Team:Washington/Gram Positive/Design
From 2010.igem.org
(→Using FoldIt to Make a Better Hydrolase) |
|||
| Line 1: | Line 1: | ||
| - | + | ||
{{Template:Team:Washington/Templates/Header}} | {{Template:Team:Washington/Templates/Header}} | ||
<html> | <html> | ||
Revision as of 00:06, 14 September 2010
Making CapD a Better Anthrax Treatment
There are two main obstacles limiting natural CapD as an Anthrax therapeutic. First, natural CapD is a difficult to express dimer, requiring an auto-cleavage to become active. Second, CapD is a better transpeptidase than poly-γ-D-glutamate hydrolase, limiting its decapsulating potential. To solve the first problem, we created a circular permuted, monomeric version of CapD that is easy to express. To improve hydrolosys, we used FoldIt, a computational toolbox, to design active sight mutations aimed to increase hydrolosis over transpeptidation.
Circular Permutation of CapD
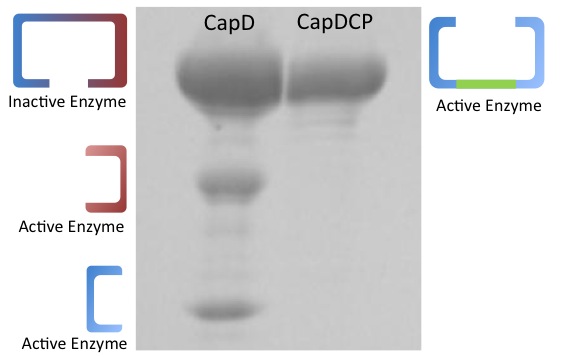
When natural CapD is first translated, the key catalytic threonine residue is buried in the active site, inaccessible to poly-γ-D-glutamate. After auto-cleavage, this critical threonine becomes the new N terminus, which can take its place in the active site. By reordering the protein so the threonine is the first residue, and putting a FoldIt designed linker between the natural N and C terminus, we make a circular permutation of CapD that we named CapD_CP. CapD_CP is a monomer, historically easier to purify and more stable than dimers.
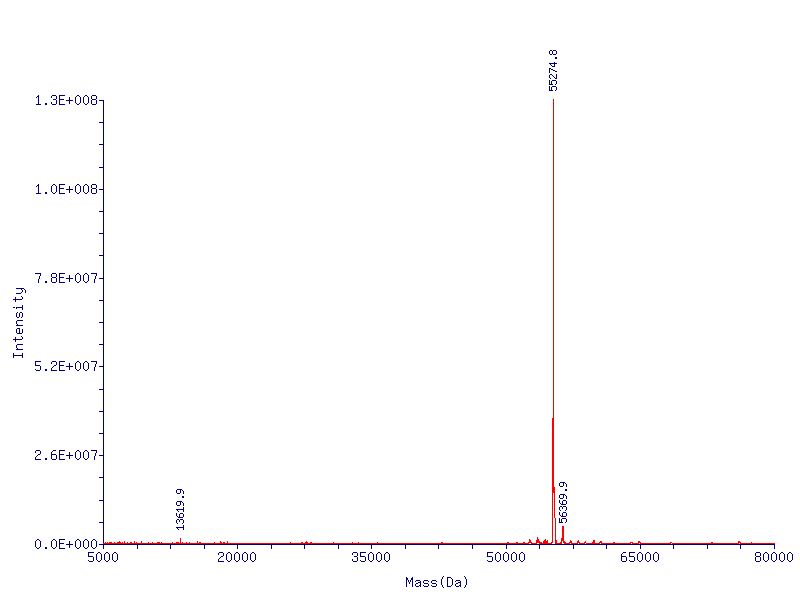
Since the first residue of any nascent protein must be methionine, we rely on E. Coli’s naturally occurring methionine aminopeptidase to remove the first methionine, making CapD_CP catalytically active. The removal of the first methionine has been verified via mass spectrometry.
Using FoldIt to Make a Better Hydrolase
To increase the hydrolytic ability of CapD_CP, we made point mutations in the active site. We focused our attention on two types of mutations. First, we created point mutations that hydrogen bond to a modeled transition state of our substrate in an attempt to stabilize the transition state. Second, we mutated the active site into a more open and polar area in an attempt to increase the ease with which water can enter and participate in a hydrolysis reaction.
To make these point mutations, we used a program called FoldIt to predict how changes in protein structure and composition will affect protein stability. FoldIt provides a 3D representation of a protein's crystal structure which can be manipulated. Manipulation functions include point mutations, insertions, deletions, repacking of side chains, and backbone movement, which FoldIt then assesses for stability. This allows the user to quickly interact with a protein and easily predict how mutations will affect a protein.
 "
"