Team:Peking/Project/Bioabsorbent/MBPExpression
From 2010.igem.org

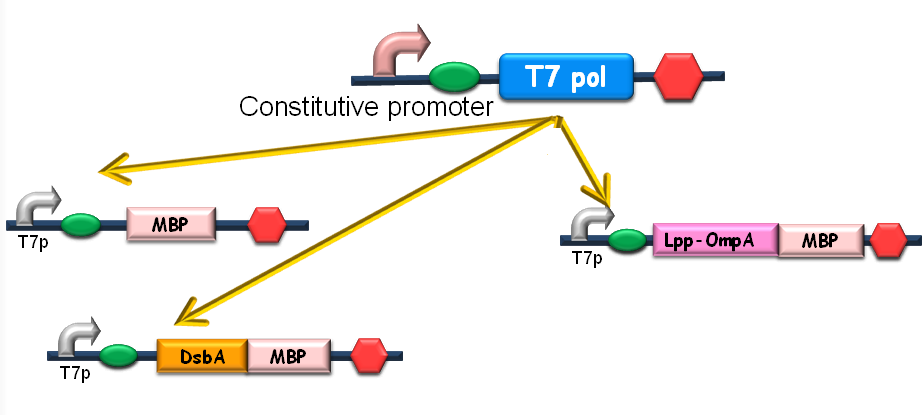
Since our ultimate goal is to design a high-performance and less energy-consuming bioabsorbent, the MBP will be an excellent candidate for the absorbent effector. MBP was then fused with DsbA, a periplasmic translocated protein with a signal sequence at its N-terminal, and OmpA, a membrane protein, to construct periplasmic MBP and surface displayed MBP, in order to maximize the bacterial capability of mercury binding. Finally, mercury MBP was constitutively expressed on surface, periplasm and cytosol of E.coli cells via carefully designed genetic circuits, to guarantee the maximum of Hg absorption (Fig 1).
Figure1. The genetic circuit we designed to guarantee the maximum of Hg absorption. The production of T7 RNA polymerase is constitutive. T7 polymerases will active high rating transcription at T7 promoters. Thus Hg (II) will be highly effectively accumulated by substantial amount of MBPs which are translocated to cytosol, periplasm and cell surface of the bacteria.
Cytoplasmic Expression of MBP
To design the module of cytosol expression, MBP in pSB3K3 was then to coloned into pSB1A2, prefixed by T7 promoter and RBS BBa_B0034 (Fig 2), verified by DNA sequencing.

Fig 2. Construction of the cytosol expression module. MBP was cloned into the pSB1A2 step by step, prefixed by T7 promoter and RBS.

Figure 2. Scheme of MBP construction and the predicted MBP structure. (A) Linear structure of MerR coding region. (B) Construction of MBP. The MBP was constructed by fusing two copies of metal binding domain of MerR in tandem with a flexible SSG bridge. Blue bars indicate the dimerization helix in the metal binding domain of MerR, and gray bars indicate other α-helices of MerR. The red line indicates the SSG linker, and the blue and green lines indicate the loop after the dimerization loop and the region after the loop, respectively. Orange dots indicate cysteines involved in Hg(II) binding[3]. (C) Predicted structure of resulted metal binding peptide. Mercury ions are indicated as black balls in metal binding pockets.
Earlier work provides us a deeper insight into the metal recognition mechanism of MerR family. Kathryn et al. constructed a hybrid regulatory protein containing the DNA binding domain of MerR and the Metal binding domain of ZntR, which showed an altered specificity to MerR’s promoter but responding to zinc[4], which suggested that the DNA binding domain and metal binding domain of MerR family function modularly to some extent. Since our bioabsorbent needs high metal binding affinity and selectivity, we then took a closer look at the metal binding mechanism and the structure of the C-terminal binding domain.
MerR family TFs share a high similarity at the C-terminal metal binding domain (Fig. 3), which indicates a similar metal recognition mechanism and metal-protein complex structure. The key factor of the remarkable selectivity and sensitivity seems to be the preorganization of geometries suited for specific metal ions through folding of the metal-binding domains in these proteins [5]. To be specific, previous mutagenesis study shows that MerR dimer binds one Hg(II) ion in a bridge fashion between the two monomers[6], and Hg (II) adopts a three-coordinate Hg-(S-Cys)3-binding mode by extended X-ray absorption fine structure (EXAFS) spectroscopy[7], though there is no crystal structure available. As is known, single α-helix has a high tendency to be oligomerized. The outer electron configuration of Hg (II) is 5d106s0, and it tends to form a complex, especially with sulfur. All these evidences above indicate that the C-terminal metal binding domain can act as a mercury accumulator without the help of the N-terminal DNA binding domain. Luckily, Qiandong Zeng et al. constructed an N-terminal deletion mutant (contain only residue 80-120) that can form a stable dimer and retain high affinity for Hg (II) [8]. This work reinforces the idea of tandeming two metal binding domains together to make a high performance and less energy consuming metal binding peptide.
Figure 3. Structure based sequence similarity of various metal responsive regulators of MerR family. CadR: cadmium, CueR: copper, ZntR: zinc, MerR: mercury, PbrR:lead. Those amino acids showed in bold are metal binding sites according to solved crystal structures [2] or mutagenesis analysis [6]. The metal binding site of CadR or PbrR can be speculated based on the high similarity of MerR family and the geometry of ligand field of metal ions.
METAL BINDING PEPTIDE (MBP) CONSTRUCTION
To achieve the goal of making a high performance MBP, we constructed a single polypeptide consisting of two dimerization helixes and metal binding loops of MerR, to form an antiparallel coiled coil MBP mimicking the dimerized metal binding domains of the wild-type as described in Fig 2. We amplified the N-terminal and C-terminal of MBP directly from full length MerR by PCR, and then cloned them into the backbone together in one step (Fig. 4).
A
B
Figure 3. MBP Construction procedure. A: Standard part. B. Expression detection part.
To be specific, the entire coding region of the MBP for standard part was amplified by PCR from full length MerR with two pairs of primers. Two of these primers encoded a three-residue bridge, SSG, which does not occur in MerR and was added to afford some flexibility in the loop connecting the two dimerization helix (fig. 1). The two PCR products were digested with EcoR I / BamH I, or BamH I / Pst I and cloned into EcoR I / Pst I -digested pSB1K3 in one step (fig. 3A), which was verified by DNA sequencing.
Based on the same strategy, MBP-His6 was constructed by using two different pairs of primers, which is used for MBP expression test by western blot. The two PCR products were digested with Nde I / BamH I, or BamH I / Xho I and cloned into Nde I / Xho I -digested pET 21a, which contains a region encoding six histines, in one step to construct pET 21a – MBP (fig. 3B), which was verified by DNA sequencing.
Reference
1.Chen, S., and D. B. Wilson. 1997. Construction and characterization of Escherichia coli genetically engineered for bioremediation of Hg2-contaminated environments. Appl. Environ. Microbiol. 63:2442–2445.
2.Chen, S., and D. B. Wilson. 1997. Genetic engineering of bacteria and their potential for Hg2 bioremediation. Biodegradation 8:97–103.
3.Chen, S., and D. B. Wilson. 1997. Construction and characterization of Escherichia coli genetically engineered for bioremediation of Hg2-contaminated environments. Appl. Environ. Microbiol. 63:2442–2445.
4.Luirink, J., and B. Dobberstein. 1994. Mammalian and Escherichia coli signal recognition particles. Mol. Microbiol. 11:9–13.
5.Debarbieux, L., and J. Beckwith. 1998. The reductive enzyme thioredoxin acts as an oxidant when it is exported to the Escherichia coli periplasm. Proc.Natl. Acad. Sci. USA. 95:10751–10756.
6.Francisco, J. A., Earhart, C. F. & Georgiou, G. (1992). Transport and anchoring of beta-lactamase to the external surface of Escherichia coli. Proc Natl Acad Sci U S A 89, 2713–2717.
7.Francisco, J. A., Campbell, R., Iverson, B. L. & Georgiou, G. (1993). Production and fluorescence-activated cell sorting of Escherichia coli expressing a function antibody fragment on the external surface. Proc Natl Acad Sci U S A 90, 10444–10448
8.Yamaguchi, K., Yu, F. & Inouye, M. (1988) Cell 53, 423-432.
9.Jie Qin,Lingyun Song,Hassan Brim, Michael J. Daly and Anne O. Summers(2006) Hg(II) sequestration and protection by the MerR metal-binding domain(MBD).Microbiology 15, 709–719
 "
"