Team:Imperial College London/Modelling/Signalling
From 2010.igem.org
(Difference between revisions)
m |
m |
||
| Line 65: | Line 65: | ||
{| style="background:#e7e7e7;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;margin- top:5px;padding: 2px;" cellspacing="5"; | {| style="background:#e7e7e7;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;margin- top:5px;padding: 2px;" cellspacing="5"; | ||
|- | |- | ||
| - | |[[Image:IC_Signalling_Results2.png|270px]] | + | |[[Image:IC_Signalling_Results2.png|270px]] [[Image:IC_Signalling_Results3.png|270px]] [[Image:IC_Signalling_Results4.png|260px]] |
|- | |- | ||
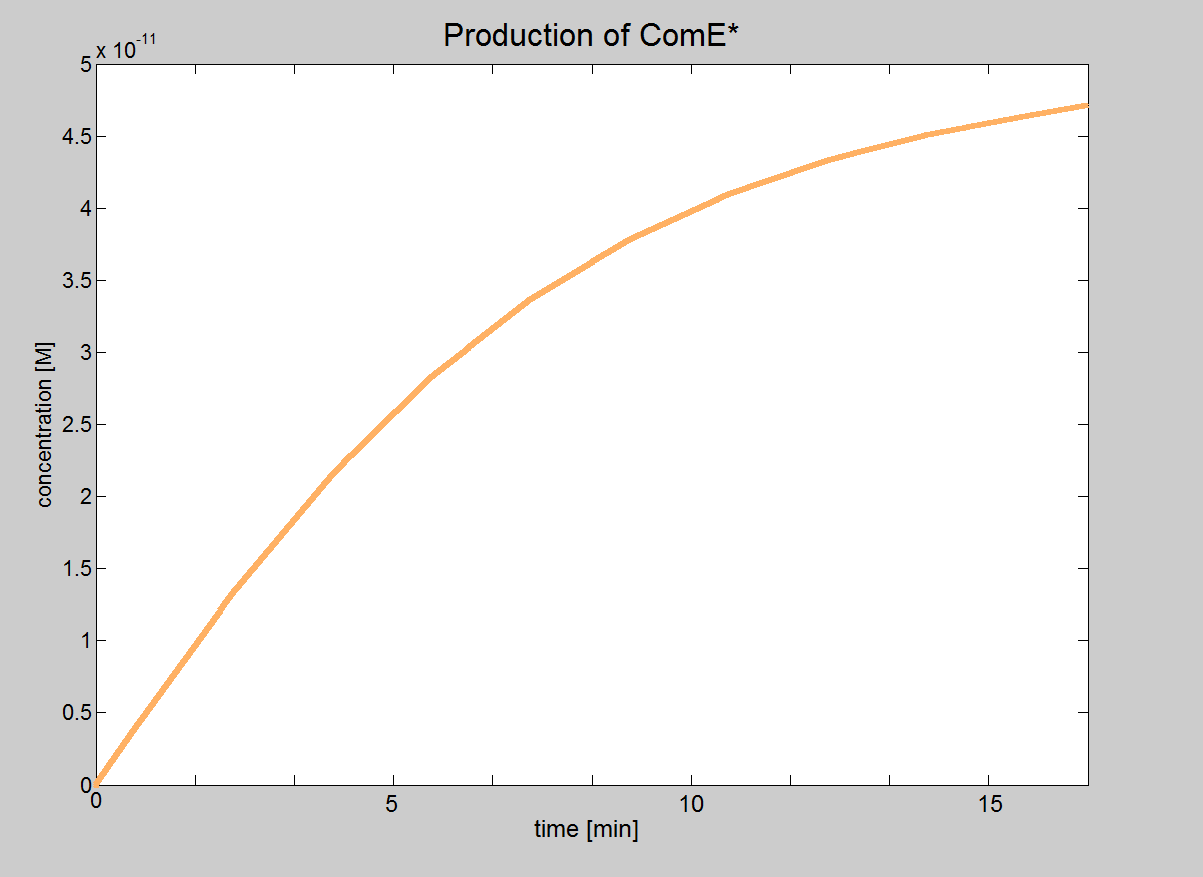
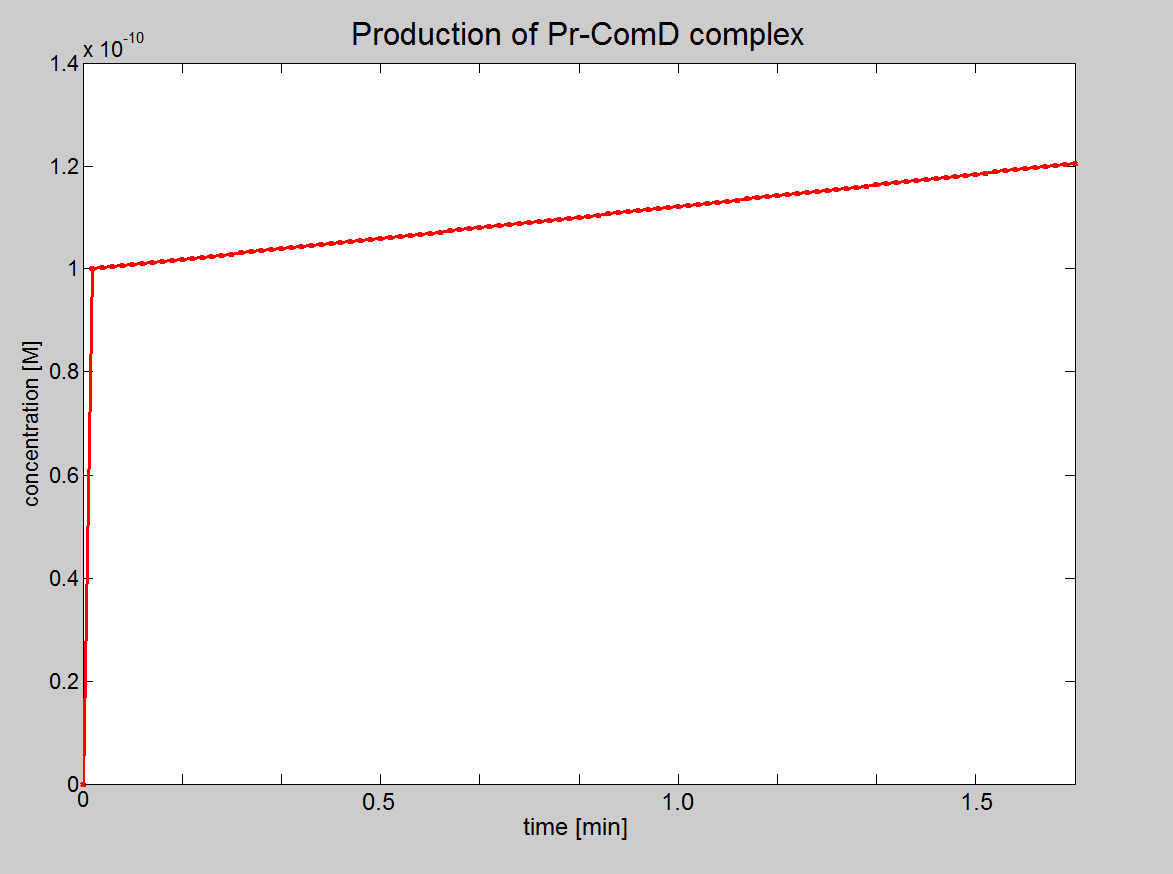
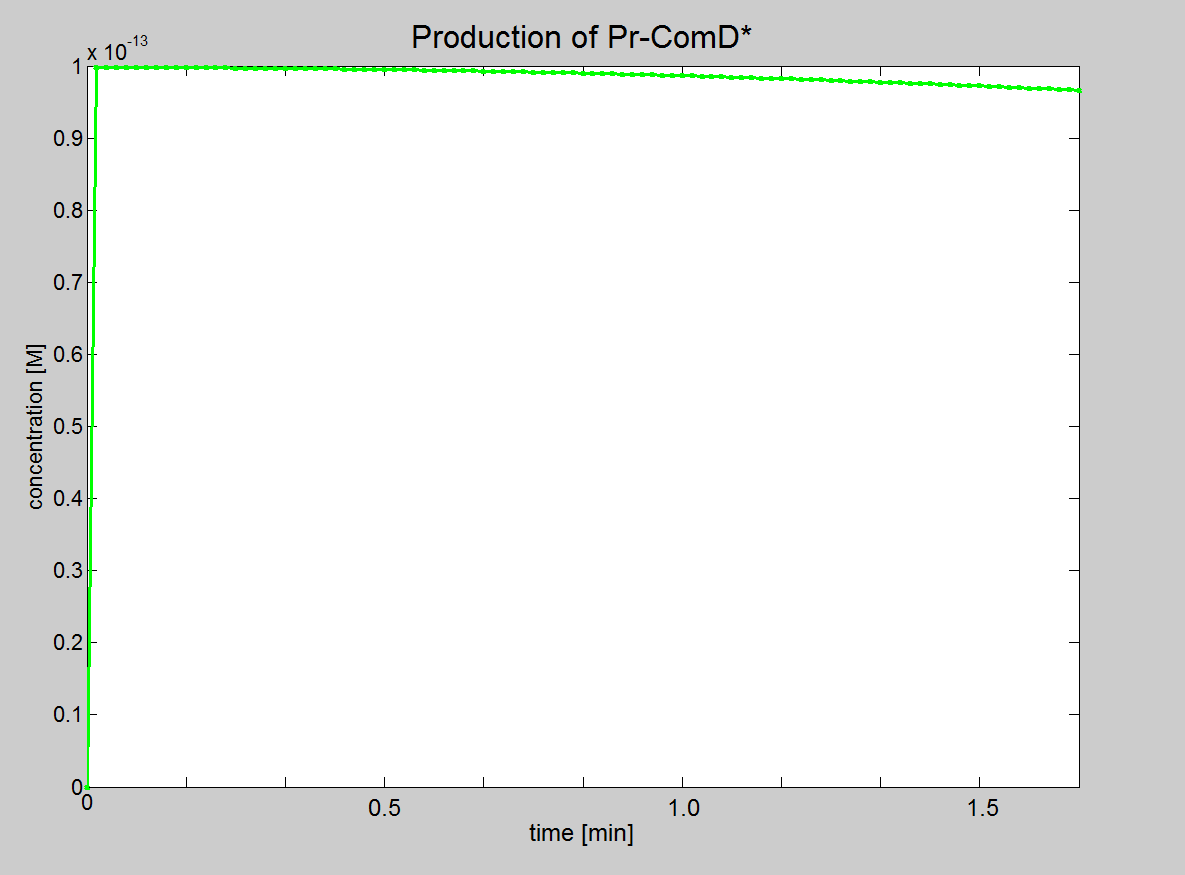
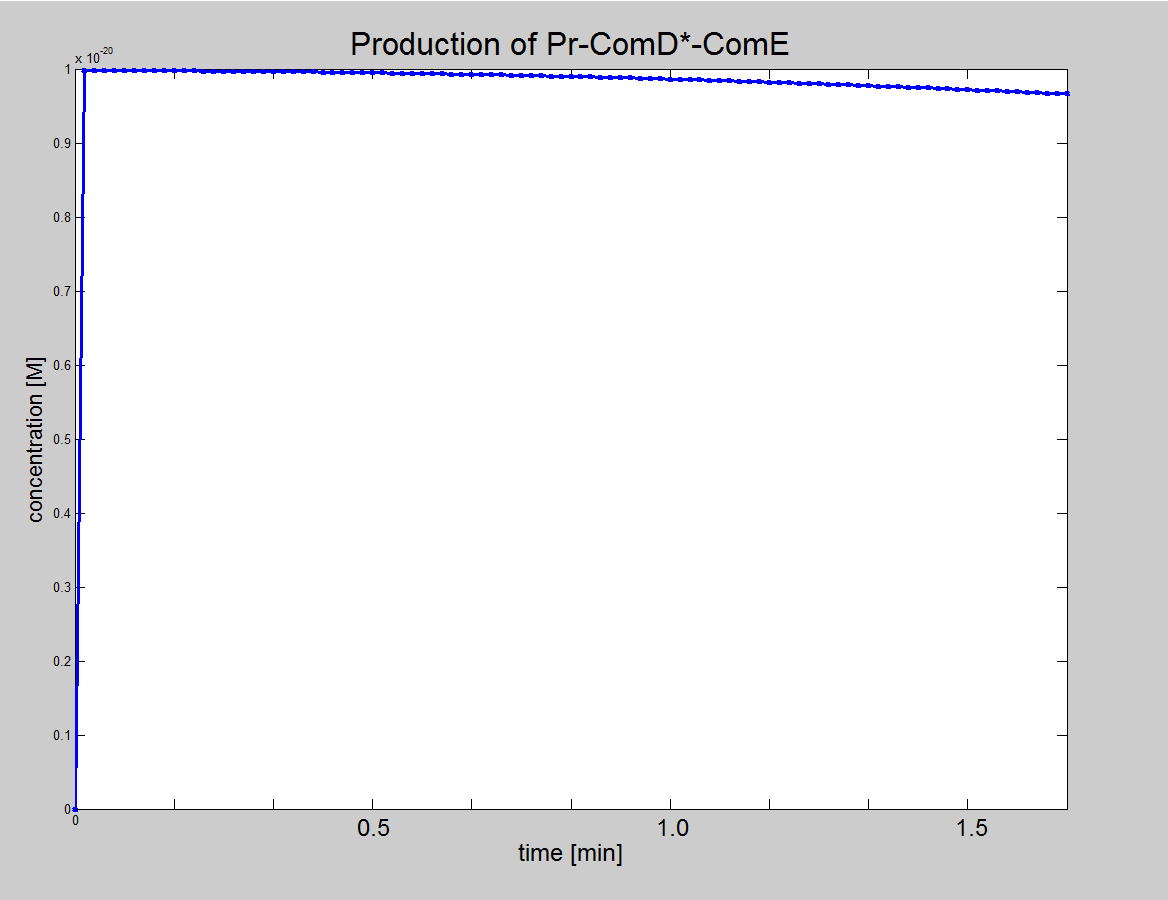
|1. Graph showing the production of Pr-ComD complex. 2. Graph showing the production of phosphorylated Pr-ComD* complex. 3. Graph showing the production of Pr-ComD*-ComE complex. Notice the steep increase of concentration for each of these graphs, which could be due to the high k<sub>1,2,3,4</sub> values. | |1. Graph showing the production of Pr-ComD complex. 2. Graph showing the production of phosphorylated Pr-ComD* complex. 3. Graph showing the production of Pr-ComD*-ComE complex. Notice the steep increase of concentration for each of these graphs, which could be due to the high k<sub>1,2,3,4</sub> values. | ||
Revision as of 18:33, 25 October 2010
| Modelling | Overview | Detection Model | Signaling Model | Fast Response Model | Interactions |
| A major part of the project consisted of modelling each module. This enabled us to decide which ideas we should implement. Look at the Fast Response page for a great example of how modelling has made a major impact on our design! | |
 "
"