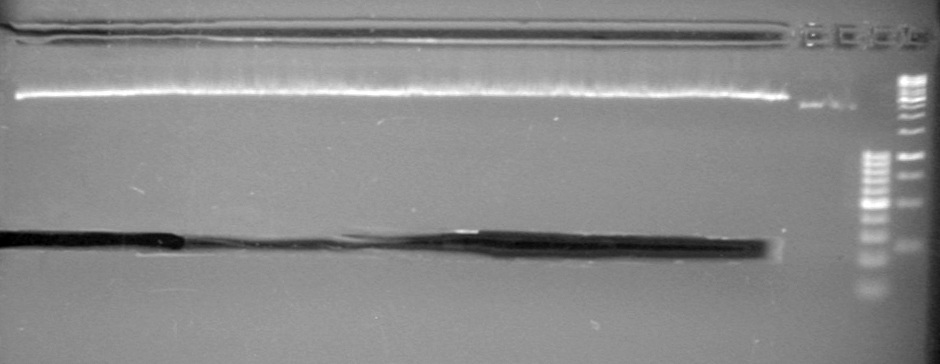
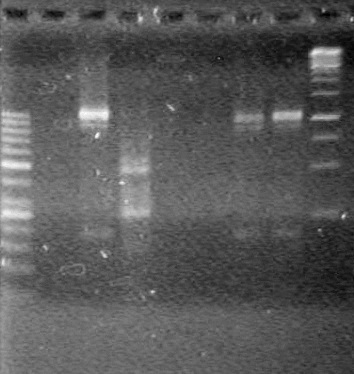
Analyzed results on a 2% agarose gel with 2 (µL) of loading dye and 10 (µL) of pDNA. Load order as follows:
| Component | Volume (µL) |
| MilliQ H2O | 15.6 or 15.8 |
| Buffer | 2 |
| pDNA | 2 |
| Enzyme | 0.20 + 0.20 |
Incubated reactions for 60 minutes at 37oC
Ligation
Reaction set up as follows:
- T4 DNA ligase - 0.25µL
- DNA 1 - 8µL
- DNA 2 - 8µL
- 10x Ligation Buffer - 2µL
- MilliQ H2O - 1.75µL
Incubated reactions overnight at room temperature.
Aug 10, 2010
(In Lab: JV)
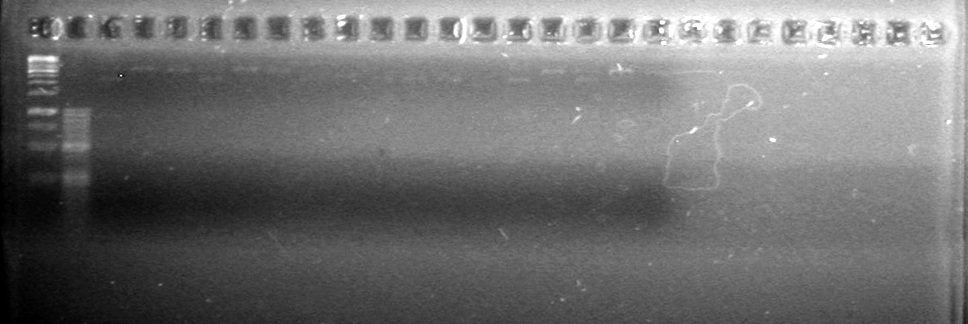
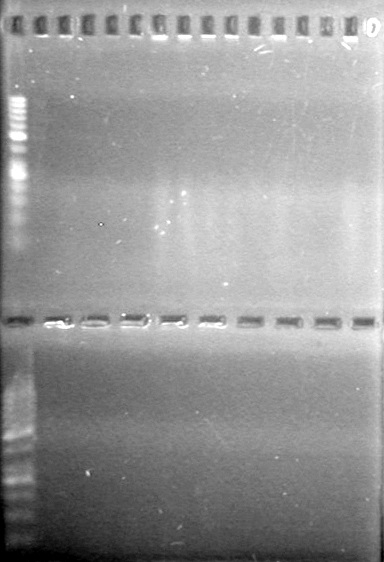
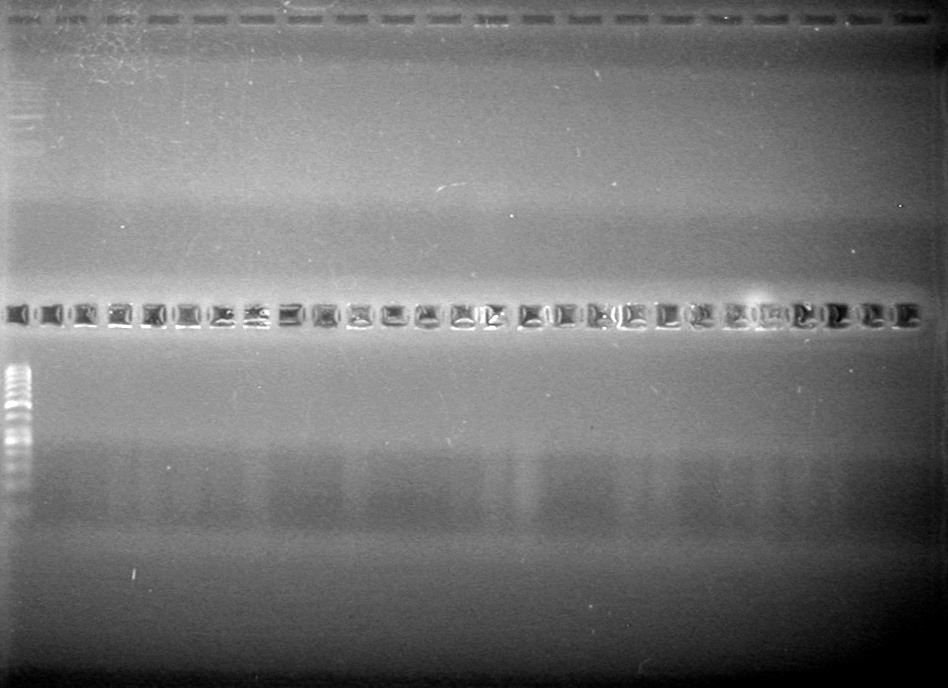
Objective: Reran large gel from Aug 9/2010.
Load order was as follows:
| Lane | Contents
|
| 1 | 50 bp ladder
|
| 2 | pBAD-mRBS 2 URD
|
| 3 | pBAD-mRBS 2 SRD
|
| 4 | pBAD-mRBS 2 DRD
|
| 5 | pLacI-sRBS 3 URD
|
| 6 | pLacI-sRBS 3 SRD
|
| 7 | pLacI-sRBS 3 DRD
|
| 8 | pLacI-mRBS 2 URD
|
| 9 | pLacI-mRBS 2 SRD
|
| 10 | pLacI-mRBS 1 DRD
|
| 11 | dT-pTet 3 URD
|
| 12 | dT-pTet 3 SRD
|
| 13 | dT-pTet 3 DRD
|
| 14 | pBAD-mRBS 1 URD
|
| 15 | pBAD-mRBS 1 SRD
|
| 16 | pBAD-mRBS 1 DRD
|
| 17 | pLacI-sRBS 2 URD
|
| 18 | pLacI-sRBS 2 SRD
|
| 19 | pLacI-sRBS 2 DRD
|
| 20 | 1 kb ladder
|
| Lane | Contents
|
| 1 | 50 bp ladder
|
| 2 | pLacI-mRBS 1 URD
|
| 3 | pLacI-mRBS 1 SRD
|
| 4 | pLacI-mRBS 1 DRD
|
| 5 | dT-pTet 1 URD
|
| 6 | dT-pTet 1 SRD
|
| 7 | dT-pTet 1 DRD
|
| 8 | mRBS-TetR 3 URD
|
| 9 | mRBS-TetR 3 SRD
|
| 10 | mRBS-TetR 3 DRD
|
| 11 | mRBS-TetR 1 URD
|
| 12 | mRBS-TetR 1 SRD
|
| 13 | mRBS-TetR 1 DRD
|
| 14 | pBAD-sRBS 2 URD
|
| 15 | pBAD-sRBS 2 SRD
|
| 16 | pBAD-sRBS 2 DRD
|
| 17 | pBAD-sRBS 1 URD
|
| 18 | pBAD-sRBS 1 SRD
|
| 19 | pBAD-sRBS 1 DRD
|
| 20 | 1 kb ladder
|
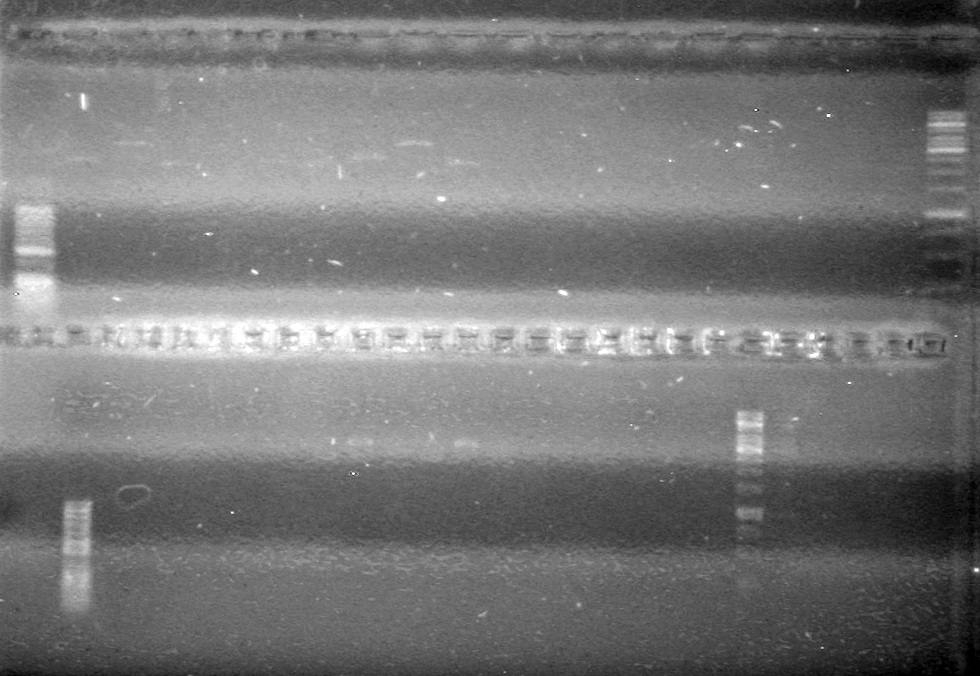

(In Lab: JV)
Objective: Determine which ligations/transformations worked from 08/04/10.
Method: Colony PCR.
A: dT-pTet (1-10)
B: pBAD-sRBS (1-10)
C: pLacI-sRBS (1-10)
D: pBAD-mRBS (1-10)
E: mRBS-TetR (1-10)
F: pLacI-mRBS (1-10)
Pick colony with pipette set at 3(µL)
Pipette colony up and dow in 20(µL) sterile Milli-Q water.
Will use 96 well plate.
PCR- Conditions:
1. 95oC for 15 min
2. 98oC for 20 sec
3. 55oC for 40 sec
4. 72oC for 2 min
5. 72oC for 20 min
6. 4oC infinitely
Reaction Mixture -
Method:
PCR: Thermocycler set to iGEM program 7
| Component | 1X(µL) | Master Mix(x65)(µL)
|
| Milli-Q H2O | 10.9 | 708.5
|
| 5X Phusion HF Buffer | 4 | 260
|
| dNTPs | 1 | 65
|
| Forward Primer (VF2) | 1 | 65
|
| Reverse Primers (VR) | 1 | 65
|
| Colony Template | 2 |
|
| Phusion polymerase | 0.17 | 11.05
|
Controls: sRBS, pTET & mRBS - will PCR these to compare size using same conditions as above.
Aug 10, 2010 Evening
(In Lab: ADS)
Objective: PCR amplify xylE from mRBS-xylE for creation of xylE BioBrick
Method: 20µL reactions
PCR- Conditions:
1. Initial Denaturation 98oC for 30 sec
2. Denaturation 98oC for 10 sec
3. Anneal (51oC, 55oC,59.8oC, 64.6oC, 69.1oC, 71oC) for 30 sec
4. Extend 72oC for 30 sec
5. Final Extend 72oC for 10 min
6. Held 4oC for 30 hours
6 tubes in gradient PCR.
| Component | 1X(µL) | Master Mix(x6.5)(µL)
|
| Milli-Q H2O | 8.8 | 57.2
|
| 5X Phusion HF Buffer | 4 | 26
|
| dNTPs | 2 | 13
|
| Forward Primer | 2 | 13
|
| Reverse Primer | 2 | 13
|
| Template DNA | 1 | 6.5
|
| Polymerase | 0.2 | 1.3
|
Ran samples on 1.5% agarose gel (1X TAE) for 60 minutes at 100V.
Results:

Aug 11, 2010
(In Lab: AS, JV, TF)
Objective: Determine what transformations have the correct insert from Aug. 4, 2010.
Method: Colony PCR. Changes - Used pipette tip instead of toothpick.
PCR: Thermocycler set to iGEM PFUTEST
1. pLacI-mRBS: 4 (A - E) 50(µL)
2. pBAD-mRBS: 5 (A - E) 20(µL)
3. mRBS-TetR: 6 (A -E) 20(µL)
Controls - mRBS
| Component | 1X(µL) | Master Mix(x7.5)(µL)
|
| Milli-Q H2O | 9.8 | 73.5
|
| 10x Pfu Buffer with MgSO4 | 2 | 15
|
| dNTPs | 2 | 15
|
| Forward Primer (VF2) | 2 | 15
|
| Reverse Primer (VR) | 2 | 15
|
| Template DNA | 2 |
|
| Pfu polymerase | 0.2 | 1.5
|
Added 18µL Master Mix to each reaction tube.
(In Lab: ADS)
Objective: Transform mRBS-xylE BioBrick into DH5alpha.
Transformation -
A) Thawed 50(µL) Sub-Cloning Efficiency DH5alpha Competent Cells on ice.
B) Gently mixed cells and then aliquoted 100(µL) into chilled polypropylene tubes.
C) Added 2(µL) of BioBrick to cells. Added 5(µL) of pUC19 DNA to 100(µL) cells to determine efficiency.
D) Incubated cells on ice for 30 minutes.
E) Heat shocked cells for 45 seconds in a 42oC water bath.
F) Placed on ice for 5 minutes.
G) Added 0.4 mL of room temperature SOC medium.
H) Shook at 225 rpm for 1 hour.
I) Diluted control cells 1:100 with SOC medium.
J) Spread 100(µL) of this dilution on LB-Amp agar plates
K) Spread 50 and 250(µL) of experimental cells on LB-Cam agar plates.
L) Incubated overnight at 37oC
Aug 12, 2010
(In Lab: JV)
Objective: Determine if Adam's colony PCRs' from Aug. 11, 2010 worked.
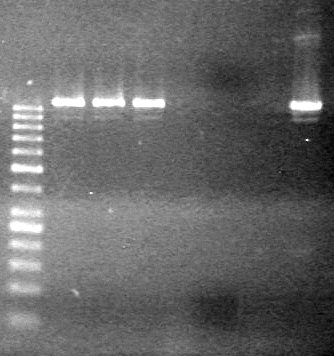
Method: Samples were run on a 2.5% agarose gel (1X TAE) for 1 hour at 100V.
GEL PICTURE!
Results: Lanes 2, 3, 4 and 8 showed PCR amplification. Colonies chosen don't show the correct insert size.
(In Lab: JV)
Objective: Screen for colonies with the correct insert from Aug. 4, 2010 transformations.
Method: Colony PCR. Changes - Used pipette tip instead of toothpick. Put colony in 20(µL) autoclaved Milli-Q water.
PCR: Thermocycler set to iGEM PFUTEST
1. pLacI-sRBS: A (11 - 17)
2. pBAD-sRBS: B (11 - 17)
3. dT-pTet: C (11 - 17)
4. pLacI-mRBS: D (11-17)
5. pBAD-mRBS: E (11-17)
6. mRBS-TetR: F (11-17)
Controls - mRBS, pTet, sRBS
| Component | 1X(µL) | Master Mix(x47)(µL)
|
| Milli-Q H2O | 9.8 | 460.6
|
| 10x Pfu Buffer with MgSO4 | 2 | 94
|
| dNTPs | 2 | 94
|
| Forward Primer (VF2) | 2 | 94
|
| Reverse Primer (VR) | 2 | 94
|
| Template DNA (Cell Lysate) | 2 | 94
|
| Pfu polymerase | 0.2 | 9.4
|
Added 18µL Master Mix to each reaction tube.
Analyzed PCR products on 2.5% TAE gel.
Results:
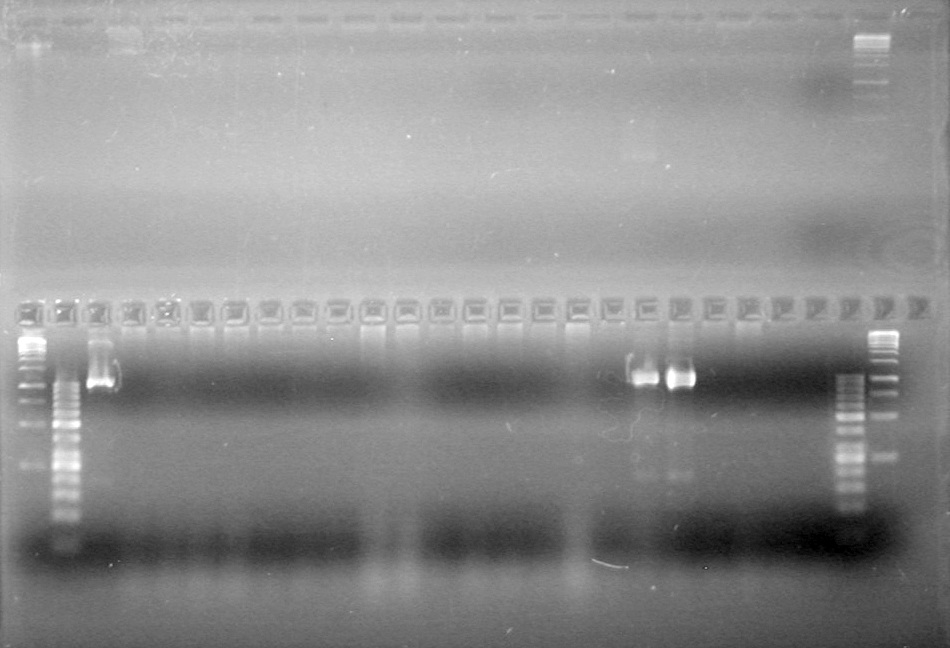
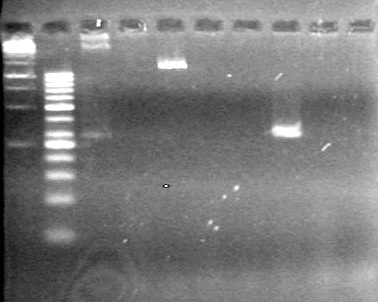
(In Lab: ADS, KG)
Objective: Perform PCR on lumazine, mms6, xylE plasmids with prefix and suffix primers (these will tell us exact size without subtracting VF2/VR regions). If right will ligate into pET28a plasmids.
Method: Plasmids used included: 6 mms6 maxipreps. 4 lumazine maxipreps. 5 xylE maxipreps. 1 mRBS maxiprep
PCR: Thermocycler set to iGEM Program #11 PFU - P/S
PCR- Conditions:
1. Initial Denaturation 95oC for 3 min
2. Denaturation 95oC for 30 sec
3. Anneal (54oC) for 30 sec
4. Extend 72oC for 3 min
5. Final Extend 72oC for 15 min
6. Held 4oC infinitely
(25 cycles)
| Component | 1X(µL) | Master Mix(x16.5)(µL)
|
| Milli-Q H2O | 9.8 | 161.7
|
| 10x Pfu Buffer with MgSO4 | 2 | 33
|
| dNTPs | 2 | 33
|
| Forward Primer (Prefix) | 2 | 33
|
| Reverse Primer (Suffix Antisense) | 2 | 33
|
| Template DNA (Cell Lysate) | 2 |
|
| Pfu polymerase | 0.2 | 3.3
|
Added 18µL Master Mix to each reaction tube.
Analyzed PCR products on 2.5% TAE gel run at 100 V for 35 minutes.
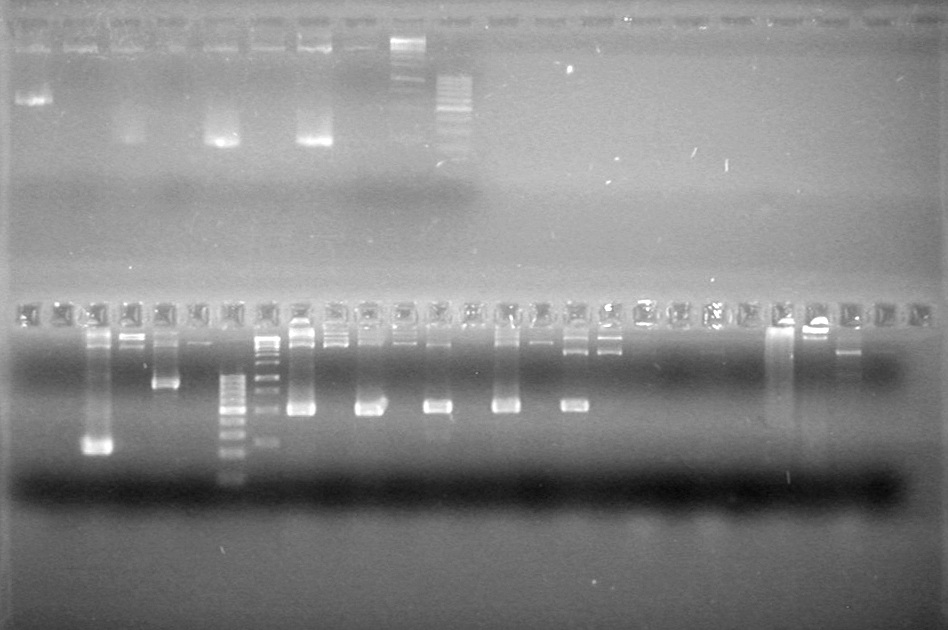
Aug 13, 2010
(In Lab: AS)
Objective: PCR amplify minipreps prepared on Aug 9/2010 to screen for properly assembled BioBricks.
| Component | 1X(µL) | Master Mix(x12.5)(µL)
|
| Milli-Q H2O | 41.85 | 523.1
|
| 10x Pfu Buffer with MgSO4 | 5 | 62.5
|
| dNTPs | 1 | 12.5
|
| Forward Primer (VF2) | 0.5 | 6.25
|
| Reverse Primer (VR) | 0.5 | 6.25
|
| Template DNA | 1 |
|
| Pfu polymerase | 0.15 | 1.888
|
Added 49µL Master Mix to each reaction tube.
Results:
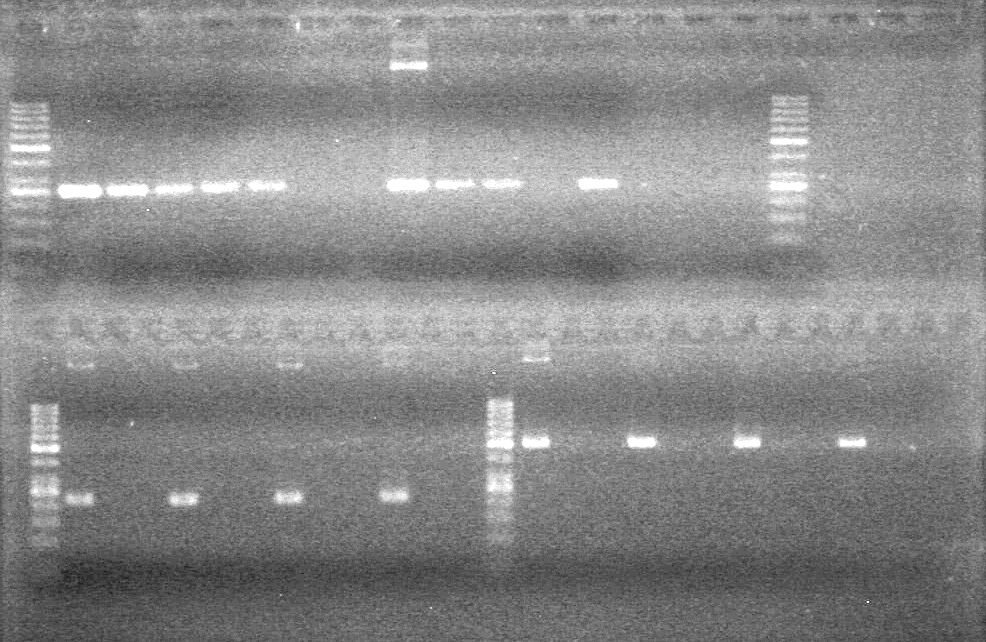
(In Lab: AS)
Objective: PCR amplify pSB1A3, pSB1T3 and pSB1C3 for use in future 3 part assembly and subsequent growth for glycerol stock.
| Component | 1X(µL) | Master Mix(x13.5)(µL)
|
| Milli-Q H2O | 14.9 | 52.15
|
| 10x Pfu Buffer with MgSO4 | 2 | 7
|
| dNTPs | 1 | 3.5
|
| Forward Primer (SB-prep-2) | 0.7 | 2.45
|
| Reverse Primer (SB-prep-3p) | 0.7 | 2.45
|
| Template DNA | 0.5 |
|
| Pfu polymerase | 0.2 | 0.7
|
PCR- Conditions:
1. 94oC for 30 sec
2. 94oC for 30 sec
3. 55oC for 30 sec
4. 68oC for 3 min
5. 68oC for 10 min
6. 4oC infinitely
(36 cycles)
Added 19.5µL Master Mix to each reaction tube.
Analyzed products on 1% TAE agarose gel which ran for 60 minutes at 100 V.
Results:
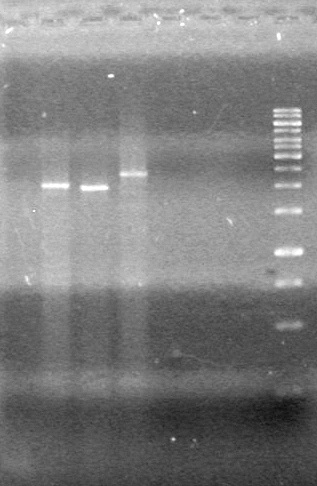
Aug 14, 2010
(In Lab: AS)
2.5% agarose gel(1x TAE)
| lane | contents
|
| 1 | pBAD-mRBS 1
|
| 2 | pBAD-mRBS 2
|
| 3 | pBAD-sRBS 1
|
| 4 | pBAD-sRBS 2
|
| 5 | mRBS-TetR 1
|
| 6 | mRBS-TetR 3
|
| 7 | 50 bp Ladder
|
| 8 | dT-pTet 1
|
| 9 | dT-pTet 3
|
| 10 | pLacI-mRBS 1
|
| 11 | pLacI-mRBS 2
|
| 12 | pLacI-sRBS 2
|
| 13 | pLacI-sRBS 3
|
| 14 | MT
|
| 15 | MT
|
| 16 | MT
|
| 17 | MT
|
| 18 | MT
|
| 19 | MT
|
| 20 | MT
|
| 21 | No Lanes
|
| 22 | No Lanes
|
| 23 | No Lanes
|
| 24 | No Lanes
|
| 25 | No Lanes
|
| 26 | No Lanes
|
| 27 | No Lanes
|
| lane | contents
|
| 1 | K249001
|
| 2 | K249004
|
| 3 | K249005
|
| 4 | K249006
|
| 5 | MT
|
| 6 | K249008
|
| 7 | K249008 (Qiagen)
|
| 8 | K249014
|
| 9 | K249017
|
| 10 | 50 bp Ladder
|
| 11 | 1
|
| 12 | 2
|
| 13 | 3
|
| 14 | 4
|
| 15 | 5
|
| 16 | xylE-dT
|
| 17 | Lumazine-dT
|
| 18 | pLacI-sRBS
|
| 19 | MT
|
| 20 | MT
|
| 21 | MT
|
| 22 | MT
|
| 23 | MT
|
| 24 | MT
|
| 25 | MT
|
| 26 | MT
|
| 27 | MT
|
GEL PICTURE!
(In Lab: AS)
Objective: Insert mms6 and lumazine into pET28a using NotI restriction site.
Method: 1. Reserve 1(µL) of dirty PCR product for analysis. 2. Clean up PCR products using Qiagen prep. 3. Restrict 4 mms6 maxipreps and 4 lumazine maxipreps.
Restriction Reactions:
For lumazine and mms6 -
| Ingredient | 1X(µL) | Master Mix(x8.5)(µL)
|
| MilliQ H20 Water | 7.8 | 66.30
|
| Orange Buffer (10x) | 2 | 17
|
| pDNA | 10 |
|
| NotI | 0.2 | 1.7
|
Added 10 (µL) to each tube.
For pET28a -
| Ingredient | Reaction Mix(µL)
|
| MilliQ H20 Water | 4.8
|
| Orange Buffer (10x) | 5
|
| pDNA | 40
|
| NotI | 0.2
|
Incubated at 37oC.
Analyzed results on 2.5% TAE agarose gel which ran at 100 V for 50 minutes.
GEL PICTURE!!!
Results: Lost all the DNA in the column clean-up step and will have to re-do.
Aug 14, 2010 Evening
(In Lab: AS)
Objective: Assemble mms6-dT and lumazine-dT using three antibiotic assembly.
Method: 1. PCR amplify BioBricks (Prefix/Suffix) 2. Restrict BioBricks 3. Ligate BioBricks into psB1C3 4. Confirm ligation by PCR analysis (VF2/VR) 5. Transform ligation mixes 6. Screen colonies with Colony PCR
PCR: Thermocycler set to iGEM program 11
| Component | 1X(µL) | Master Mix(x10.5)(µL)
|
| Milli-Q H2O | 10.8 | 113.4
|
| 10x Pfu Buffer with MgSO4 | 2 | 21
|
| dNTPs | 2 | 21
|
| Forward Primer (Prefix) | 2 | 21
|
| Reverse Primers (Suffix Antisense) | 2 | 21
|
| Template DNA | 1 |
|
| Pfu polymerase | 0.2 | 2.1
|
Added 19 (µL) to each tube.
Restriction Reactions:
For lumazine and mms6 -
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 11.6 | 63.8
|
| Red Buffer (10x) | 2 | 11
|
| pDNA | 6 |
|
| EcoRI | 0.2 | 1.1
|
| SpeI | 0.2 | 1.1
|
Added 14 (µL) to each tube.
For dT -
| Ingredient | Reaction Mix(µL)
|
| MilliQ H20 Water | 58
|
| Tango Buffer (10x) | 10
|
| pDNA | 30
|
| XbaI | 1
|
| PstI | 1
|
For psB1C3 -
| Ingredient | Reaction Mix(µL)
|
| MilliQ H20 Water | 70
|
| Orange Buffer (10x) | 10
|
| pDNA | 30
|
| EcoRI | 1
|
| PstI | 1
|
| DpnI | 1
|
Also cut lumazine, mms6 and dT with one enzyme for two part, PCR amplification and subsequent ligation into pSB1X3.
For lumazine and mms6, CUT with SpeI -
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 15.8 | 86.9
|
| Tango Buffer (10x) | 2 | 11
|
| pDNA | 2 |
|
| SpeI | 0.2 | 1.1
|
Added 18 (µL) to each reaction.
For dT, CUT with XbaI-
| Ingredient | Reaction Mix(µL)
|
| MilliQ H20 Water | 79
|
| Tango Buffer (10x) | 10
|
| pDNA | 10
|
| XbaI | 1
|
Incubated at 37oC for 1.5 hours. Heat killed enzymes for 20 minutes at 80oC.
Ligation Reactions:
3 Part: Lumazine/mms6 + dT + psB1C3
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 11.0 | 64.9
|
| T4 Ligase Buffer (10x) | 2 | 11
|
| Plasmid (psB1C3) | 2 | 11
|
| Part 1 (Lumazine/mms6) | 2 |
|
| Part 2 (dT) | 2 | 11
|
| T4 DNA Ligase | 0.2 | 1.1
|
2 Part: Lumazine/mms6 + dT
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 13.8 | 75.9
|
| T4 Ligase Buffer (10x) | 2 | 11
|
| Part 1 (Lumazine/mms6) | 2 |
|
| Part 2 (dT) | 2 | 11
|
| T4 DNA Ligase | 0.2 | 1.1
|
Added 18(µL) to each rxn tube. Incubated 1 hour and overnight at room temperature ( 25oC).
Screening via PCR amplification : Thermocycler set to iGEM program 11
3 Part: Lum/mms6 + dT + pSB1C3
| Component | 1X(µL) | Master Mix(x5.5)(µL)
|
| Milli-Q H2O | 33.8 | 185.9
|
| 10x Pfu Buffer with MgSO4 | 5 | 27.5
|
| dNTPs | 2 | 11
|
| VF2 Primer | 2 | 11
|
| VR Primer | 2 | 11
|
| Template DNA | 5 |
|
| Pfu polymerase | 0.2 | 1.1
|
Added 45 (µL) MM to each tube.
2 Part: Lum/mms6 + dT
| Component | 1X(µL) | Master Mix(x5.5)(µL)
|
| Milli-Q H2O | 6.8 | 37.4
|
| 10x Pfu Buffer with MgSO4 | 2 | 11
|
| dNTPs | 2 | 11
|
| Prefix Primer | 2 | 11
|
| Suffix Antisense Primer | 2 | 11
|
| Template DNA | 5 |
|
| Pfu polymerase | 0.2 | 1.1
|
Added 15 (µL) MM to each tube.
Aug 15, 2010
(In Lab: ADS)
Objective: Insert mms6 and lumazine into pET28a using NotI restriction site.
Method: 1. Reserve 1(µL) of dirty PCR product for analysis. 2. Clean up PCR products using Qiagen prep. 3. Restrict 3 mms6 maxipreps and 2 lumazine maxipreps.
Restriction Reactions:
For lumazine and mms6 -
| Ingredient | 1X(µL) | Master Mix(x10.5)(µL)
|
| MilliQ H20 Water | 11.8 | 123.9
|
| Orange Buffer (10x) | 2 | 21
|
| pDNA | 6 |
|
| NotI | 0.2 | 2.1
|
Added 14 (µL) to each tube.
For pET28a -
| Ingredient | Reaction Mix(µL)
|
| MilliQ H20 Water | 4.8
|
| Orange Buffer (10x) | 5
|
| pDNA | 40
|
| NotI | 0.2
|
Incubated at 37oC for 1.5 hours. Heat killed enzymes at 80oC for 20 minutes.
Ligation Reactions:
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 13.9 | 75.9
|
| T4 Ligase Buffer (10x) | 2 | 11
|
| Plasmid (pET28a) | 2 | 11
|
| Cut out part (Lumazine/mms6) | 2 |
|
| T4 DNA Ligase | 0.2 | 1.1
|
Added 18(µL) to each tube. Incubated 1 hour and overnight at room temperature.
Aug 15, 2010 Evening
(In Lab: AS)
Objective: Assemble Lum-dT & mms6-dT using BioBrick standard assembly.
Method: Obtain plasmid DNA from maxipreps.
Restriction Reactions:
For lumazine and mms6 -
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 7.6 | 41.8
|
| Red Buffer (10x) | 2 | 11
|
| pDNA | 10 |
|
| EcoRI | 0.2 | 1.1
|
| SpeI | 0.2 | 1.1
|
For dT -
| Ingredient | Reaction Mix(µL)
|
| MilliQ H20 Water | 38
|
| Orange Buffer (10x) | 10
|
| pDNA | 50
|
| XbaI | 1
|
| EcoRI | 1
|
Incubated at 37oC for 1.5 hours. Heat killed enzymes for 20 minutes at 80oC.
Ligation Reactions:
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 13.8 | 75.9
|
| T4 Ligase Buffer (10x) | 2 | 11
|
| Plasmid (dT) | 2 | 11
|
| Cut out part (Lumazine/mms6) | 2 |
|
| T4 DNA Ligase | 0.2 | 1.1
|
Added 18(µL) MM to each rxn tube. Incubated at one hour and overnight at room temperature.
Screening via PCR amplification : Thermocycler set to iGEM program 11
| Component | 1X(µL) | Master Mix(x5.5)(µL)
|
| Milli-Q H2O | 36.8 | 185.9
|
| 10x Pfu Buffer with MgSO4 | 5 | 27.5
|
| dNTPs | 2 | 11
|
| VF2 Primer | 2 | 11
|
| VR Primer | 2 | 11
|
| Template DNA | 2 |
|
| Pfu polymerase | 0.2 | 1.1
|
Added 45 (µL) MM to each tube.
Analyzed PCR products of BioBrick standard assembly; 3 part (or 3 antibiotic) assembly; and 3 part (3AB) Intermediate/2 part assembly on a 2% TAE agarose gel.
GEL PICTURE!
(In Lab: ADS)
Objective: Produce large quantities of pSB1A3, pSB1C3 and pSB1T3.
Method:
| Component | 1X(µL) | Master Mix(x3.2)(µL)
|
| Milli-Q H2O | 75 | 240
|
| 10x Pfu Buffer with MgSO4 | 10 | 32
|
| dNTPs | 5 | 16
|
| Primer (SB-prep-2Ea) | 3.5 | 11.2
|
| Primer (SB-prep-3P) | 3.5 | 11.2
|
| Template DNA | 2 |
|
| Pfu polymerase | 1 | 3.2
|
Added 98(µL) to each tube. Used PLASRB PCR Protocol from Aug. 13, 2010.
Aug 16, 2010
(In Lab: KG)
Objective: Confirm overnight ligations done on August 15, 2010.
Method:
Screening via PCR amplification : Thermocycler set to iGEM program 4
| Component | 1X(µL)
|
| Milli-Q H2O | 33.8
|
| 10x Pfu Buffer with MgSO4 | 5
|
| dNTPs | 2
|
| VF2 Primer | 2
|
| VR Primer | 2
|
| Template DNA | 5
|
| Pfu polymerase | 0.2
|
3AB Master Mix
| Component | Master Mix(x5.5)(µL)
|
| Milli-Q H2O | 185.9
|
| 10x Pfu Buffer with MgSO4 | 27.5
|
| dNTPs | 11
|
| Prefix Primer | 11
|
| Suffix Primer | 11
|
| Template DNA | 2
|
| Pfu polymerase | 1.1
|
BBS/pSB1C3 Master Mix
| Component | Master Mix(x11)(µL)
|
| Milli-Q H2O | 371.8
|
| 10x Pfu Buffer with MgSO4 | 55
|
| dNTPs | 22
|
| VF2 Primer | 22
|
| VR Primer | 22
|
| Template DNA |
|
| Pfu polymerase | 2.2
|
Analyzed PCR products of overnight BioBrick standard assembly; 3 part (or 3 antibiotic) assembly; and 3 part (3AB) Intermediate/2 part assembly on a 2% TAE agarose gel.
GEL PICTURE!
Aug 16, 2010 Evening
(In Lab: KG, AS)
Objective: Restriction of PCR Products (mms6-dT, lumazine-dT). Restriction is necessary for ligation into plasmid backbone pSB1C3
Method:
Restriction Reactions:
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 7.6 | 41.8
|
| Orange Buffer (10x) | 2 | 11
|
| pDNA | 10 |
|
| PstI | 0.2 | 1.1
|
| SpeI | 0.2 | 1.1
|
Incubated at 37oC for 1.5 hours. Heat killed enzymes for 2 minutes at 80oC.
Ligation Reactions:
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 13.8 | 75.9
|
| T4 Ligase Buffer (10x) | 2 | 11
|
| Plasmid Backbone (pSB1C3) | 2 | 11
|
| pDNA | 2 |
|
| T4 DNA Ligase | 0.2 | 1.1
|
(In Lab: KG)
Objective: Transformations of insertions of mms6 or lumazine into pET28a.
Method: used Competent Cell Transformation protocol
- changes:
- used 50µL aliquottes of DH5&alpha
- did not pipette up and down once, the cells were just swirled 3 times
- added 400µL SOC media, shoock at 370C for 90 min
- platted 250µL and 150µL
| results
| | contents | &250µL | 150µL
|
| + control(pUC19) | good | good
|
| mms6 | good | good
|
| mms6-2 | good | good
|
| mms6 | good | x
|
| Lumazine | good | x
|
| Lumazine | good | x
|
Aug 17, 2010
(In Lab: JV)
Objective: Confirm ligations done on August 16, 2010.
Method:
Screening via PCR amplification : Thermocycler set to iGEM program 4
| Component | 1X(µL) | Master Mix(x5.5)(µL)
|
| Milli-Q H2O | 12.8 | 70.4
|
| 10x Pfu Buffer with MgSO4 | 2 | 11
|
| dNTPs | 1 | 5.5
|
| VF2 Primer | 1 | 5.5
|
| VR Primer | 1 | 5.5
|
| Template DNA | 2 |
|
| Pfu polymerase | 0.2 | 1.1
|
Ran a 2% Agarose gel in 1X TAE buffer for 65 minutes at 100V.
GEL PICTURE!
Aug 17, 2010 Evening
(In Lab: AS)
Objective: Repeat ligation of mms6-dT and lumazine-dT to pSB1C3.
Method: Use already restricted mms6-dT and lumazine-dT. Restrict pSB1C3 PCR product with EcoRI and PstI.
Restriction Reactions:
| Ingredient | 1X(µL)
|
| MilliQ H20 Water | 28.6
|
| Orange Buffer (10x) | 6
|
| pDNA (pSB1C3) | 25
|
| PstI | 0.2
|
| EcoRI | 0.2
|
Incubated at 37oC for 1.5 hours. Heat killed enzymes for 20 minutes at 80oC.
Ligation Reactions:
| Ingredient | 1X(µL) | Master Mix(x5.5)(µL)
|
| MilliQ H20 Water | 13.8 | 75.9
|
| T4 Ligase Buffer (10x) | 2 | 11
|
| Part 2 (pSB1C3) | 2 | 11
|
| Part 1 (mms6-dT/Lum-dT) | 2
|
| T4 DNA Ligase | 0.2 | 1.1
|
Added 18(µL) MM to each rxn tube. Incubated overnight at room temperature.
Objective: Ligation confirmation by PCR. 2 different PCR reaction conditions were utilized. Believe PstI is not being heat inactivated.
Method:
PCR 1 - Show complete insertion of mms6-dT, lumazine-dT into pSB1C3. Both EcoRI and PstI ligations occurred.
| Component | 1X(µL) | Master Mix(x11)(µL)
|
| Milli-Q H2O | 33.8 | 371.8
|
| 10x Pfu Buffer with MgSO4 | 5 | 55
|
| dNTPs | 2 | 22
|
| Forward VF2 Primer | 2 | 22
|
| Reverse VR Primer | 2 | 22
|
| Template DNA | 5 |
|
| Pfu polymerase | 0.2 | 2.2
|
Added 45(µL) MM to each rxn tube.
PCR 2 - Show PstI is not heat killed and only EcoRI ligation occurred.
| Component | 1X(µL) | Master Mix(x16)(µL)
|
| Milli-Q H2O | 33.8 | 540.8
|
| 10x Pfu Buffer with MgSO4 | 5 | 80
|
| dNTPs | 2 | 32
|
| Forward VF2 Primer | 2 | 32
|
| Reverse VR Primer | 2 | 32
|
| Template DNA | 5 |
|
| Pfu polymerase | 0.2 | 3.2
|
Added 45(µL) MM to each rxn tube.
Objective: Analyze tendency of PstI to be heat killed.
Hypothesis: Ligation will be counteracted by PstI digestion even following 20 minute heat kill at 80oC .
Method: Restrict plasmid DNA with PstI (and EcoRI control). Reactions stopped by heat killing. Plasmids re-ligated with T4 DNA Ligase. Ligation confirmed with PCR. If PCR product is present then enzyme heat killed but if there is no PCR Product then enzyme was not heat killed.
Restriction Reactions:
| Ingredient | 1X(µL)
|
| MilliQ H20 Water | 12.8
|
| Orange Buffer (10x) | 2
|
| pDNA (dT) | 5
|
| PstI | 0.2
|
| EcoRI (control only) | 0.2
|
Incubated at 37oC for 1.5 hours. Heat killed enzymes for 20 minutes at 80oC.
Ligation Reactions:
| Ingredient | 1X(µL)
|
| MilliQ H20 Water | 15.8
|
| T4 Ligase Buffer (10x) | 2
|
| Restriction Mix | 2
|
| T4 DNA Ligase | 0.2
|
Incubated at room temperature overnight.
PCR to confirm Ligation
| Component | 1X(µL) | Master Mix(x2.2)(µL)
|
| Milli-Q H2O | 33.8 | 74.36
|
| 10x Pfu Buffer with MgSO4 | 5 | 11
|
| dNTPs | 2 | 4.4
|
| Forward VF2 Primer | 2 | 4.4
|
| Reverse VR Primer | 2 | 4.4
|
| Template DNA | 5 |
|
| Pfu polymerase | 0.2 | 0.44
|
Added 45(µL) to each reaction tube.
Aug 18, 2010
(In Lab: JV)
Objective: Confirmation of ADS' analysis of PstI's tendency to be heat killed.
Method: Will PCR ligation reactions from different assembly methods and EcoRI and PstI controls.
| Component | 1X(µL)
|
| Milli-Q H2O | 33.8
|
| 10x Pfu Buffer with MgSO4 | 5
|
| dNTPs | 2
|
| Forward VF2 Primer | 2
|
| Reverse VR Primer | 2
|
| Template DNA | 5
|
| Pfu polymerase | 0.2
|
Ran on cycle 4 of thermocycler.
Viewed PCRs on 2% TAE agarose gel that ran at 110 V for 30 minutes.
GEL PICTURE!!!
(In Lab: HB)
Objective: Analyze tendency of PstI to be heat killed.
Method: Restrict plasmid DNA with PstI (and EcoRI control). Reactions stopped by heat killing. Plasmids re-ligated with T4 DNA Ligase. Ligation confirmed with PCR. If PCR product is present then enzyme heat killed but if there is no PCR Product then enzyme was not heat killed.
Restriction Reactions:
| Ingredient | 1X(µL)
|
| MilliQ H20 Water | 12.8
|
| Orange Buffer (10x) | 2
|
| pDNA (dT) | 5
|
| PstI | 0.2
|
| EcoRI (control only) | 0.2
|
Incubated at 37oC for 1.5 hours. Heat killed enzymes for 20 minutes at 80oC.
Ligation Reactions:
| Ingredient | 1X(µL)
|
| MilliQ H20 Water | 15.8
|
| T4 Ligase Buffer (10x) | 2
|
| Restriction Mix | 2
|
| T4 DNA Ligase | 0.2
|
Incubated at room temperature overnight.
PCR to Confirm Ligation
| Component | 1X(µL) | Master Mix(x6.5)(µL)
|
| Milli-Q H2O | 33.8 | 219.7
|
| 10x Pfu Buffer with MgSO4 | 5 | 32.5
|
| dNTPs | 2 | 13
|
| Forward VF2 Primer | 2 | 13
|
| Reverse VR Primer | 2 | 13
|
| Template DNA | 5 |
|
| Pfu polymerase | 0.2 | 1.3
|
Added 45(µL) to each reaction tube.
Used thermocycler iGEM Program 4.
3 different treatments: 1. Restricted, heat killed and ligated. 2. Restricted and heat killed. 3. Restricted and not heat killed.
Aug 19, 2010
(In Lab: JV)
Objective: Determine if any transformations from Aug 16, 2010 have the correct insert.
Method: Pick colonies and incubate at 37oC in LB Media with Kan overnight. Use QIAGEN method to extract plasmid DNA. Restrict plasmid DNA to determine if mms6 or lumazine has correctly ligated into pET-28(A).
Restriction Reactions:
mms6 RESTRICTED -
| Ingredient | 1X(µL) | Master Mix(x31)(µL)
|
| MilliQ H20 Water | 15.75 | 488.25
|
| Red Buffer (10x) | 2 | 62
|
| Template DNA | |
|
| Enzyme (EcoRV) | 0.25 | 7.75
|
mms6 UNRESTRICTED -
| Ingredient | 1X(µL) | Master Mix(x31)(µL)
|
| MilliQ H20 Water | 16 | 496
|
| Red Buffer (10x) | 2 | 62
|
| Template DNA | |
|
lumazine RESTRICTED -
| Ingredient | 1X(µL) | Master Mix(x31)(µL)
|
| MilliQ H20 Water | 15.75 | 488.25
|
| Tango Buffer (10x) | 2 | 62
|
| Template DNA | |
|
| Enzyme (EcoRV) | .25 | 7.75
|
lumazine UNRESTRICTED -
| Ingredient | 1X(µL) | Master Mix(x31)(µL)
|
| MilliQ H20 Water | 16 | 496
|
| Tango Buffer (10x) | 2 | 62
|
| Template DNA | |
|
Added 18(µL) to each restriction digest reaction. Incubated at 37oC for 1.5 hours. Heat killed enzymes for 20 minutes at 80oC.
Samples were run on a 2% agarose gel in 1X TAE Buffer.
GEL PICTURE!
(In Lab: HB)
Objective: Run a PCR testing VF2/Suffix and Prefix/VR. A colony PCR of lumazine and mms6 in pET28a. A PCR of 15 ligation tests.
(In Lab: JV)
Objective: Screen for colonies with the correct insert from Aug. 4, 2010 transformations.
Method: Colony PCR. Changes - Used pipette tip instead of toothpick. Put colony in 20(µL) autoclaved Milli-Q water.
PCR: Thermocycler set to iGEM Program 4
| Component | 1X(µL)
|
| Milli-Q H2O | 9.8
|
| 10x Pfu Buffer with MgSO4 | 2
|
| dNTPs | 2
|
| Forward Primer (VF2) | 2
|
| Reverse Primer (VR) | 2
|
| Template DNA (Cell Lysate) | 2
|
| Pfu polymerase | 0.2
|
Added 18µL Master Mix to each reaction tube.
Method: Control test of primer combinations.
Combination 1 - VF2/Suffix
Combination 2 - Prefix/VR
Combination 3 - Prefix/Suffix
dT maxiprep used as known template DNA source.
PCR Combination 1:
| Component | 1X(µL)
|
| Milli-Q H2O | 9.8
|
| 10x Pfu Buffer with MgSO4 | 2
|
| dNTPs | 2
|
| Forward Primer (Prefix) | 2
|
| Reverse Primer (Suffix) | 2
|
| Template DNA (dT) | 2
|
| Pfu polymerase | 0.2
|
PCR Combination 2:
| Component | 1X(µL)
|
| Milli-Q H2O | 9.8
|
| 10x Pfu Buffer with MgSO4 | 2
|
| dNTPs | 2
|
| Forward Primer (Prefix) | 2
|
| Reverse Primer (VR) | 2
|
| Template DNA (dT) | 2
|
| Pfu polymerase | 0.2
|
PCR Combination 3:
| Component | 1X(µL)
|
| Milli-Q H2O | 9.8
|
| 10x Pfu Buffer with MgSO4 | 2
|
| dNTPs | 2
|
| Forward Primer (VF2) | 2
|
| Reverse Primer (suffix) | 2
|
| Template DNA (dT) | 2
|
| Pfu polymerase | 0.2
|
Method: PCR of 15 ligation tests.
| Component | 1X(µL) | Master Mix(x19.5)(µL)
|
| Milli-Q H2O | 9.8 | 191.1
|
| 10x Pfu Buffer with MgSO4 | 2 | 39
|
| dNTPs | 2 | 39
|
| Forward VF2 Primer | 2 | 39
|
| Reverse VR Primer | 2 | 39
|
| Template DNA | 2 |
|
| Pfu polymerase | 0.2 | 3.9
|
Added 18(µL) to each tube.
(In Lab: JV)
Analyzed HB's PCR products on 2% TAE gel which ran for 60 minutes at 100 V.
GEL PICTURE!!!
Aug 20, 2010
(In Lab: JV)
Objective: Determine if attempts to PCR amplify plasmid backbone were successful.
Method: Ran samples on 1% agarose gel with 1X TAE buffer for 50 minutes at 100 V.
Results: DNA concentration was good, however there was no evidence of an insert into pET-28(A).
GEL PICTURE!!!
Aug 23, 2010
(In Lab: JV)
Objective: Obtained part <partinfo>BBa_K118021</partinfo> and <partinfo>BBa_I716462</partinfo>.
Method: Used competent cell transformation protocol.
(In Lab: HB)
Objective: Restrict 18 maxiprepped parts to quantify DNA.
Method: Restrict all 18 parts and run on a 1% agarose gel with unrestricted parts.
Restriction Reactions:
Restriction Mix -
| Ingredient | 1X(µL) | Master Mix(x19)(µL)
|
| MilliQ H20 Water | 12.8 | 243.2
|
| Orange Buffer (10x) | 2 | 38
|
| Template DNA | 5 |
|
| EcoRI | 0.2 | 3.8
|
Unrestricted Mix -
| Ingredient | 1X(µL) | Master Mix(x19)(µL)
|
| MilliQ H20 Water | 13 | 247
|
| Orange Buffer (10x) | 2 | 38
|
| Template DNA | 5 |
|
15(µL) added to each rxn tube.
Incubated at 37oC for 1.5 hours.
GEL PICTURE!
(In Lab: AV)
Objective: To PCR amplify pSB1T2, pSB1C3, pSB1A3 and run on 1% agarose gel.
Method:
| Component | 1X(µL) | Master Mix(x6.5)(µL)
|
| Milli-Q H2O | 14.9 | 96.85
|
| 10x Pfu Buffer with MgSO4 | 2 | 13
|
| dNTPs | 1 | 6.5
|
| SB-prep-2 Primer | 0.7 | 4.55
|
| SB-prep-3p Primer | 0.7 | 4.55
|
| Template DNA | 0.5 |
|
| Pfu polymerase | 0.2 | 1.3
|
Added 19.5(µL) to each tube.
Ran on program 6 of thermocycler.
Results: Nothing visible on gel, therefore will have to try loading a larger volume of PCR product.
Aug 24, 2010
(In Lab: AV)
Objective: To re-run a 1% agarose gel of PCR products from Aug 23, 2010.
Results: Nothing appeared on gel, therefore the PCR was unsuccessful.
Aug 25, 2010
(In Lab: JV)
Objective: Create amounts of pSB1C3.
Method:
| Component | 1X(µL) | Master Mix(x6.5)(µL)
|
| Milli-Q H2O | 14.9 | 96.85
|
| 10x Pfu Buffer with MgSO4 | 2 | 13
|
| dNTPs | 1 | 6.5
|
| SB-prep-2 Primer | 0.7 | 4.55
|
| SB-prep-3p Primer | 0.7 | 4.55
|
| Template DNA | 0.5 |
|
| Pfu polymerase | 0.2 | 1.3
|
Added 19.5(µL) to each tube.
PCR- Conditions:
1. 95oC for 5 min
2. 94oC for 30 sec
3. 55oC for 30 sec
4. 68oC for 4 min
5. 68oC for 10 min
6. 4oC infinitely
(36 cycles)
Aug 25, 2010
(In Lab: FM)
Objective: Gradient PCR of xylE.
Method:
| Component | 1X(µL) | Master Mix(x10)(µL)
|
| Milli-Q H2O | 5.8 | 58
|
| 10x Pfu Buffer with MgSO4 | 1 | 10
|
| dNTPs | 1 | 10
|
| Standard Prefix or Fusion Prefix Primer | 0.5 | 5
|
| Standard Suffix or Fusion Suffix Primer | 0.5 | 5
|
| Template DNA | 1 | 10
|
| Pfu polymerase | 0.2 | 2
|
Gradient Temperatures:
58.5oC, 60.5oC, 62.3oC, 64.1oC, 65.9oC, 67.7oC, 69.4oC, 71.1oC
(In Lab: JS)
Objective: Run a 2% agarose gel of gradient PCR of xylE.
GEL PICTURE!!!
Aug 26, 2010
(In Lab: KG)
Objective: Do PCR from Aug 25, 2010 to compare PCR of part from registry and our pSB1C3.
Method:
| Component | 1X(µL) | Master Mix(x2)(µL)
|
| Milli-Q H2O | 14.9 | 29.8
|
| 10x Pfu Buffer with MgSO4 | 2 | 4
|
| dNTPs | 1 | 2
|
| SB-prep-26 Primer | 0.7 | 1.4
|
| SB-prep-3P Primer | 0.7 | 1.4
|
| Template DNA | 0.5 |
|
| Pfu polymerase | 0.2 |
|
PCR- Conditions:
1. 95oC for 5 min
2. 94oC for 30 sec
3. 55oC for 30 sec
4. 68oC for 4 min
5. 68oC for 10 min
6. 4oC infinitely
(36 cycles)
Ran a 1% agarose gel of gradient PCR of xylE.
Was stained in ethidium bromide for too long.
Aug 31, 2010
(In Lab: DM)
Objective: Overexpression test of xylE part (BBa_K118021) in DH5alpha cells in m9 minimal media.
Method: Overexpression
0.5 M Catechol was made, to allow for the addition of 1(µL) of this stock solution to be added to the samples taken during the overexpression.
Upon addition of catechol, the solutions turned yellow!
1 mL samples were taken for SDS-PAGE analysis.
Optical density readings at 600 nm and absorbance readings at 375 and 275 nm were recorded for flasks containing either glucose or sucrose.
(In Lab: ADS)
Objective: PCR amplify plasmid backbones (pSB1C3, pSB1A3 and pSB1T3).
Method: Use PCR product from Aug 13, 2010 as template DNA.
| Component | 1X(µL) | Master Mix(x3.5)(µL)
|
| Milli-Q H2O | 14.4 | 52.4
|
| 10x Pfu Buffer with MgSO4 | 2 | 7
|
| dNTPs | 1 | 3.5
|
| SB-prep-26 Primer | 0.7 | 2.45
|
| SB-prep-3P Primer | 0.7 | 2.45
|
| Template DNA | 1 |
|
| Pfu polymerase | 0.2 | 0.7
|
Added 19(µ:L) to each tube.
Phusion polymerase was used and apparently in the other previously unsuccessful PCRs, instead of Pfu polymerase.
PCR- Conditions:
1. 95oC for 5 min
2. 94oC for 30 sec
3. 55oC for 30 sec
4. 68oC for 4 min
5. 68oC for 10 min
6. 4oC infinitely
(36 cycles)
Analyzed PCR on 1% TAE agarose gel which ran for 60 min at 100 V.
Objective: Obtain new sources of pSB1T3, pSB1C3 and pSB1A3 plasmid backbone. Backbone from registry used up.
Method: pSB1A3 can be obtained from anything team already possesses. pSB1C3 can be obtained from BBa_J04450 (RFP) in kit. Don't have tetracycline plates so cannot obtain pSB1T3. Used competent cell transformation protocol.
Method: pSB1A3 can be obtained from anything team already possesses. pSB1C3 can be obtained from BBa_J04450 (RFP) in kit. Don't have tetracycline plates so cannot obtain pSB1T3. Used Competent Cell Transformation protocol.
- changes:
- used 50µL aliquottes of DH5&alpha
- did not pipette up and down once, the cells were just swirled 3 times
- added 500µL SOC media, shoock at 370C for 90 min
- platted 250µL and 150µL
| Results
| | contents | &250µL | 150µL
|
| + control(pUC19) | good | good
|
| J04450 | X (TMTC) | X(TMTC)
|
Prepared overnight cultures for minipreps. Added 5(µL) of 35 mg/mL Chloramphenicl to 5 mL of LB Media. Inoculated media with cells from single colony picked from transformation plate. Incubated overnight at 37oC with shaking.
 "
"