Team:Tsinghua/project/outline/m2
From 2010.igem.org
(→Strategy 1 Bacterial Based Microarray) |
(→Strategy 2 ToxR-based Transmembrane Signaling Pathway Method) |
||
| Line 61: | Line 61: | ||
<br/> | <br/> | ||
| + | <html><a name="m2s2"></a></html> | ||
===Strategy 2 '''ToxR-based Transmembrane Signaling Pathway Method'''=== | ===Strategy 2 '''ToxR-based Transmembrane Signaling Pathway Method'''=== | ||
Revision as of 14:08, 27 October 2010

Module II
Strategy 1 Bacterial Based Microarray
Outline of screening desired antibody by bacterial based microarray
Microarray technique is a kind of pretty mature biomedical technology applied to diverse areas from pharmacology research to clinical use, such as genetics diagnose. Due to the advantage that large amount of various substances can be screened in microarray, microarray can be used for selection of desired antibodies with specific affinity in our project. However, current existing microarray technology rely on specific DNA or protein attached substrate and our project lean on bacteria expressing specific surface protein used for screening for desired antibody. Therefore, traditional protocols and materials used in microarray will be adjusted in our project. We believe that our adjustment will improve the efficiency of antibody screening and help our project achieve its ultimate goal.
Specific description of the details of the experiments
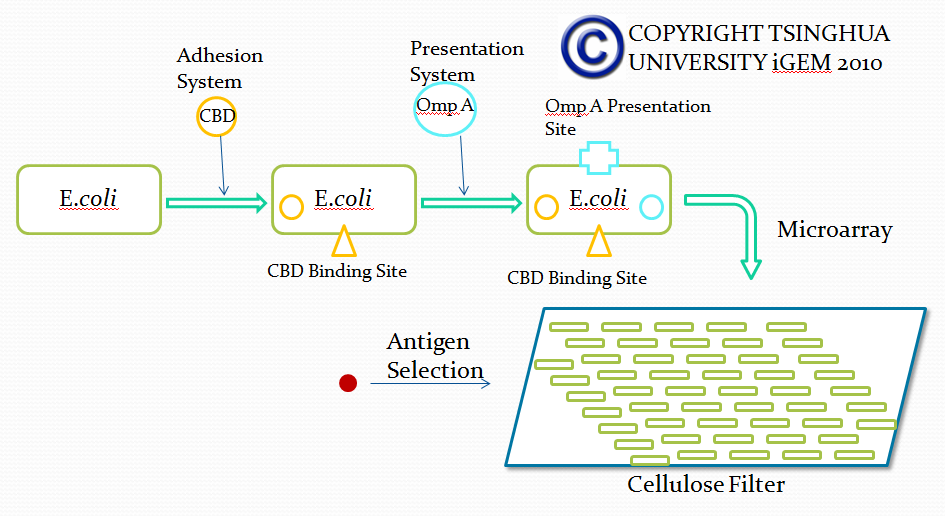
In this part, we want to transform two batches of E coli with vectors carrying specific genes expressing various kinds of antibody achieved by the recombination methods in our project and specific antigen or some other protein used to select antibody from the aforementioned mix of antibodies. By fusion antibody and antigen gene with the genes of membrane integral displaying protein called OmpA, we can manage to display our protein to the surface of the bacteria. Therefore, the selection will process through interaction between antibody and antigen at the surface of the bacteria. Then, by linking the gene of display protein to the gene of one kind of protein called cellulose binding domain which can bind to cellulose, we are able to anchor the bacteria expressing CBD to the surface of the specific microarray coated with cellulose substrate. The whole process is illustrated in the following figure.
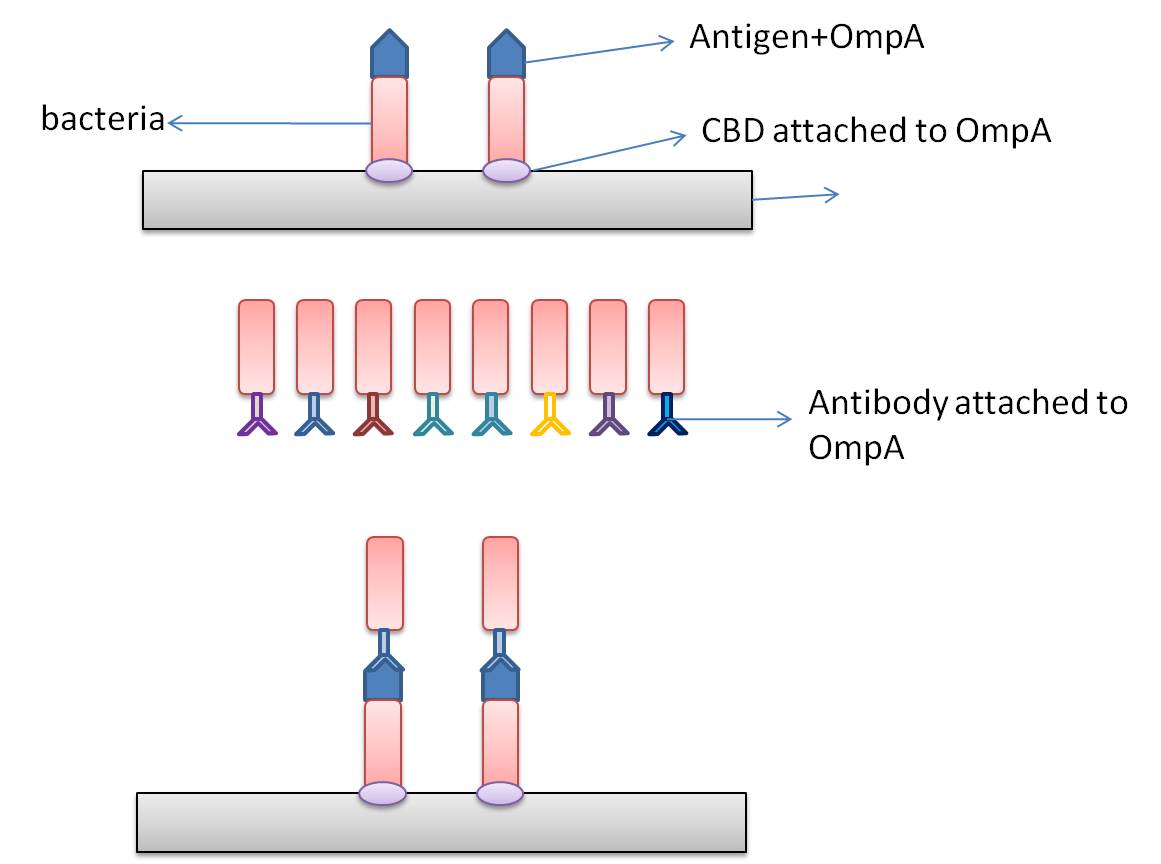
Information concerning the specific proteins
Cellulose binding domain
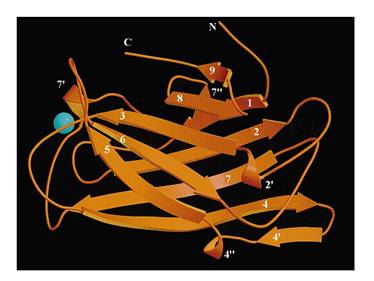
The protein used in this part comes from CipC gene found in Clostridium cellulolyticum. CipC gene encodes a protein called cohesion which participate in degradation of cellulose for the benefits of the bacteria Clostridium cellulolyticum. The protein encoded in this gene contains a specific domain called cellulose binding domain which bind to cellulose with pretty high binding affinity and thus facilitate the degradation of the cellulose. The structure of CBD has been resolved shown in the figure below. Our purpose is to utilize the ability of CBD to bind to cellulose and express CBD in E coli and thus force it to bind to microarray coated with cellulose substrate.
Displaying protein OmpA
At present, two kinds of membrane protein are known, that is, alpha helix protein from cytoplasmic membrane and beta-barrel protein in bacteria outer-membrane, exemplified by porin protein. There are two membrane layers in gram-negative bacteria, separately called inner membrane and outer membrane. OmpA is located in outer membrane and belongs to beta-barrel category. This protein is responsible for normal physiological functions in gene regulation in E coli. Because OmpA is located in outer membrane, intensive investigation has been conducted in OmpA because OmpA plays an important role in signal transduction and much work has focused on the mechanism how OmpA was transported from cytosol out of the membrane. Because of the clarified function of the protein, some bioengineers managed to utilize its function to display protein to achieve certain purposes. Therefore, we try to take advantage of the properties of OmpA to display protein, that is, to display CBD, antibody and antigen. The structure of OmpA has been determined, illustrated in the following figure.
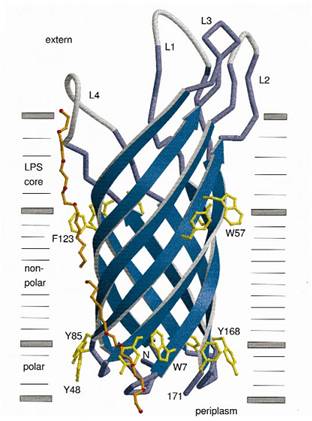
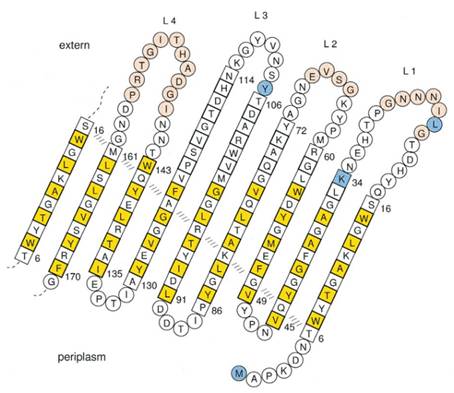
Previous work has demonstrated that 46-159 amino acid is adequate to anchor OmpA to membrane and from the topology figure, we can find out that the fragment of AA 46-159 comprise the transmembrane part of the protein, as the red box indicates. Besides, to ensure the transport of OmpA from cytosol to cell membrane, we need to add a signal peptide to the N terminal of OmpA. Previous work found that one kind of lipoprotein in E coli contains a signal peptide consisting of nine amino acids, which ensures the successful transport of the protein outside the membrane. Mining through the parts provided by iGEM, we found one part which provides exactly the same sequence we wants. This part contains N terminal signal peptide and the sequence AA 46-159 and the linker region downstream of OmpA which ensures the folding of the attached domain, such as CBD. The engineered OmpA is illustrated as following.
The selection of antibodies
Due to commercialization of antibody, we have no access to the cDNA of these antibodies. Therefore, we managed to synthesize the cDNA.
Design of microarray
Microarray is a solid substrate based 2 dimensinoal array, which can be utilized to detect large sum of bio-substance. After decade of developing, various kinds of microarrays have been invented, such as DNA microarray, protein microarray, tissue microarray and so on. Different microarrays can be used to detect different components based on different substrates. Our project belongs to cellular microarray, aiming to select different kinds of antibodies based on the interaction between different proteins displayed on bacteria membrane.
This part of our project aims to attach CBD-expressing bacteria to the surface of microarray coated with cellulose. Weizmannn Institute has developed zephyrin-based microarray, in which the microarray plate is dotted with cellulose. Thus, this innovation provides potential for application in our project.
Strategy 2 ToxR-based Transmembrane Signaling Pathway Method
Propose of This Step
In order to select a specific antibody from a large number of antibodies, we studied the formation of mammalian antibodies. In the process of antibody production in mammals, mediated Annexin antibodies-mediated antibody activation play an important role. This section hopes to use a membrane receptor of E. coli to simulate this process, to achieve the purpose of screening.
Principle and Background
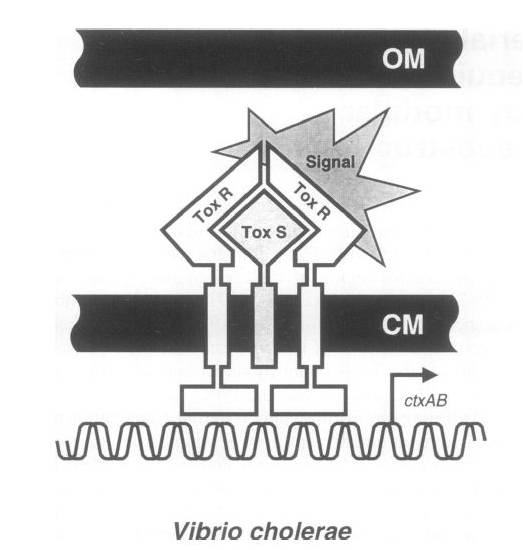
To achieve this, a ToxR-based two-hybrid system was introduced into E.coli cells. The Vibrio cholerae transcriptional regulator ToxR is anchored in the cytoplasmic membrane by a single transmembrane segment, its C-terminal domain facing the periplasm. Most of its N-terminal cytoplasmic domain shares sequence similarity with the winged helix–turn–helix (wHTH) motif of OmpR-like transcriptional regulators. The ToxR protein of Vibrio cholerae regulates the expression of several virulence factors that play important roles in the pathogenesis of cholera.
ToxR stimulates transcription from the cholera toxin gene promoter ctx by direct binding to DNA element ITITTGAT present in different isolates of Vcholerae in three to eight tandemly repeated copies upstream of the ctxAB structural genes.
Transcription activation is thought to be initiated by environmental stimuli which cause the periplasmic ToxR domain to form a homodimer. This, in turn, tethers together the two cytoplasmic ToxR domains, which can now bind to the control region of the ctx promoter. A second membraneassociated protein, ToxS (Mr 19 000), is required for maximal activation of the ctx promoter; most plausibly it stabilizes the ToxR dimer by direct contact.
Figure Shows the Original structure of ToxR system in Vibrio cholerae
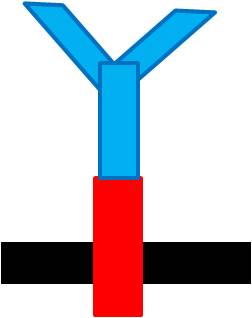
Fusion protein was made between the inner- and trans-membrane parts of ToxR and of the recombinant antibody.
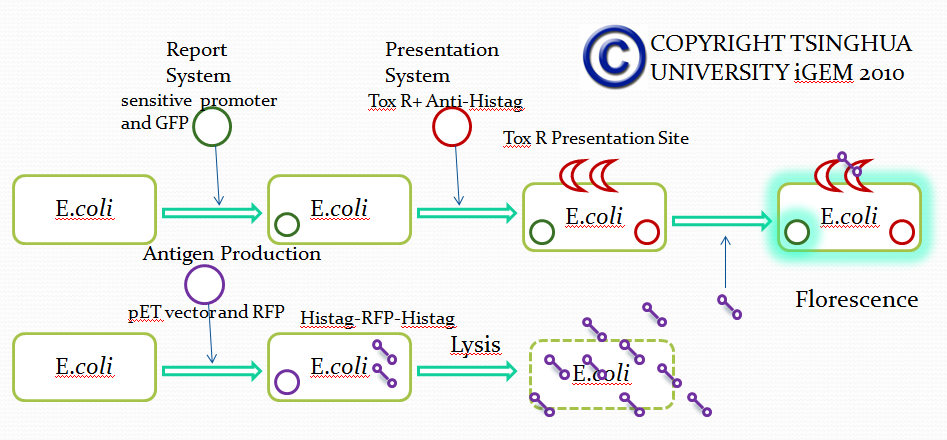
When the specific antigens recognized by antibodies, antibodies with the same antigen recognize site will be close to each other through the antigen,inner-membrane part of “Antibody-ToxR” Protein dimerize, ctx promoter active, Resistance gene starts to be transcribed.
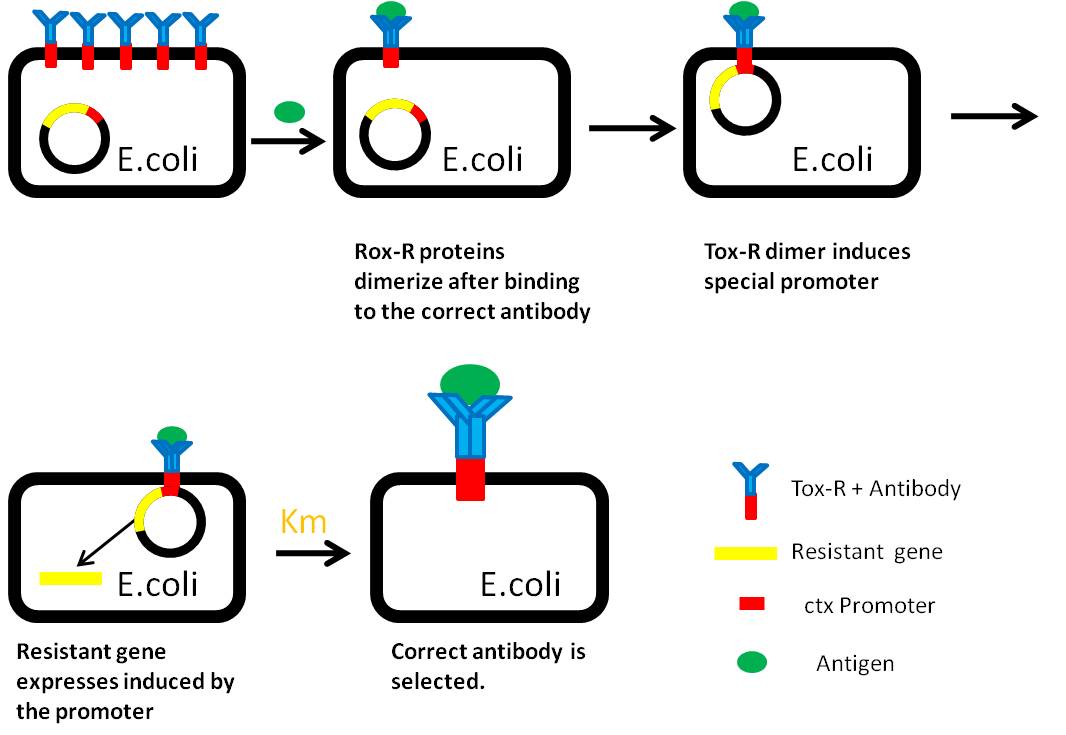
Use “Anti-Histag Antibody” and “Protein with His-tag” as “antibody” and “antigen” to verify the effect of this system.
Antibody: ToxR + Anti-Histag Antibody scFv Light Chain
Plus: ToxR + Anti-Histag Antibody scFv Heavy Chain
Antigen: His tag + Protein A + Histag
Protocol
We need to achieve the construction of the selection vector under the control of ctx promoter, then induce to the bacteria. Measure the expression of the two genes and measure the distribution in membrane. Use the exogenous protein carried His-tag as the antigen. Treat with antibiotics. Compare the expression level between the experiment group and the control, then define the effect of our system indirectly.
Reference:A ToxR-based two-hybrid system for the detection of periplasmic and cytoplasmic protein–protein interactions in Escherichia coli: minimal requirements for specific DNA binding and transcriptional activation
Strategy 3
Cooperation with Macquarie Australia
 "
"







 TOP
TOP