Team:Washington/Gram Positive/Test
From 2010.igem.org
(→Test: Enzyme Assay) |
(→Protein Gel) |
||
| Line 38: | Line 38: | ||
[[Image:Washington_Confirming_CP_activity_revised3.jpg|thumb|750px|left|Figure 1: Confirming the activity of CapD_CP by comparing it to two CapD_CP knockouts, T2A and T2V.]] | [[Image:Washington_Confirming_CP_activity_revised3.jpg|thumb|750px|left|Figure 1: Confirming the activity of CapD_CP by comparing it to two CapD_CP knockouts, T2A and T2V.]] | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
=='''Continuing with CapD_CP'''== | =='''Continuing with CapD_CP'''== | ||
Revision as of 23:32, 27 September 2010
More descriptive titles, verification that your CapD_CP was a monomer ( gel) should go before testing catalytic ability
Contents |
Test: Enzyme Assay
After we have the CapD_CP mutants, we tested our mutants for their catalytic activity using our fluorescence-based enzyme assay scheme. Fluorescence-based enzyme assay measures the rate at which fluorescence in the testing media is released and the amount fluorescence depends on the rate at which fluorophore-quencher linkage is disrupted. Our substrate PDGA contains a linked fluorophore-quencher component. The faster fluorophore-quencher component is cleaved, the higher the amount of fluorescence is released and thus the greater the enzymatic activity observed.

Result: Data Analysis
CapD_CP Activity Validation
Before we predict which mutations increase hydrolysis capability, we need to validate that the circularly permuted version of CapD has measurable activity for further assessments. We also hypothesize a threonine residue in the catalytic site of CapD_CP plays an important role in the catalysis reaction and mutating it will eliminate all enzymatic activity. Thus we created two mutants, T2V and T2A, to act as negative controls. The result (figure 1 below) of this assay confirms our hypothesis that CapD_CP has enzymatic activity to the two catalytic knockouts. The relatively flat activity curves of the knockout mutants confirm the hypothesis of the threonine's role in the catalytic site.
Continuing with CapD_CP
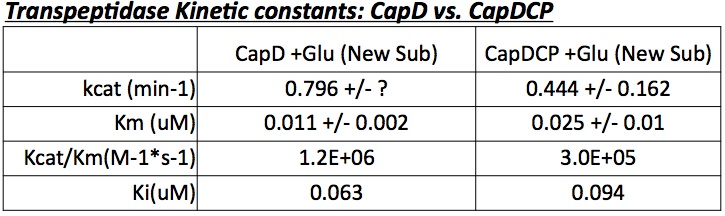
Based on the gel's results, CapD_CP was used for future mutant designs. The reason is two-fold. First, CapD_CP is easier to quantify due to its single band characteristic. Second, testing with the three unequal and ambiguous bands in CapD would prove problematic since determining which band to measure for active enzyme is nearly impossible. After validating the activity of CapD_CP, more analysis was done pertaining enzymatic properties. Compiling a Michaelis-Menten profile, the data provides properties of Kcat, Km, Kcat/Km and Ki (see figure 3) for the CapD and CapD_CP enzymes. Based on the gel result and the kinetic properties, evidence shows the two versions are very similar, thus all mutants will be done in CapD_CP.
Analyzing CapD_CP
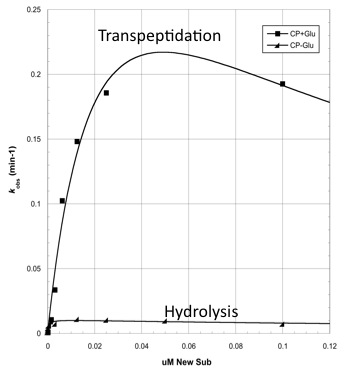
The two abilities of CapD_CP are transpeptidation and hydrolysis. Based on the Kcat and Km (see figure 4) values of the two, we conclude that CapD_CP is a weak binder and efficient catalyst for the transpeptidation reaction. In terms of hydrolysis, it shows strong binding but slow catalysis, thus our mutant designs focus on increasing the catalytic efficiency.
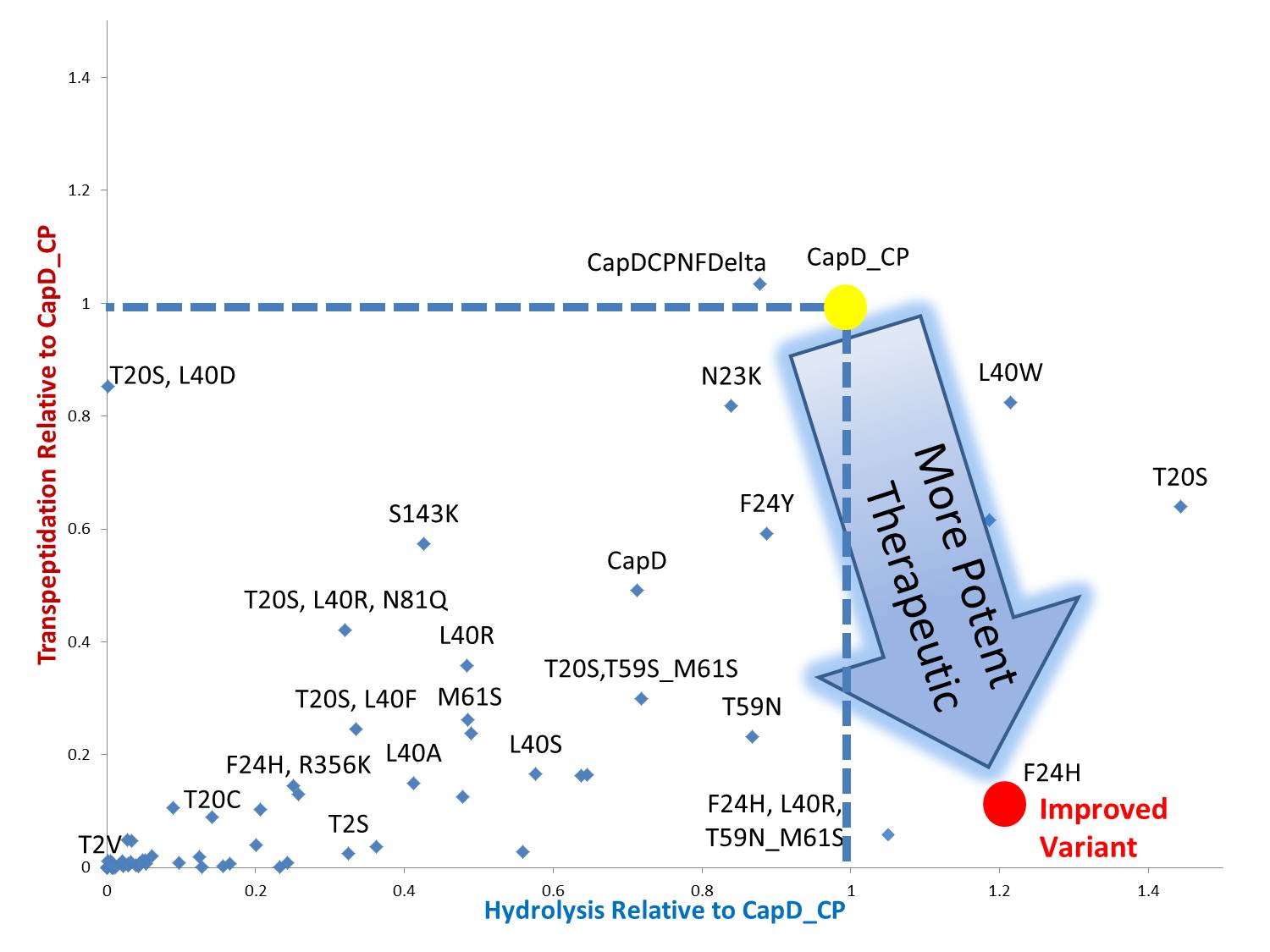
Mutant Designs
By standardizing the activity slope of each design relative to CapD_CP, a scatterplot easily portrays the qualities of each mutant. Several designs show negative catalytic curves similar to the catalytic knockouts. Some immediately show a negative activity curve meaning decrease in transpeptidation, hydrolysis, or both. T20S is a promising mutant hydrolase design.
 "
"