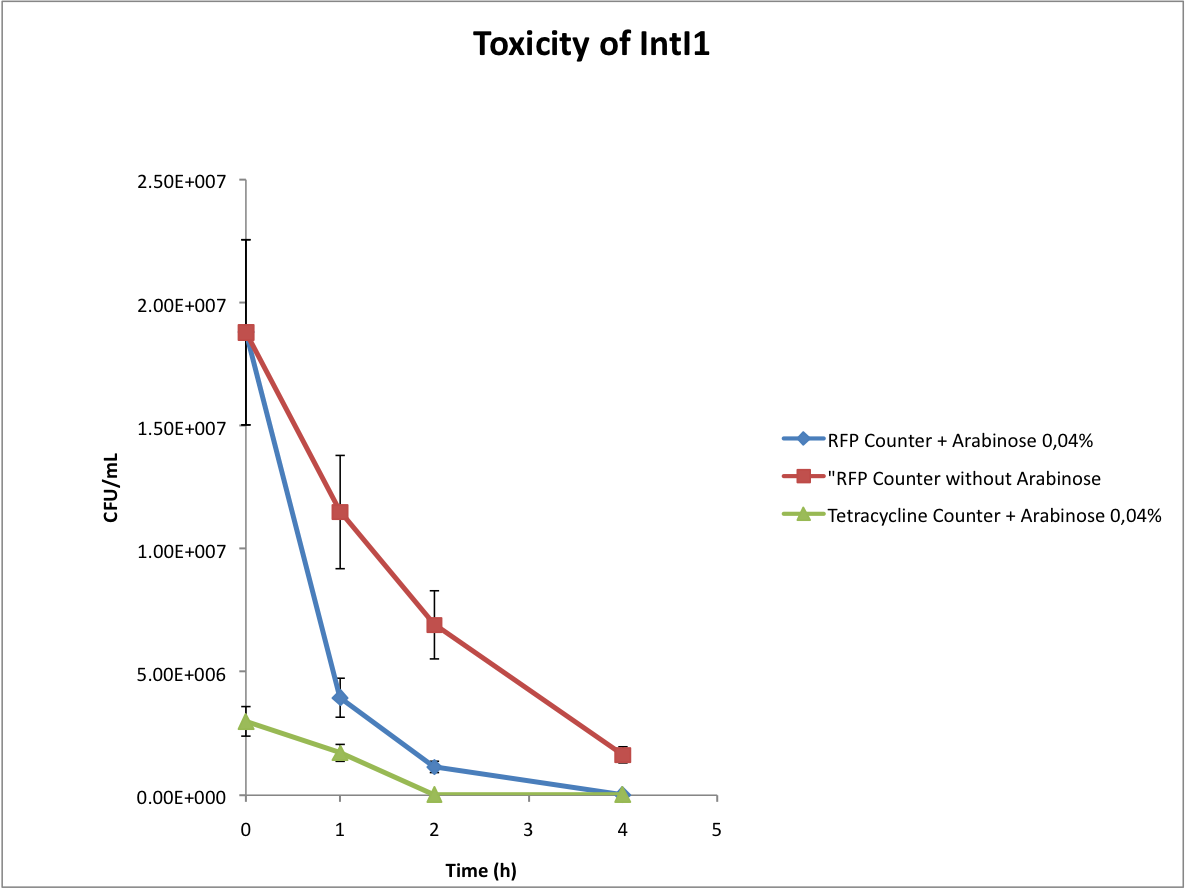
Characterisation of the integrase toxicity
When doing the experiments, we noted that the number of colonies on our plates abnormally decreased with time. We deduced that the expression of the integrase is toxic for the cells, as you can see below.
Test of the microfluidic device
We have replicated the original pattern from the mask and made PDMS chips. We have managed to test the microfluidic chip. It works : the cells can get trapped inside the chambers, and then grow inside the chambers filling them up. The fact that the cells grow to a high density is important because we need the culture to be dense enough for the quorum sensing to work.
Here are the videos of our microfluidic device we have taken :
 "
"