Team:Imperial College London/Results/Fast Response Module
From 2010.igem.org
(New page: {{:Team:Imperial_College_London/Templates/Header}} {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:25px;" cel...) |
|||
| Line 1: | Line 1: | ||
{{:Team:Imperial_College_London/Templates/Header}} | {{:Team:Imperial_College_London/Templates/Header}} | ||
| - | { | + | {{:Team:Imperial_College_London/Templates/ResultsHeader}} |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
|style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Fast Response Module | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Fast Response Module | ||
Latest revision as of 12:52, 27 October 2010
| Experimental Results | Exp 1 | Exp 2 | Exp 3 | Exp 4 | Exp 5 | Exp 6 | Exp 7 |
| Testing is a fundamental stage of the engineering design cycle and is a crucial part of charactrising BioBrick Standard Biological Parts so that other people can benefit from our work. We've compiled all our results on this page, detailing how the experiments were carried out and the significance of the data. | |
| Fast Response Module |
| Catechol 2,3 Dioxygenase (C2,3O) |
|
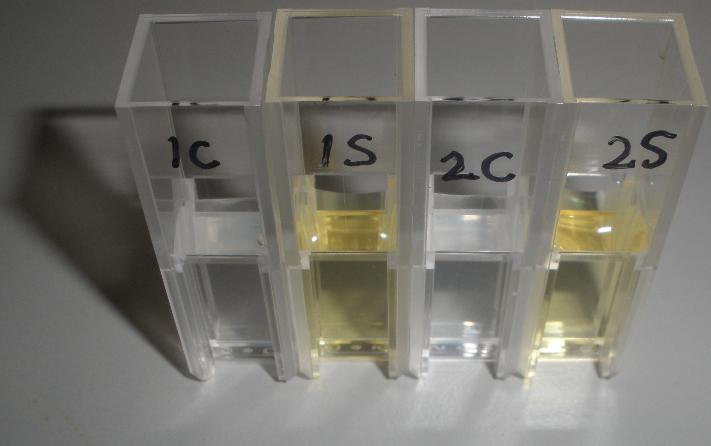
The XylE gene, originating from the TOL plasmid of Pseudomonas putida, encodes the catechol 2,3-dioxygenase (C2,3O) enzyme, the final choice for the output signal mediator of our system. Colonies of cells that express XylE gene product become yellow within seconds after selection plates are sprayed with catechol, a colorless substrate, that is converted by C2,3O to the yellow product, 2-hydroxymuconic semialdehyde. This reporter system is ideal for our project as it has very fast kinetics, gives a visual output by naked eye and the substrate used is inexpensive. |
| Experiment 2 | The threshold value of catechol assay | ||
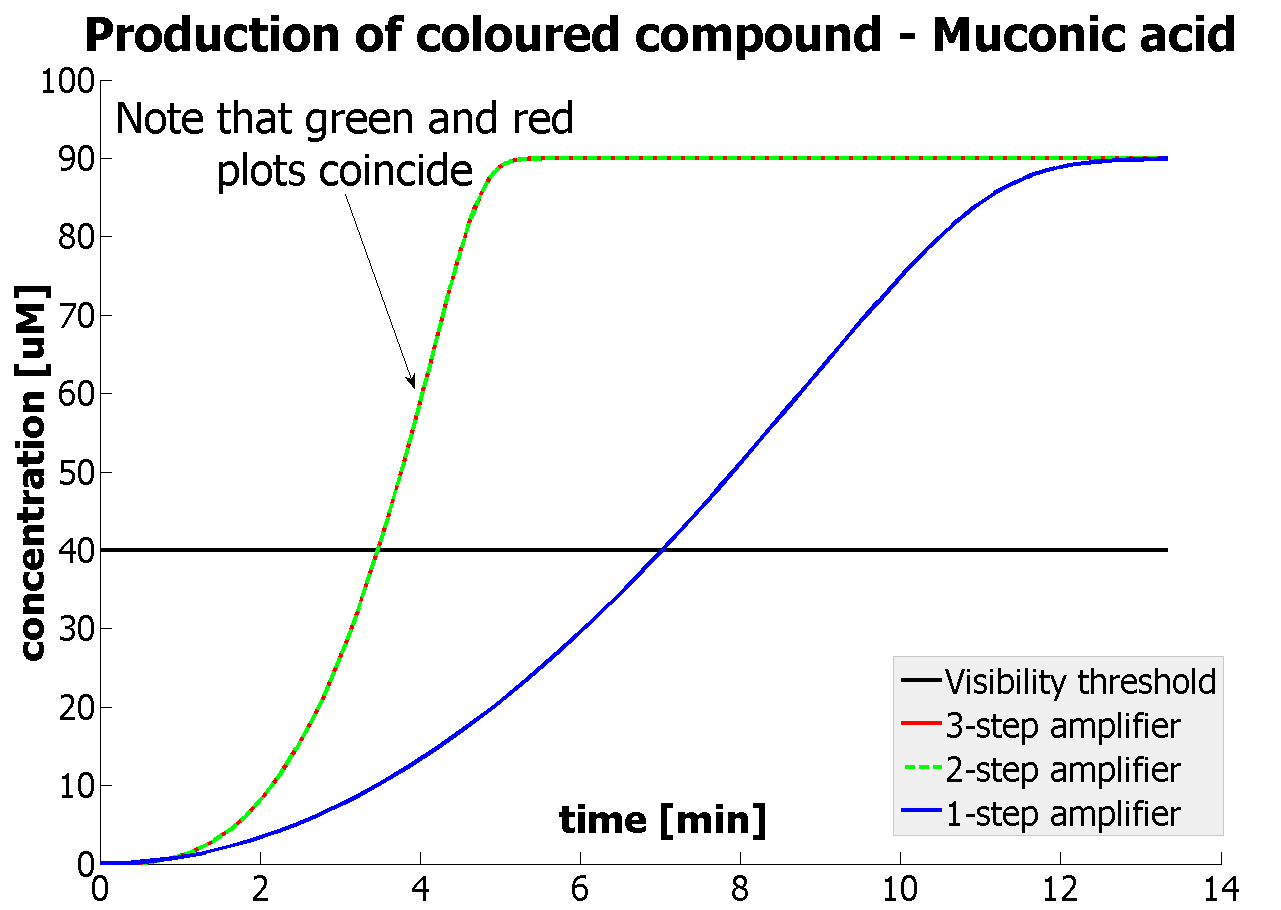
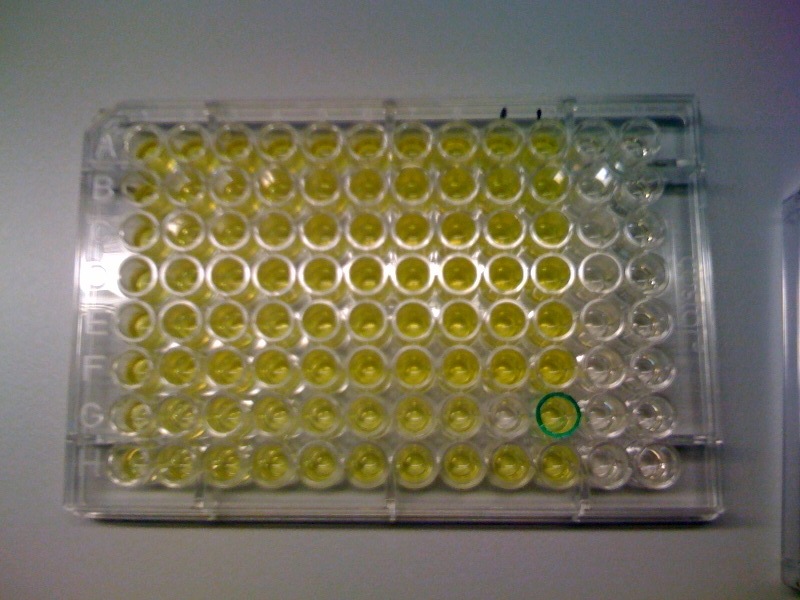
The threshold concentration of catechol needed for a person to identify a yellow positive test was found to be 30-40μM. This result was fed back to the modelling team and allowed them to constrain the concentration values of catechol to be used. Furthermore, it facilitated the determination of conditions for 2-step amplification performing better than 1-step amplifier. If you want to know more on output amplification modelling, please follow the link. |
| Experiment 3 | Characterizing kinetic parameters of C2,3O in whole cells | ||||||
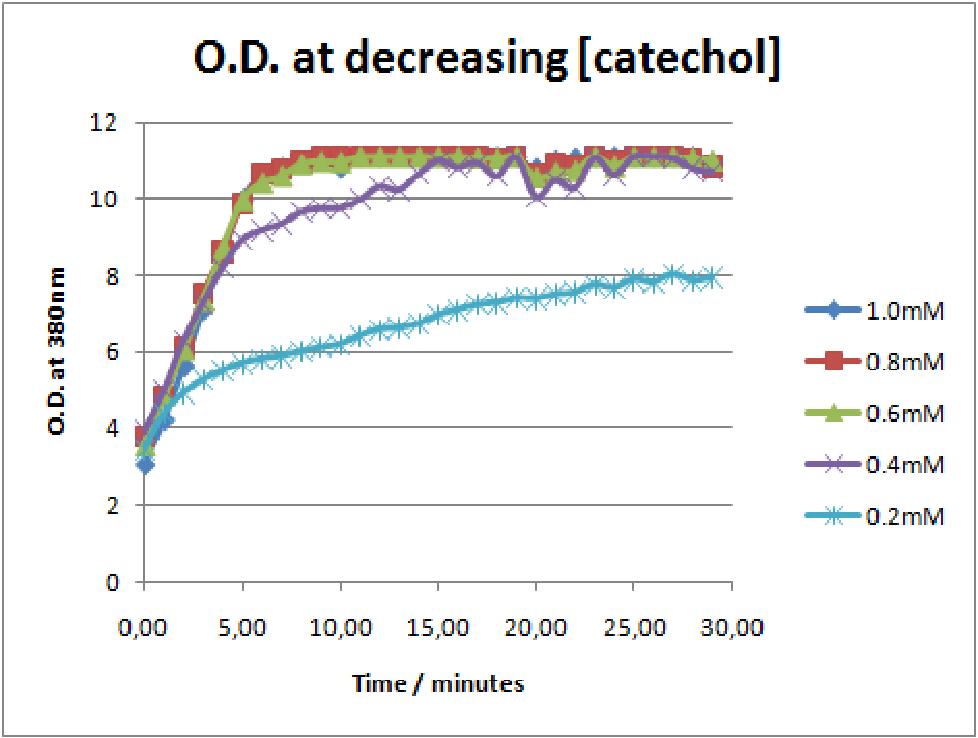
The results of the experiment confirmed previous hints of the huge catalytic turnover rate of the C(2,3)0 enzyme. The graph on the left illustrates production of yellow product per min. The slope at the initial stages of the reaction (first minutes) would represent the rate of the reaction, that is the initial Velocity. By calculating initial velocities of the reaction at different substrate concentrations, one can plot a Michaelis-Menten curve to extract kinetic parameters such as Vmax, Km and catalytic turnover number. Several attempts were made to determine these parameters on whole cells. Unfortunately due to the actual nature of the enzyme, velocity of reaction and technical limitations of lab equipement this has been proven impossible. To start with one of the limitation was that the color output and the extinction coefficient of the yellow product is so large, that the system saturates within the first minutes after catechol addition to cell cultures. The spectrophotometer and plate reader reaches its detection maximum reading value from the start of the reaction. Also due to the high velocity of the reaction, in the time frame of about 10 seconds that are needed to load the plate into the plate reader after catechol addition, the reaction has already proceeded too far. So initial velocity measurements of the reactions needed for construction of Michaelis-Menten curve are inaccurate. These problem could be overcome by decreasing total enzyme concentration. In whole cell cultures this would be possible by decreasing cell density in each well as each cell contain an average C2,3O enzyme concentration. However, previous experiments showed that cell cultures with optical densities lower than 0.5 O.D. introduce serious perturbations in the readings of the spectrophotometer and plate reader detectors. Despite these limitations some useful data were extracted out of these assays. More specifically we acquired data that delineate the course of the reaction in terms of yellow product production over time at various catechol concentrations. The results of one of these assays is presented in the figure below. In order to extract data that will allow characterization of the kinetic parameters of catechol dioxygenase enzyme our lab team proposes purification of the protein from cell lysate, several fold dilution, and in vitro characterization. |
| Experiment 4 | Assaying cell-growth in presence of Catechol | |
|
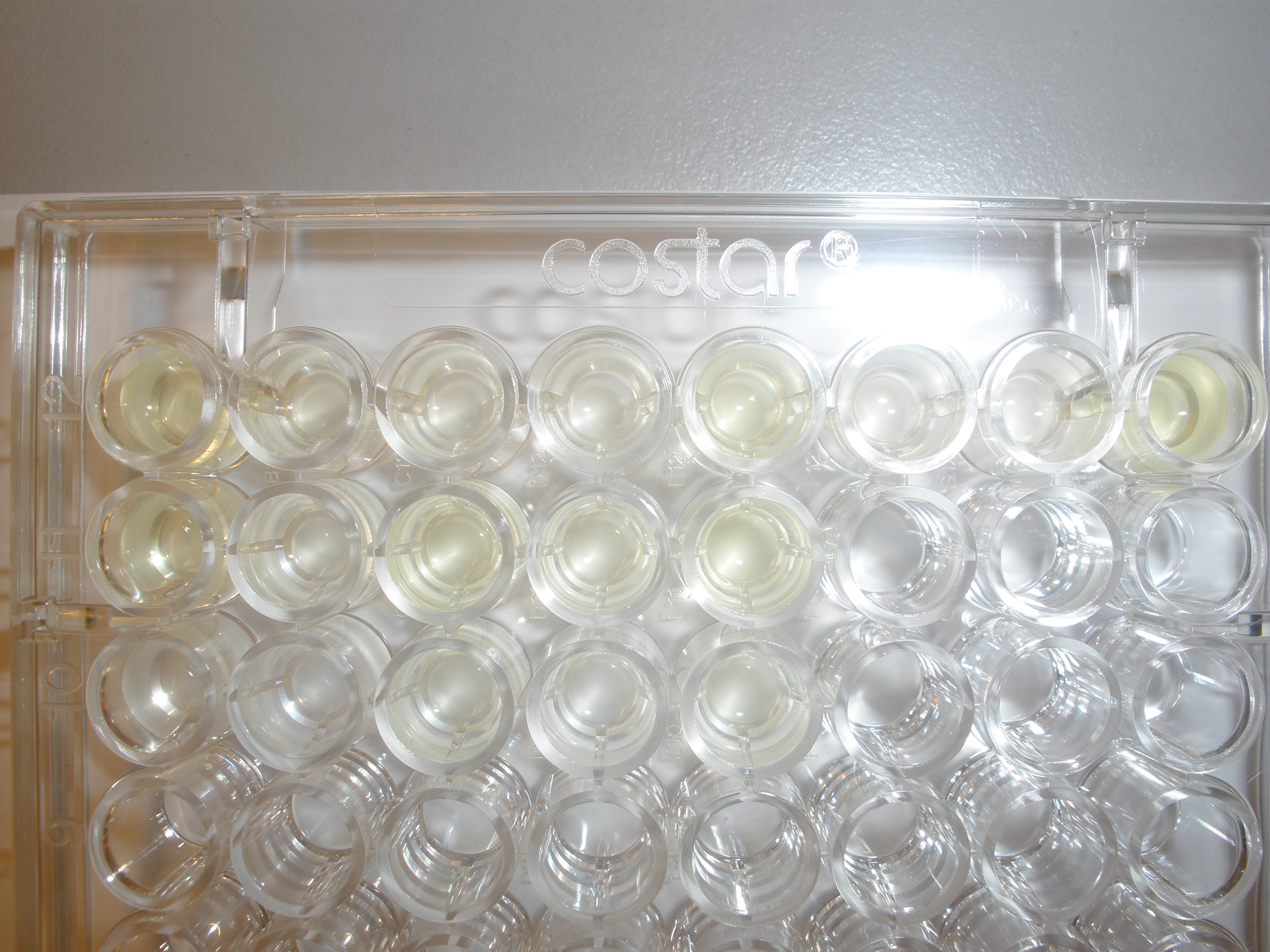
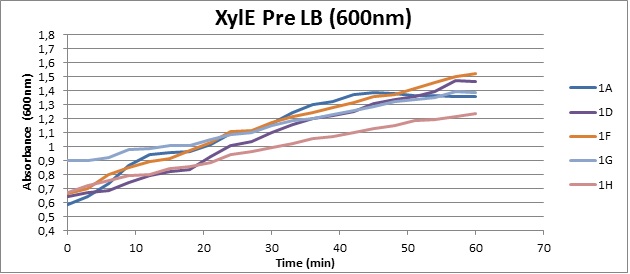
Assaying the growth behaviour of E.coli Top10 cells transformed with either the XylE gene promoted by Pveg or alternatively as control, transformed with a CMR vector of identical size, we were able to determine if and what effect either Catechol itself, or the breakdown product 2-hydroxymuconic semialdehyde had the expected growth-inhibiting effects on the cells. Top10 cells were transformed with XylE and CMR respectively contained on the PSB1C3 vector.Experiments were performed both in LB and M9 medium. The 100ml wells were prepared to display an array of 0 to 2mM [Catechol] exposure. To determine cell-growth behavior we measured absorbance at 600nm using a POLARstar Omega plate-reader set to take measurements every 3 minutes over one hour, without catechol. Catechol was added to all wells after the pre-growth assay was completed and measurements of optical density were taken in 3 minute intervals over 3 hours. Growth without Catechol Pre-growth assays showed growth behavior as expected for cells growing normally. | |
|
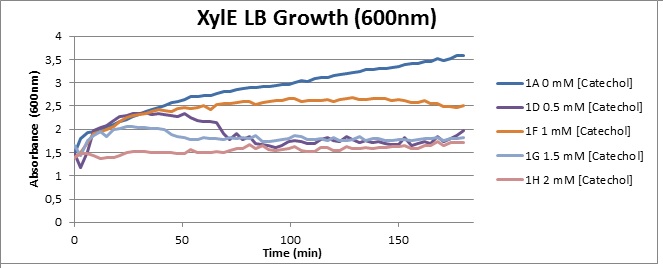
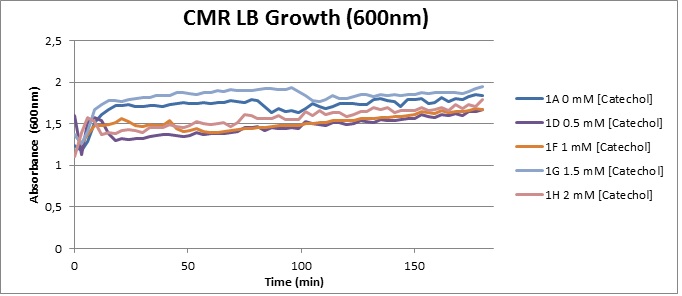
Growth with Catechol (LB) The addition of catechol had distinctive effects on the XylE expressing cells growing in LB medium. While at 0% catechol growth-behavior did not show a significant change (dark blue), even the lowest concentration of 0.25% catechol appeared to drastically reduce cell-survival (red). In contrast, CMR-control cells did not change their growing behavior in the presence of catechol. | |
|
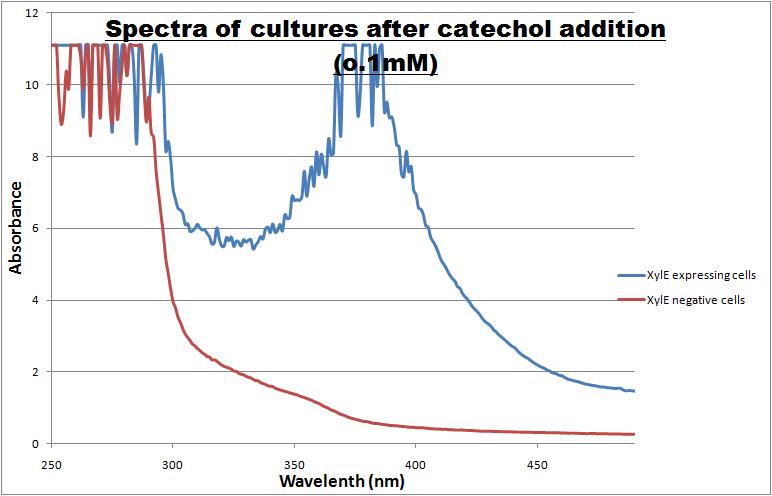
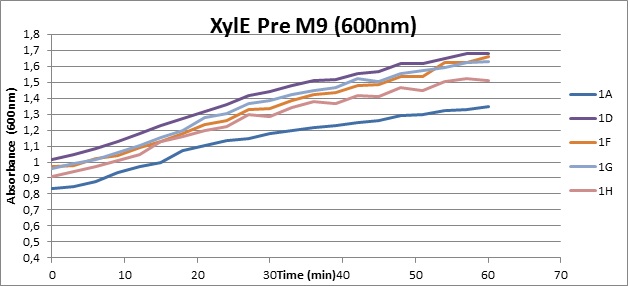
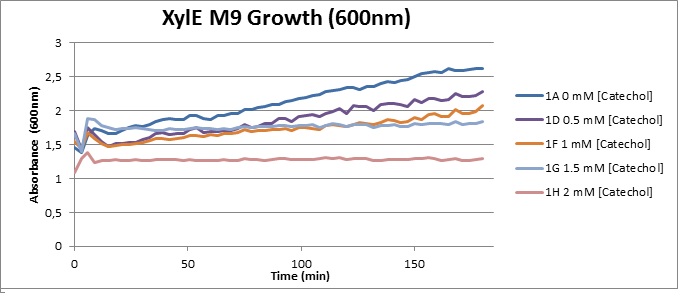
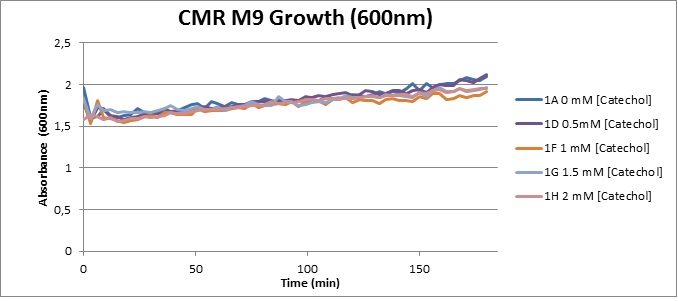
From this we conclude that in LB medium, the breakdown product of catechol, 2-hydroxymuconic semialdehyde, has a lethal effect on E. coli. (M9) Cells growing in M9 medium appeared more resistant to the effects of catechol. Even though absorbance at 380 nm increased significantly in well containing XylE expressing cells, indicating strong turnover of Catechol by C2,3O (4), Catechol did not appear to influence growing behavior in generalizable fashion: while a significant increase in the variance in the sample could be determined. Such was not observed for CMR expressing cells exposed to the same conditions. However, we were not able to establish a clear trend of growing-behavior in the presence of catechol as in case of the LB – XylE samples. | |
|
We conclude that the breakdown product of Catechol has a strong, deleterious effect on XylE expressing cells in an M9 medium, however, cells appear more able to resist the effects of 2-hydroxymuconic semialdehyde in M9 medium. These results support the hypothesis that the breakdown product and not catechol itself has an intense deleterious effect on cell survival. This contributes to the safety of our system, as the output module is inherently coupled to the introduction to a substance lethal to the cells. | |
| Experiment 5 | Characterizing GFP-XylE gene product in whole cell cultures | |
|
As part of the project, the team needed to identify a reporter gene that would give us the visual output. However the means of getting this visual output needed to be inducible, and not constitutively active, when the appropriate input is present. Another specification for this reporter was that it's activation did not to go through the usual route of transcription/translation, which is a slow process. We had little success of finding something suitable for our project so we decided to build our own novel system. We decided to use the gene product of XylE, catechol 2,3 dioxygenase (C2,3O). By looking at three dimensional diagrams of C2,3O and using software tools we proposed an inducible model of this enzyme. | |
|
In our construct, GFP was added to the N-terminus of C2,3O monomer. The C2,3O enzyme is an obligate homotetramer where all 4 subunits are required for its activity. We hoped that by adding GFP at the N-terminus of the each monomer, we would block the tetramerization of the enzyme, thus rendering it inactive. This modification is important as it makes our system inducible. The cleavage of the GFP by TEV protease, which is synthesised on arrival of the input signal, means that Catechol dioxygenase becomes active only when the parasite is detected. | |
|
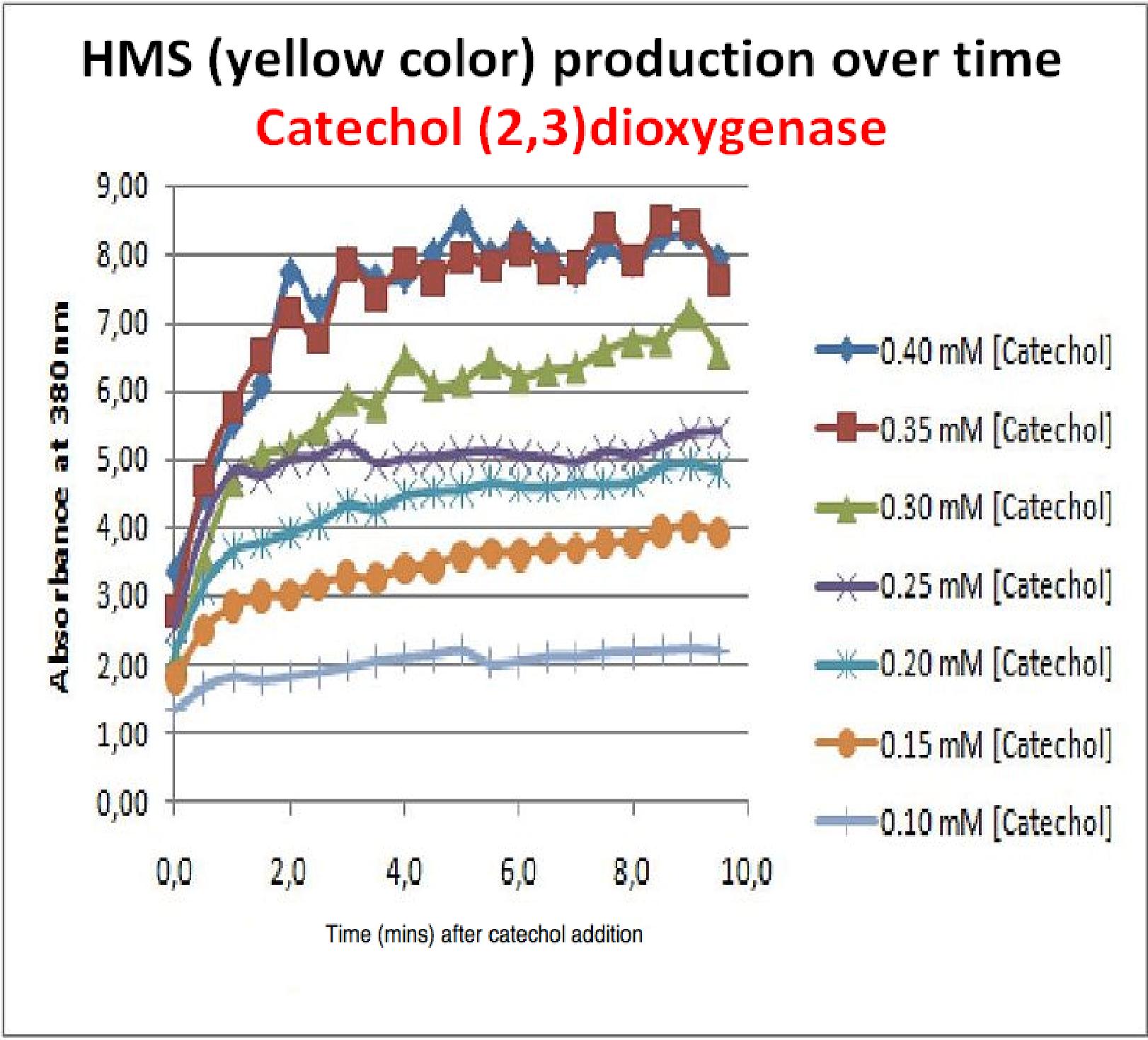
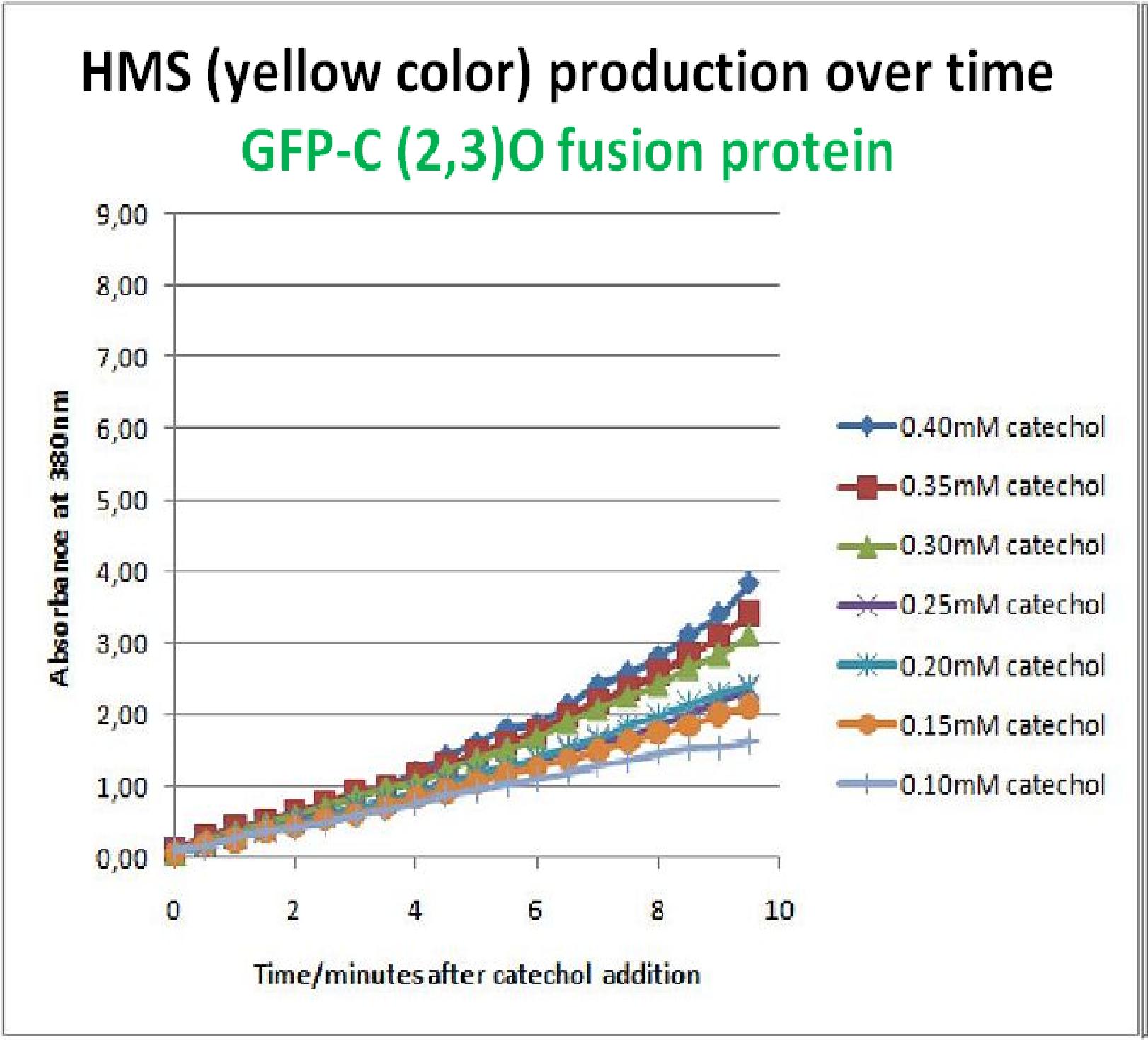
To investigate this, an experiment was designed where catechol was added to GFP-XylE transformed E.coli. The progress of the reaction was monitored by measuring production of the 2-hydroxymuconic semialdehyde (yellow), which arbsorbs light at 380nm. The product concentraton correlates well to the velocity of the reaction, if divided by time, and by using various initial catechol concentrations as a substrate, a kinetic profile of the reaction could be constructed. The experimental set up is essentially the same as described above for determining the kinetics of C2,3O. The only difference is that the E.coli cells used were the ones transformed with GFP-XylE gene construct instead of XylE gene. The results of the assay show conclusively that the N-terminal fusion of GFP gene to the XylE gene affects activity of the wild type C2,3O. Although the activity is leaky, producing very small amounts of the colored product of the reaction in the presence of catechol, the alteration in the reaction profile of the reaction is impressively large. The two graphs below are equivalent and comparable in the sense that the only different between these experiments is the E.coli cells used. The left graph represents cells expressing the original XylE gene, while the graph on the right hand side represents cultures expressing the GFP-XylE gene. These genes are both under the influence of the Pveg promoter so that we can assume equal production of protein product. | |
|
If we take 3 minutes as the reference point on the graphs, as this is where the XylE expressing cells saturate the system with the colored product, we can see that there is about a 10-fold reduction in the rate at which the protein product of GFP-XylE gene construct catalyses the reaction in comparison to the wild type XylE gene. From this we can deduce that the attached GFP protein influences tetramerisation of the Catechol dioxygenase enzyme, so that the equilibrium: 4 monomers (inactive) <-----> tetramer (active) is shifted is shifted towards protein remaining as monomers and thus inactive. The results of this experiment were really encouraging for the team, as it shows that we have managed to inactivate the normally constitutively active reporter protein of our output module. The lab team now proposes an experiment where the inactive fusion protein (GFP-catechol dioxygenase) is exposed to TEV protease. Reversion of the reaction profile back to the kinetics of the wild type catechol dioxygenase would mean that we have successfully constructed an inducible reporter system. | |
| Experiment 6 | In vitro characterization of C(2,3)O in cell lysate | |
|
In a previous experiment investigating cultures of XylE expressing E.coli, the addition of catechol and monitoring the production of HMS product in the plate reader allowed us to obtain a reaction profile of the reaction. However due to technical limitations on measuring velocity of the reaction in whole cell cultures, we were not able to obtain kinetic parameters of the catechol dioxygenase enzyme. In this experiment cell cultures of XylE expressing cells were lysed to obtain cell lysate with catechol dioxygenase activity for determination of kinetic parameters of catechol dioxygenase in vitro. | |
|
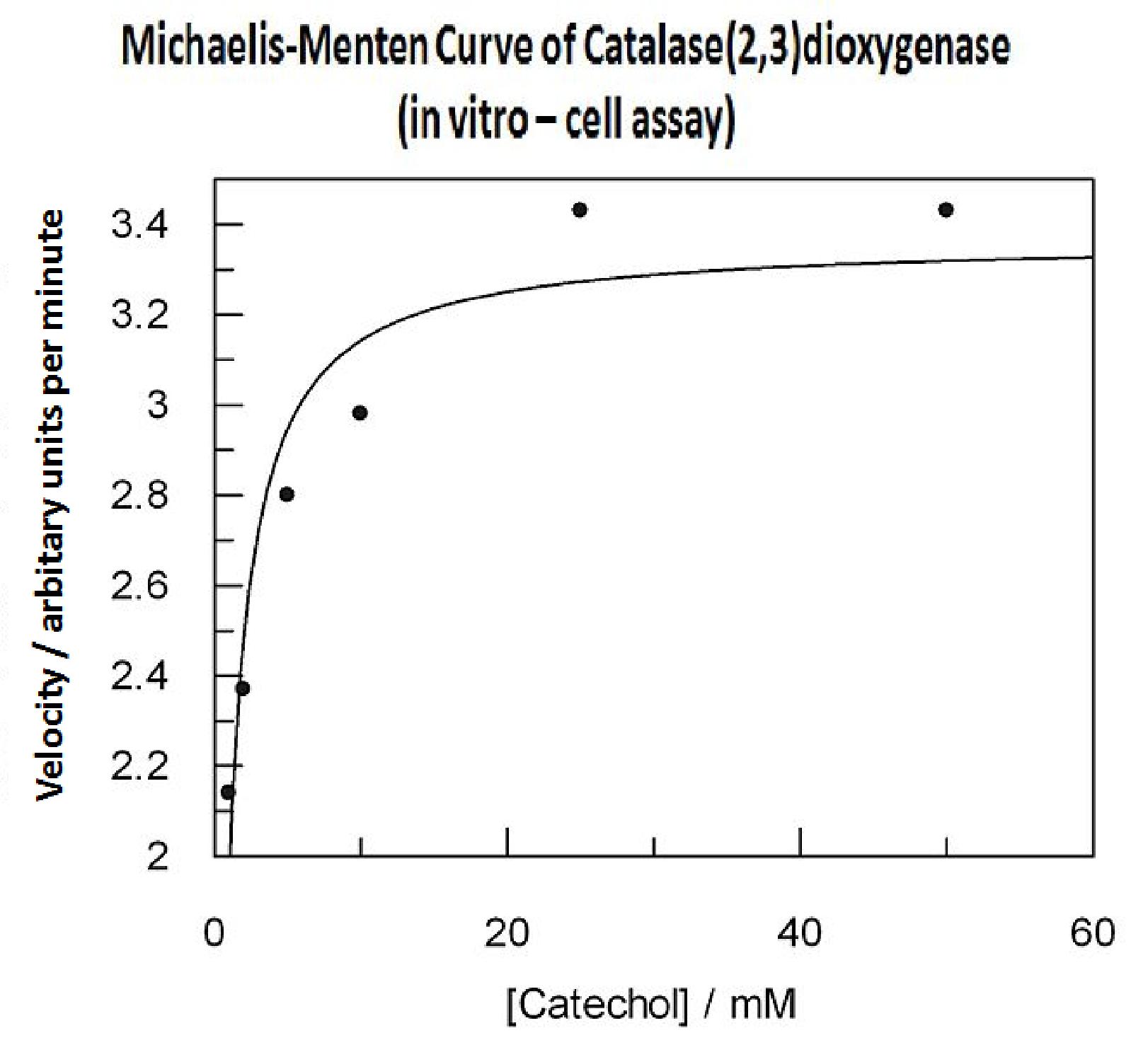
In this experiment cell lysate was assayed with increasing catechol concentrations. The reaction was monitored by measuring color output of the reaction in the plate reader. The rate at which the yellow HMS product appears is directl proportional to the velocity of the reaction. The cell lysate was prepared from cell culures of XylE expressing cells. Obtaining cell lysate with catechol assay activity allowed us to overcome the technical limitations of over saturating the system too quickly in previous experiment. Diluting the cell lysate will result in lower concentration of total enzyme in the assay, thus less substrate is converted to product per unit of time. This allowed us to take more readings at various time points before the system saturates the detector of the reader. In order to determine what the appropriate dilutions of the cell lysate for the assay, a pre-test was essential where various cell lysate dilutions were tested on their dioxygenase activity in respect to the ability of the detector to receive the signal. Data collected from the assay allowed us to construct a Michaelis-Menten curve for the in vitro kinetics of C2,3O in cell lysate. The cell lysate was obtained from a 100ml overnight culture and the lysate was diluted by a factor of 20-fold in order to obtain a suitable concentration of total enzyme for the plate reader assay. The concentrations of catechol used to derive Michaelis-Menten curve were 1, 2, 5, 10, 25, 50 mM, prepared from 1M stock catechol. | |
|
Michaelis-Menten curve was drawn using velocity values calculated from data collected from the catechol assay. The velocity values were calculated from the rate slope at the initial stages of the reaction, as this is the only time when substrate concentration values are accurate. The plot was delineated by non-linear regression analysis using GraFit software tool. The calculated Km is 0.71mM catechol (with a Vmax of 3.37 in O.D. arbitrary units for this dilution of cell lysate). | |
 "
"