Team:Washington/Gram Positive/Build
From 2010.igem.org
(→Annealing to ssDNA) |
(→Order Oligonucleotides) |
||
| Line 32: | Line 32: | ||
==='''Order Oligonucleotides'''=== | ==='''Order Oligonucleotides'''=== | ||
| - | To mutate our wild-type gene, we used [https://2010.igem.org/Team:Washington/Protocols/Kunkel Kunkel's mutagenesis protocol]. Kunkel’s is a site-directed mutagenesis, requiring knowledge of wild-type sequences. After the desired mutation is modeled using [ | + | To mutate our wild-type gene, we used [https://2010.igem.org/Team:Washington/Protocols/Kunkel Kunkel's mutagenesis protocol]. Kunkel’s is a site-directed mutagenesis, requiring knowledge of wild-type sequences. After the desired mutation is modeled using [https://2010.igem.org/Team:Washington/Project/Tools/FoldIt/ FoldIt], we order a mutation's antisense oligonucleotides from [http://www.idtdna.com/ Integrated DNA Technologies]. Oligonucleotides are short segments of nucleotide primers which contain one or more mutations and will anneal to single-stranded DNA (ssDNA) of a plasmid containing the CapD expression gene. The result should be a double stranded plasmid which holds the [https://2010.igem.org/Team:Washington/Project/Tools/FoldIt/ FoldIt]-designed mutation. |
==='''Generate ssDNA'''=== | ==='''Generate ssDNA'''=== | ||
Revision as of 18:56, 17 September 2010
Contents |
Build (Gram Positive)
Mutate DNA
Order Oligonucleotides
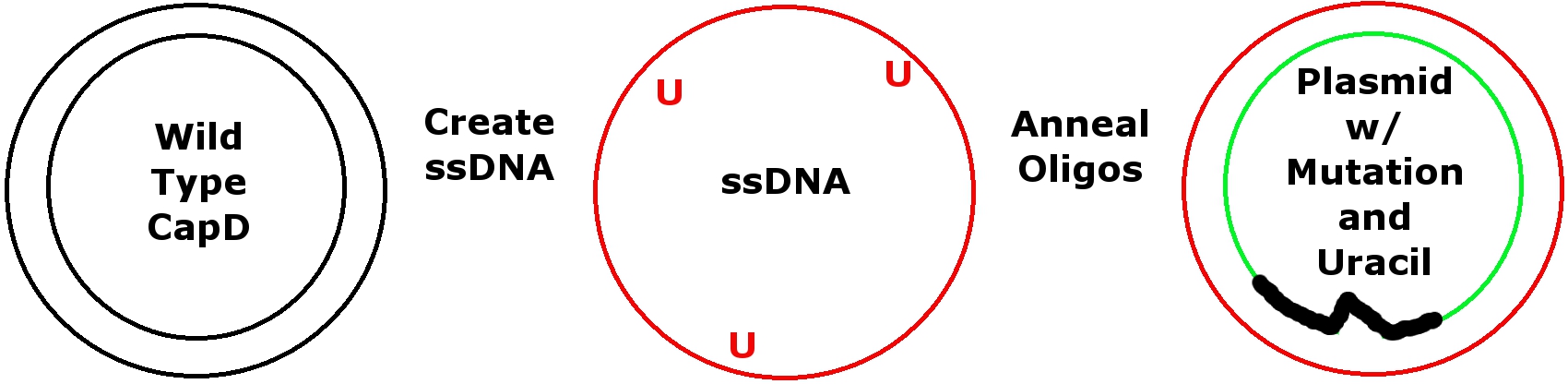
To mutate our wild-type gene, we used Kunkel's mutagenesis protocol. Kunkel’s is a site-directed mutagenesis, requiring knowledge of wild-type sequences. After the desired mutation is modeled using FoldIt, we order a mutation's antisense oligonucleotides from [http://www.idtdna.com/ Integrated DNA Technologies]. Oligonucleotides are short segments of nucleotide primers which contain one or more mutations and will anneal to single-stranded DNA (ssDNA) of a plasmid containing the CapD expression gene. The result should be a double stranded plasmid which holds the FoldIt-designed mutation.
Generate ssDNA
In order to anneal oligonucleotides, CapD expression gene ssDNA must be obtained by transforming CJ236 cells with a plasmid containing the CapD gene. These cells use Uracil instead of Thymine, and are unable to destroy Uracil containing DNA. Colonies are then picked and M13K07 helper phage is introduced. The phage will use the cells to reproduce and copy the plasmid containing the CapD expression gene to daughter phages. The phage will produce one strand of DNA using reverse transcriptase, creating ssDNA. We use the Miniprep protocol to harvest the ssDNA from the phage.
Annealing to ssDNA
Received oligonucleotides lack activity-inducing phosphates. Adding phosphates by kinasing readies them for annealing to ssDNA. The oligonucleotide binds to a specified location on the ssDNA except where the mutation is. This creates an area where the oligonucleotide will not bond to the template ssDNA.
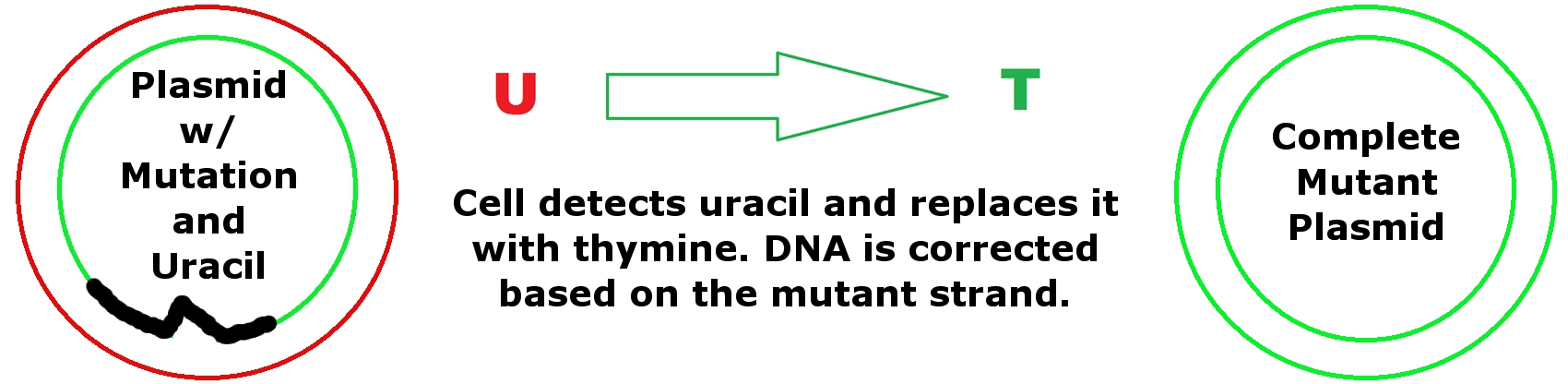
Synthesize the Plasmid
Using DNA polymerase, the rest of the missing strand is synthesized. Finally the area in the plasmid where the mutation is fixed because the former ssDNA strand that contains uracil is destroyed by the cell we transform our mutant plasmid into. Since the cell uses the other strand (the strand with the Oligonucleotides) to fix the DNA, the result is a complete mutant plasmid. Plasmids are then sent for sequencing by [http://www.genewiz.com/ GENEWIZ] to confirm mutations.
Grow Protein
Transform E. coli with mutant plasmid
E. coli is transformed with our mutant plasmid. This completes mutant plasmid because these cells are able to remove uracil and replace it with thymine based on the complementary strand (which contains the mutation).
Grow cells
Inoculated E. coli is grown in terrific broth (TB) until 600nm optical density reaches desired range.
Protein Production
By introducing Isopropyl β-D-1-thiogalactopyranoside (IPTG), an allolactose mimic, we induce E. Coli to produce our protein. IPTG binds with the lac inhibitor protein and activates the lac operon, turning on the CapD gene and causing production of our mutant protein.
Harvest Protein
Spin down cells
Using a centrifuge, cells and media are spun to separate cells from media.
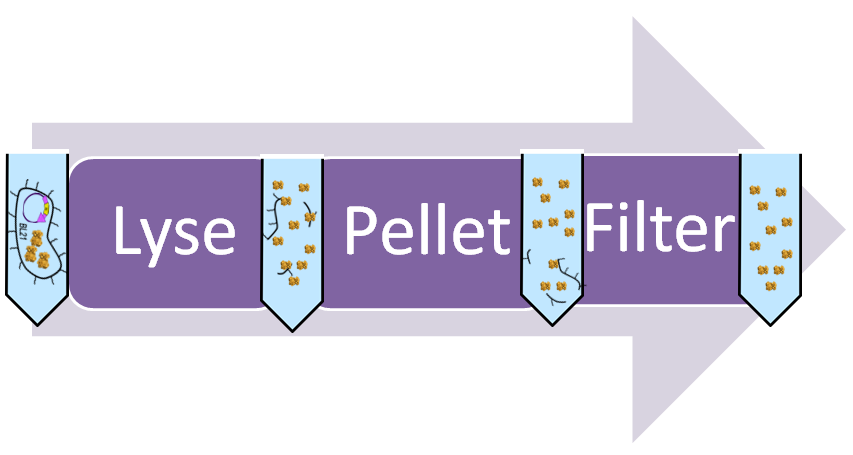
Lyse cells
The supernatant (media) is emptied and the cells at the bottom are lysed open. The result is a slurry containing all the cell’s proteins and DNA. Lysis is then spun down and the supernatant, containing all the proteins, is collected. Among the proteins is CapD.
Purify proteins
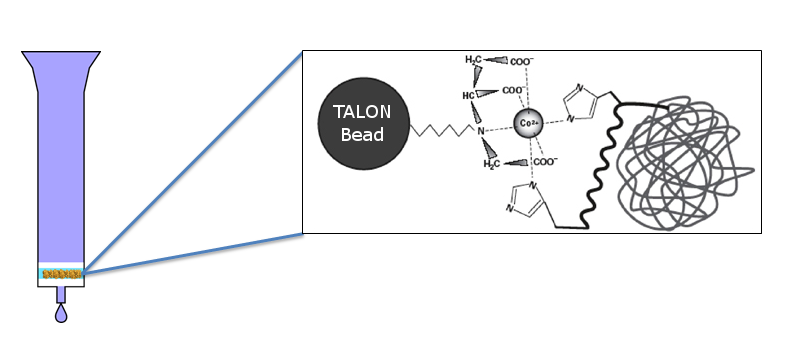
To purify the protein, we run the supernatant collected from lysis through a column containg TALON resin cobalt beads. CapD's designed histidine tags bind to the beads, whilst everything else flows through. Finally, the CapD is eluted with imidazole, a histidine without a backbone, which outcompetes the affinity to bind. The result is our purified mutant CapD.
 "
"