Team:Imperial College London/Modelling/Protein Display/Results and Conclusion
From 2010.igem.org
(Difference between revisions)
m |
(updating text under the figure) |
||
| Line 10: | Line 10: | ||
|[[Image:IC_Protein_display.png|600px]] | |[[Image:IC_Protein_display.png|600px]] | ||
|- | |- | ||
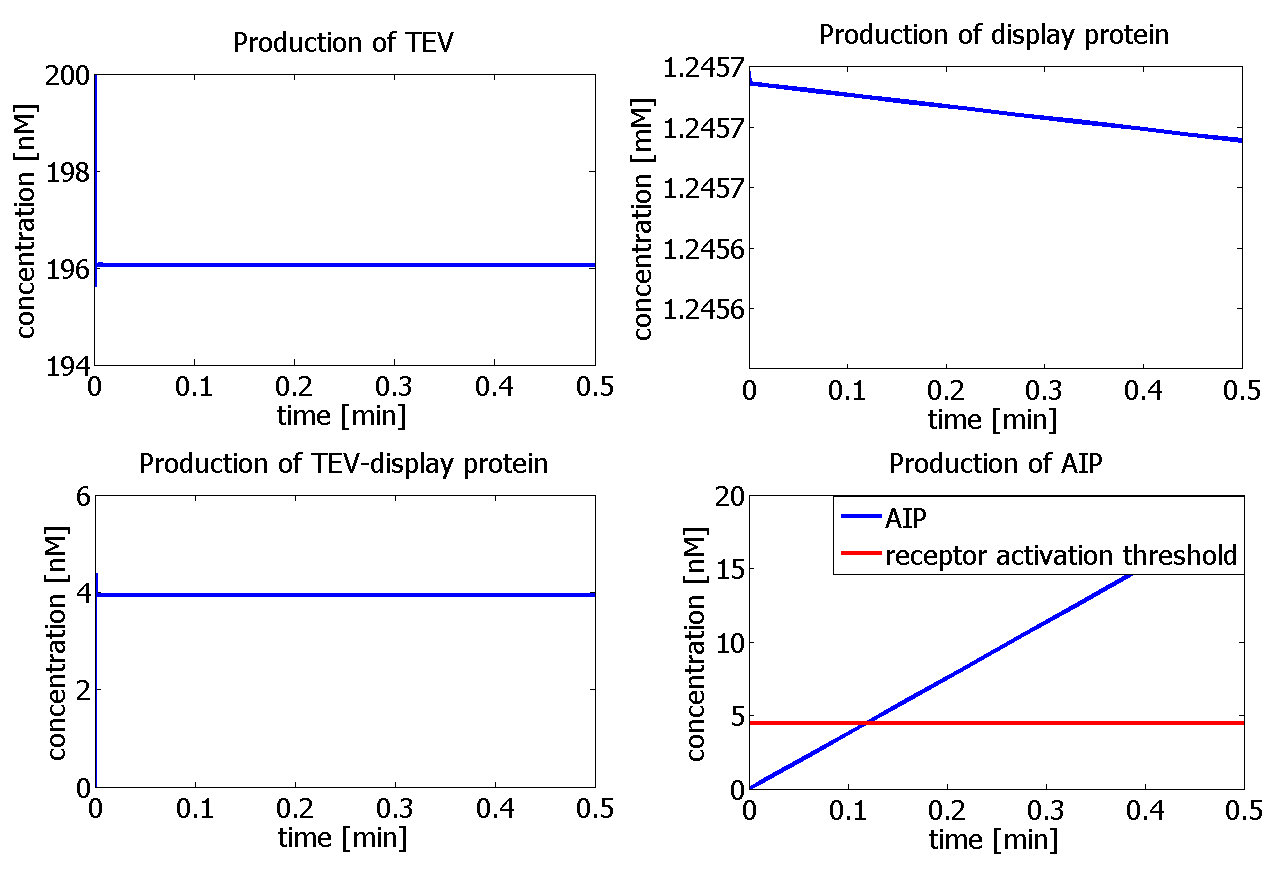
| - | |<html>Graphs showing the simulation using [TEV]<sub>0</sub> = | + | |<html>Graphs showing the simulation using [TEV]<sub>0</sub> = 400 nM. The graph on the right hand-side<br/> bottom shows that the AIP threshold (red line) is reached after 11 s!</html> |
|} | |} | ||
</div> | </div> | ||
Revision as of 14:09, 26 October 2010
| Modelling | Overview | Detection Model | Signaling Model | Fast Response Model | Interactions |
| A major part of the project consisted of modelling each module. This enabled us to decide which ideas we should implement. Look at the Fast Response page for a great example of how modelling has made a major impact on our design! | |
| Objectives | Description | Results | Constants | MATLAB Code |
| Results and Conclusion | ||||
Risk of False positives It was pointed out that we should assess the risk of false positive activation of the receptor. We are particularly concerned about the display protein not binding to the cell wall, but instead diffusing into the extra-cellular environment. In order to be able to assess the risk of false positives, we need to do further research into the affinity of AIP with attached linker and transmembrane proteins for the receptor as compared to the affinity of the AIP itself for the receptor. This paper [1] might have some information on affinity comparison. We need to know how proteins are being transported from intracellular to transmembrane space. Understanding this concept could give us an idea of what could go wrong. References
|
 "
"