Team:Imperial College London/Modelling/Protein Display/Detailed Description
From 2010.igem.org
(Menu fix) |
m |
||
| Line 7: | Line 7: | ||
|- | |- | ||
|<html> | |<html> | ||
| - | This model consists of | + | This model consists of seven parts that had to be developed: |
<ol> | <ol> | ||
<li>Identification of all active and relevant elements of the isolated part of the system.</li> | <li>Identification of all active and relevant elements of the isolated part of the system.</li> | ||
<li>Identification of interactions between the identified elements.</li> | <li>Identification of interactions between the identified elements.</li> | ||
| - | <li> | + | <li>Determining the threshold concentration of Auto Inducing Peptide (AIP) needed for activation of the receptor.</li> |
| - | <li>Determining the volume of cell wall.</li> | + | <li>Determining the volume of the cell wall.</li> |
<li>Defining a control volume around the bacterial cell (after cleavage, the surface protein will float around in the extracellular environment).</li> | <li>Defining a control volume around the bacterial cell (after cleavage, the surface protein will float around in the extracellular environment).</li> | ||
| - | <li>Determining the importance of localised concentrations in a | + | <li>Determining the importance of localised concentrations in a control volume.</li> |
<li>Determining the surface protein production to estimate the maximum abundance on the bacterial surface.</li> | <li>Determining the surface protein production to estimate the maximum abundance on the bacterial surface.</li> | ||
</ol> | </ol> | ||
| Line 20: | Line 20: | ||
<h2>1. Elements of the system</h2> | <h2>1. Elements of the system</h2> | ||
<ol> | <ol> | ||
| - | <li>The surface protein consists of a cell wall binding domain, linker | + | <li>The surface protein consists of a cell wall binding domain, a linker and AIP (Auto Inducing Peptide). It is expressed by constitutive gene expression. It was assumed that the bacteria would be fully grown before carrying out the detection so the cell wall would be covered with as many surface proteins as the cell can maintain.</li> |
| - | <li>Schistosoma elastase (enzyme released by the parasite) cleaves the AIP from the cell wall binding domain at the linker site. In the laboratory we used TEV protease as we could not get hold of Schistosoma elastase, so the model was adjusted appropriately (TEV enzyme kinetic parameters were used).</li> | + | <li>Schistosoma elastase (enzyme released by the parasite) cleaves the AIP from the cell wall binding domain at the linker site. In the laboratory, we used TEV protease as we could not get hold of Schistosoma elastase, so the model was adjusted appropriately (TEV enzyme kinetic parameters were used).</li> |
<li>The ComD receptor is activated by a sufficiently high AIP concentration. Once activated, ComD signals into the cell to activate the colour response.</li> | <li>The ComD receptor is activated by a sufficiently high AIP concentration. Once activated, ComD signals into the cell to activate the colour response.</li> | ||
</ol> | </ol> | ||
| Line 34: | Line 34: | ||
<li>Product (P) = Peptide</li> | <li>Product (P) = Peptide</li> | ||
</ul><br /> | </ul><br /> | ||
| - | This enzymatic reaction can be rewritten as ordinary differential equations (ODEs), which is of similar form | + | This enzymatic reaction can be rewritten as a set of ordinary differential equations (ODEs), which is of similar form to the 1-step amplification model. |
<CENTER><img src="https://static.igem.org/mediawiki/2010/9/9c/Enz_react10.png"/></CENTER> | <CENTER><img src="https://static.igem.org/mediawiki/2010/9/9c/Enz_react10.png"/></CENTER> | ||
| - | This model is quite peculiar as we realised that its behaviour is not only dependent on time, but space as well. Namely, the cleavage reaction happens on the cell wall of bacteria. However, AIP that has already been cleaved is allowed to diffuse in any direction. Since we were not sure how to model this scenario, we decided to determine the importance of localised concentrations in a | + | This model is quite peculiar as we realised that its behaviour is not only dependent on time, but space as well. Namely, the cleavage reaction happens on the cell wall of the bacteria. However, AIP that has already been cleaved is allowed to diffuse in any direction. Since we were not sure how to model this scenario, we decided to determine the importance of localised concentrations in a control volume. |
<h2>3. Threshold concentration of AIP</h2> | <h2>3. Threshold concentration of AIP</h2> | ||
The optimal peptide concentration required to activate ComD is 10 ng/ml <a href="http://ukpmc.ac.uk/backend/ptpmcrender.cgi?accid=PMC40587&blobtype=pdf">[1]</a>. This is the threshold value for ComD activation. However, the minimum concentration of peptide to give a detectable activation is 0.5ng/ml. | The optimal peptide concentration required to activate ComD is 10 ng/ml <a href="http://ukpmc.ac.uk/backend/ptpmcrender.cgi?accid=PMC40587&blobtype=pdf">[1]</a>. This is the threshold value for ComD activation. However, the minimum concentration of peptide to give a detectable activation is 0.5ng/ml. | ||
<br /> | <br /> | ||
| - | The threshold for the minimal activation of the receptor is c<sub>th</sub>=4.4658×10<sup>-9</sup> mol/L. Click on the button below to uncover the | + | The threshold for the minimal activation of the receptor is c<sub>th</sub>=4.4658×10<sup>-9</sup> mol/L. Click on the button below to uncover the conversion from ng/ml to mol/L. |
<div id="wrapper"> | <div id="wrapper"> | ||
| Line 52: | Line 52: | ||
<li>The number of molecules in one ml is 10ng/3.7184×10<sup>-21</sup>g = 2.6893×10<sup>12</sup>. In a litre, the number of molecules is 2.6893×10<sup>15</sup>molecules/L.</li> | <li>The number of molecules in one ml is 10ng/3.7184×10<sup>-21</sup>g = 2.6893×10<sup>12</sup>. In a litre, the number of molecules is 2.6893×10<sup>15</sup>molecules/L.</li> | ||
<li>Dividing this value by Avogadro's constant gives the threshold concentration of c<sub>th</sub>=4.4658×10<sup>-9</sup> mol/L.</li> | <li>Dividing this value by Avogadro's constant gives the threshold concentration of c<sub>th</sub>=4.4658×10<sup>-9</sup> mol/L.</li> | ||
| - | <li>The threshold for minimal activation of receptor is 2.2329×10<sup>-10</sup> mol/L.</li> | + | <li>The threshold for minimal activation of a receptor is 2.2329×10<sup>-10</sup> mol/L.</li> |
</ul> | </ul> | ||
</div> | </div> | ||
| Line 59: | Line 59: | ||
<h2>4. Cell Wall Volume</h2> | <h2>4. Cell Wall Volume</h2> | ||
| - | + | It was necessary to calculate the volume of the cell wall as we needed it for the calculation of concentrations in the enzymatic reaction.<br/> | |
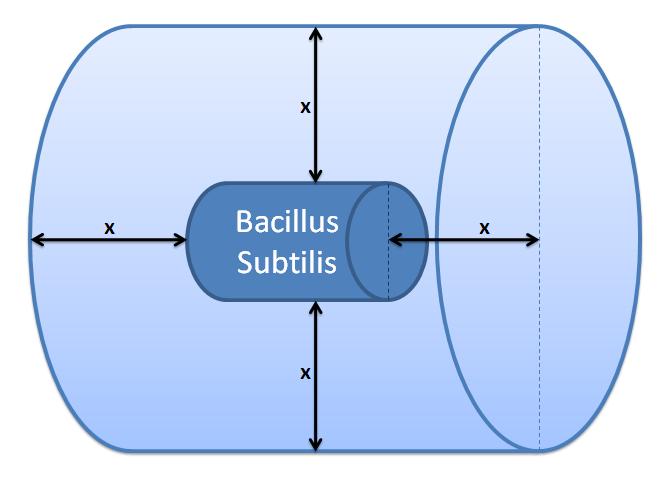
| - | Volume of B. subtilis is 2.79μm<sup>3</sup> and the thickness of cell wall is 35nm <a href="http://jb.asm.org/cgi/reprint/176/5/1413?ijkey=27dafbac7e23dee50390d3fe67d9d1bab0c6f48c">[5]</a>. In order to approximate the cell wall volume assume that B. subtilis is a sphere - not a rod. Calculate the outer radius from the total volume: 0.874μm. Now subtract the thickness of cell wall from outer radius to determine inner radius of the sphere: 0.839μm. The volume of cell wall | + | Volume of ''B. subtilis'' is 2.79μm<sup>3</sup> and the thickness of the cell wall is 35nm <a href="http://jb.asm.org/cgi/reprint/176/5/1413?ijkey=27dafbac7e23dee50390d3fe67d9d1bab0c6f48c">[5]</a>. In order to approximate the cell wall volume, assume that ''B. subtilis'' is a sphere - not a rod. Calculate the outer radius from the total volume: 0.874μm. Now subtract the thickness of the cell wall from the outer radius to determine the inner radius of the sphere: 0.839μm. The volume of the cell wall is equal to the difference between outer volume and the inner volume (calculated from the inner radius): <b>cell wall volume=0.32×10<sup>-15</sup>m<sup>3</sup></b> |
<h2>5. Control volume selection</h2> | <h2>5. Control volume selection</h2> | ||
| - | Note that product of the enzymatic reaction, AIP, is allowed to diffuse outside the cell. Hence, it is important to take into account the cell boundaries. It is worth considering whether diffusion or fluid movements will play a significant role. | + | Note that the product of the enzymatic reaction, AIP, is allowed to diffuse outside the cell. Hence, it is important to take into account the cell boundaries. It is worth considering whether diffusion or fluid movements will play a significant role. |
<br/> | <br/> | ||
<p>Initially, we defined a control volume assuming that bacteria would grow in close colonies on the plate. We realized that our initial choice of control volume was not accurate, since our bacteria are meant to be used in suspension so we had to reconsider this issue. | <p>Initially, we defined a control volume assuming that bacteria would grow in close colonies on the plate. We realized that our initial choice of control volume was not accurate, since our bacteria are meant to be used in suspension so we had to reconsider this issue. | ||
Revision as of 13:41, 27 October 2010
| Modelling | Overview | Detection Model | Signaling Model | Fast Response Model | Interactions |
| A major part of the project consisted of modelling each module. This enabled us to decide which ideas we should implement. Look at the Fast Response page for a great example of how modelling has made a major impact on our design! | |
| Objectives | Description | Results | Constants | MATLAB Code |
| Detailed Description | ||
This model consists of seven parts that had to be developed:
1. Elements of the system
2. Interactions between elementsApart from the proteins being expressed from genes, there was only one more chemical reaction identified in this part of the system. This is the cleavage of proteins, which is an enzymatic reaction:
 This enzymatic reaction can be rewritten as a set of ordinary differential equations (ODEs), which is of similar form to the 1-step amplification model.  3. Threshold concentration of AIPThe optimal peptide concentration required to activate ComD is 10 ng/ml [1]. This is the threshold value for ComD activation. However, the minimum concentration of peptide to give a detectable activation is 0.5ng/ml.The threshold for the minimal activation of the receptor is cth=4.4658×10-9 mol/L. Click on the button below to uncover the conversion from ng/ml to mol/L. Converting 10 ng/ml to 4.4658×10-9 mol/L
4. Cell Wall VolumeIt was necessary to calculate the volume of the cell wall as we needed it for the calculation of concentrations in the enzymatic reaction.Volume of ''B. subtilis'' is 2.79μm3 and the thickness of the cell wall is 35nm [5]. In order to approximate the cell wall volume, assume that ''B. subtilis'' is a sphere - not a rod. Calculate the outer radius from the total volume: 0.874μm. Now subtract the thickness of the cell wall from the outer radius to determine the inner radius of the sphere: 0.839μm. The volume of the cell wall is equal to the difference between outer volume and the inner volume (calculated from the inner radius): cell wall volume=0.32×10-15m3 5. Control volume selectionNote that the product of the enzymatic reaction, AIP, is allowed to diffuse outside the cell. Hence, it is important to take into account the cell boundaries. It is worth considering whether diffusion or fluid movements will play a significant role.Initially, we defined a control volume assuming that bacteria would grow in close colonies on the plate. We realized that our initial choice of control volume was not accurate, since our bacteria are meant to be used in suspension so we had to reconsider this issue.
Initial Choice of Control Volume
Control volume initial choice
The control volume: The inner boundary is determined by the bacterial cell (proteins after being displayed and cleaved cannot diffuse back into bacterium). The outer boundary is more time scale dependent. We have assumed that after mass cleavage of the display-proteins by TEV, many of these AIPs will bind to the receptors quite quickly (eg. 8 seconds). Our volume is determined by the distance that AIPs could travel outwards by diffusion within that short time. In this way, we are sure that the concentration of AIPs outside our control volume after a given time is approximately 0. This approach is not very accurate and can lead us to false negative conclusions (as in reality there will be a concentration gradient, with highest concentration on the cell wall).
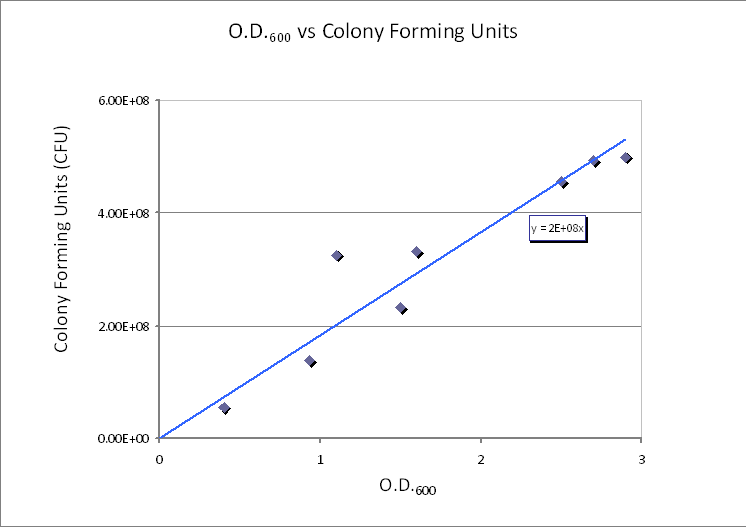
Using CFU to estimate the spacing between cells CFU stands for Colony-forming unit. It is a measure of bacterial numbers. For liquids, CFU is measured per ml. We already have data of CFU/ml from the Imperial iGEM 2008 team, so we could use this data to estimate the number of cells in a given volume using a spectrometer at 600nm wavelength. The graph below is taken from the Imperial iGEM 2008 Wiki page [4].
Side length of cubic Control volume is y = 1.26×10-4 dm = 1.26×10-5 m. Choice of Control Volume allows simplifications 6. Localised concentrationsSince we failed to determine a control volume across which concentration of AIP could be assumed to be uniform, it was deduced that localised concentrations will play an important role in this model. Hence, we tried to come up with some kind of measure of localised concentrations. As the whole reaction happens at the wall and just the final product (AIP) floats freely around, we decided just to scale the AIP concentration by a factor after completion of reaction to simulate the loss of AIPs that diffuse away from cell surface.It was arbitrarily chosen that 20% to 50% of AIPs will bind to receptors rather than diffuse away. There are several arguments that would suggest this kind of percentage:
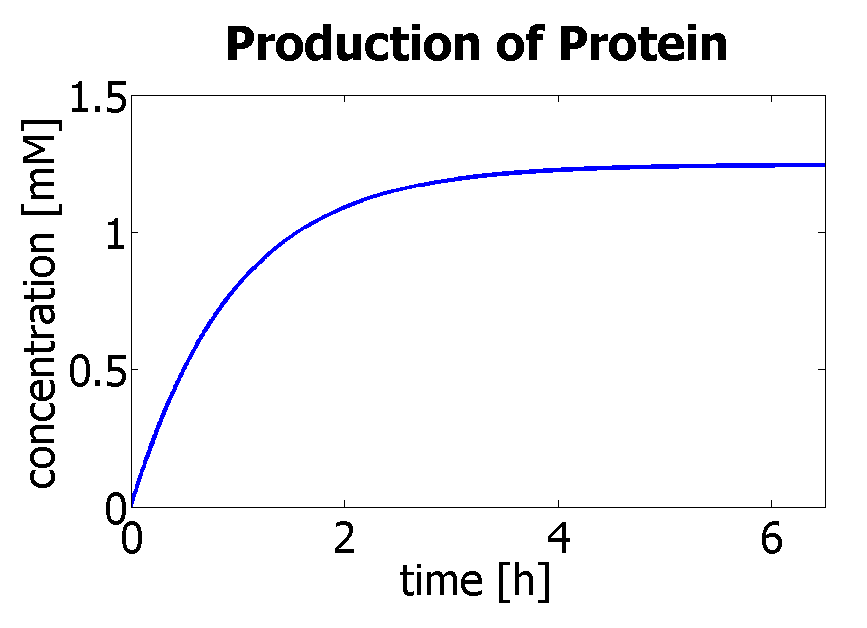
7. Protein production
Hence, we can deduce that the final concentration that the protein expression will tend to is: c = 1.24×10-3 mol/dm3 = cfinal. Therefore, we can model the protein production by transcription and translation and adjust the production constants so that the concentration will tend towards cfinal. The degradation rate was kept constant (same as used in output amplification module), and the production rate was adjusted match the final concentration to be achieved. Using a similar model to the simple production of Dioxygenase for the Output Amplification Model (Model preA), we obtain the following graph:
References
|
 "
"