Team:Washington/Tools Created/New Vectors
From 2010.igem.org
Protein Expression Vectors
Vector Design
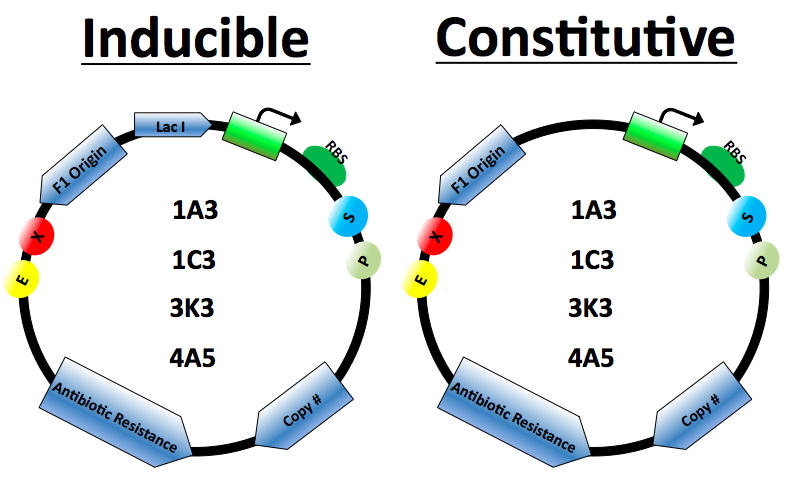
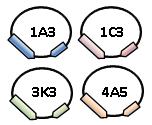
The protein expression cassette is designed to go into any biobrick plasmid backbone. It was placed in 4 different plasmid backbone from the registry pSB1C3, pSB1A3, pSB3K3,and pSB4A5 by our lab.
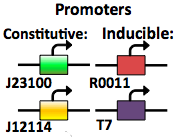
The expression vector promoters are available in constitutive and inducible variety. The Constitutive promoters are BBa J23100 and J23114. Inducible vectors include R0011 and T7, both of which are repressed by the Lac I protein.
Lac I gene inserted to ensure regulation of the Lactose inducible promoters
Antibiotics resistance based on biobrick plasmid backbone used.
The Elowitz standard RBS B0034 is used on all vectors.
Origin of replication for the M13 series of phages. When included in plasmids the cells can be infected by M13 phage which will then replicate and package the plasmid as single stranded DNA. This DNA can then be isolated as described in our Kunkel Mutagenesis protocol.
Plasmid copy number based on biobrick plasmid backbone used.
Restriction cut sites based on biobrick standards.
Building the Vectors
The expression cassettes were built using the biobricks construction tutorial.
Testing the Vectors
Protein Expression
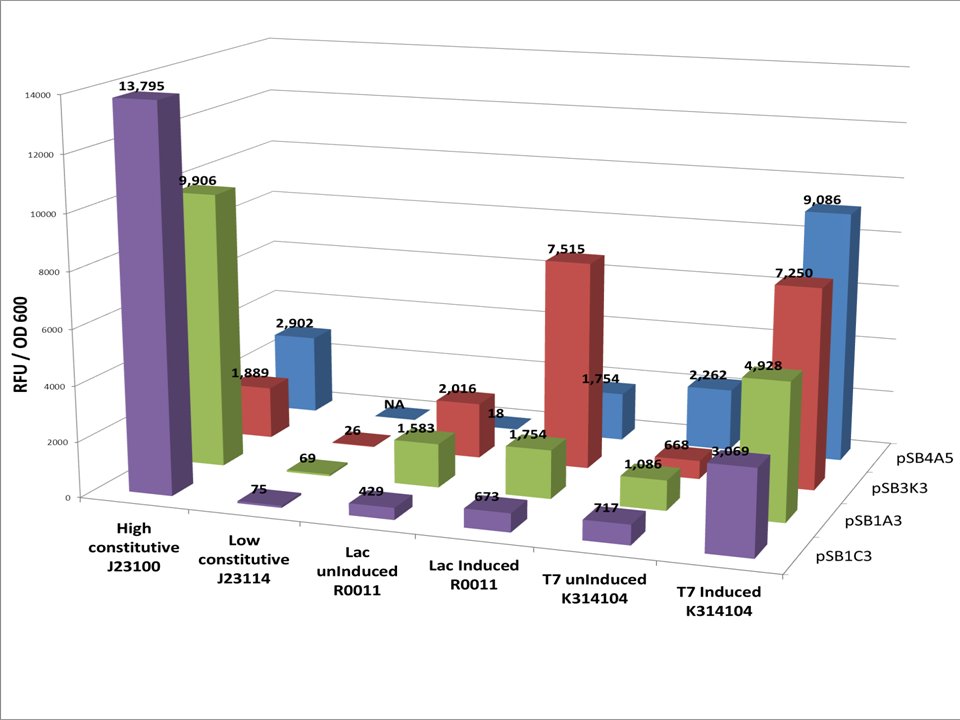
The vector cassettes was placed in 4 different plasmid backbones from the registry pSB1C3, pSB1A3, pSB3K3, pSB4A5 and GFP was placed in the protein expression area of the vector. The expression cassette was grown according to our vector assay protocol. Data was corrected for background florescence.
f1 origin
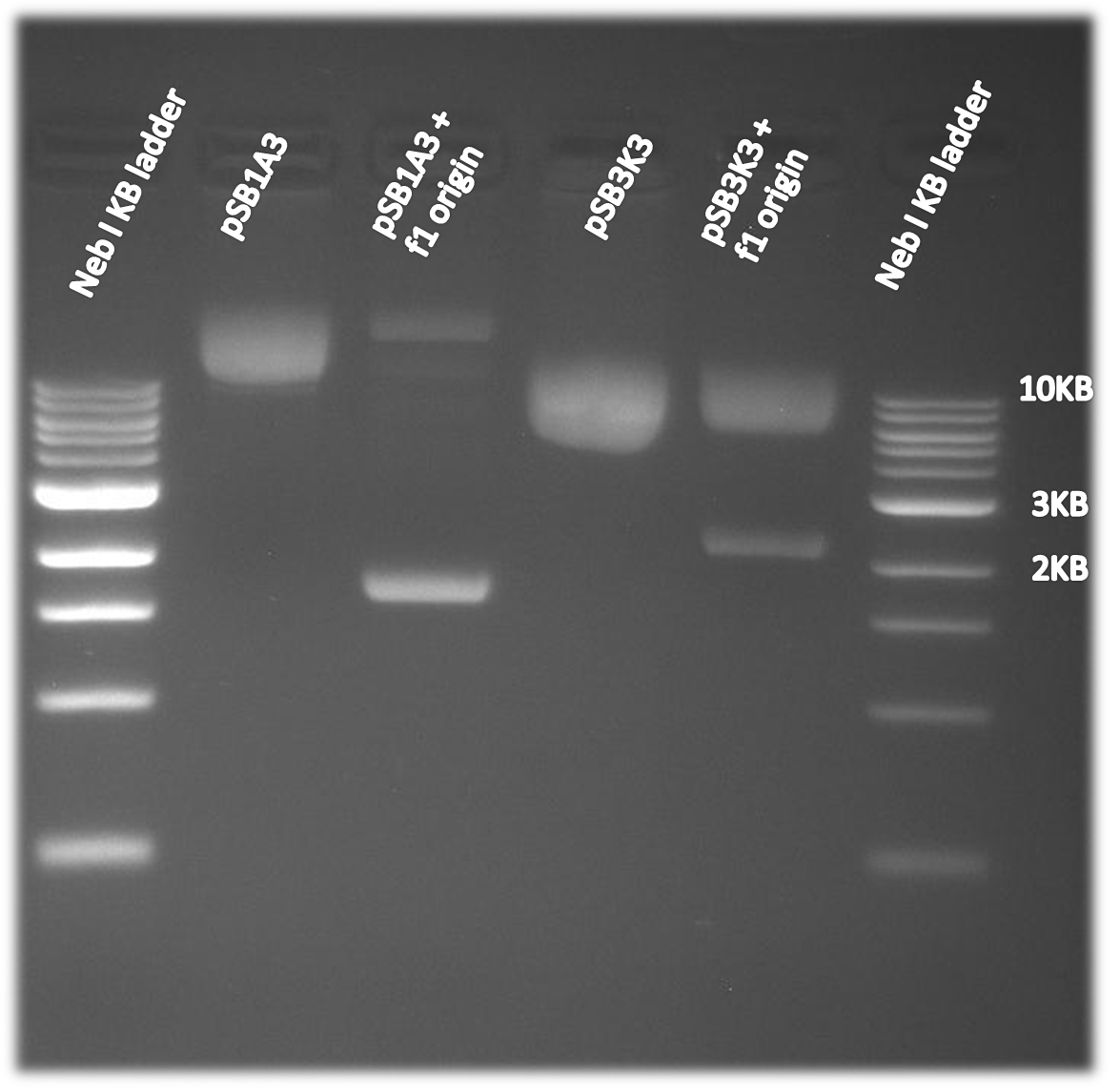
The f1 origin was tested by comparing single strand DNA harvest using part of the Kunkel Mutagenesis. CJ236 cells were infected with M13K07 helper phage. The CJ236 cells varied in the present or absence of the f1 origin on the pSB1A3 or pSB3K3 plasmid. The single stranded DNA was harvested and then run on a 1% agarose gel. As is clearly visible on the gel below single strand DNA of the plasmid (~2-3kb, depending on the plasmid) is only made when the f1 origin is present. Note that single stranded DNA runs at a slightly smaller than expected size when comparing to a double stranded DNA ladder.
 "
"