Team:Monash Australia/Modelling
From 2010.igem.org
(Difference between revisions)
(→Metabolic modelling of the ethylene generator) |
|||
| Line 6: | Line 6: | ||
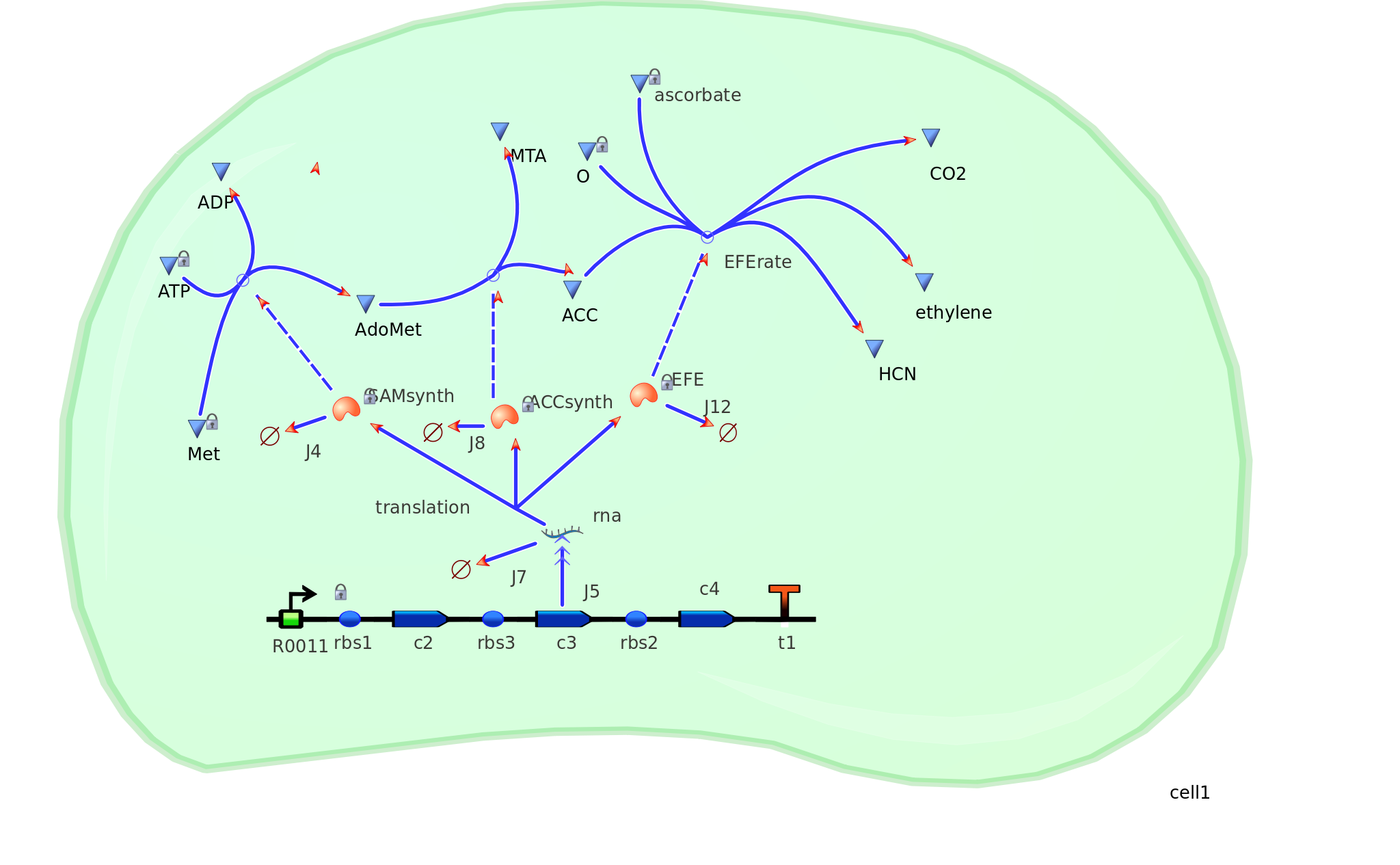
The program tinkercell was used to model the synthesis of ethylene in our e.coli cell. Where, in addition to the qualitative representation (figure 1) the program allowed, simulation of the model could be exercised inorder to measure the flux of metabolites – with respect to time - in the system given the input of values pertaining to substrate/enzyme concentration, and reaction rates were inputted. As a result, with a model, under certain parameters and assumptions made, quantitative data representing the theoretical: protein, mRNA and most importantly, ethylene outputs could be obtained for further analysis/optimization. | The program tinkercell was used to model the synthesis of ethylene in our e.coli cell. Where, in addition to the qualitative representation (figure 1) the program allowed, simulation of the model could be exercised inorder to measure the flux of metabolites – with respect to time - in the system given the input of values pertaining to substrate/enzyme concentration, and reaction rates were inputted. As a result, with a model, under certain parameters and assumptions made, quantitative data representing the theoretical: protein, mRNA and most importantly, ethylene outputs could be obtained for further analysis/optimization. | ||
| - | [[Image: | + | [[Image:Monash_modeling.png|200px|thumb|left|Figure 1: Metabolic modeling of the desired reaction in an '<i>E. coli</i>' cell]] |
| + | |||
More to come soon! | More to come soon! | ||
Revision as of 16:15, 21 October 2010

|
|
|||
Metabolic modelling of the ethylene generator
The program tinkercell was used to model the synthesis of ethylene in our e.coli cell. Where, in addition to the qualitative representation (figure 1) the program allowed, simulation of the model could be exercised inorder to measure the flux of metabolites – with respect to time - in the system given the input of values pertaining to substrate/enzyme concentration, and reaction rates were inputted. As a result, with a model, under certain parameters and assumptions made, quantitative data representing the theoretical: protein, mRNA and most importantly, ethylene outputs could be obtained for further analysis/optimization.
More to come soon!
 "
"