Team:Cambridge/References/ProjectBioluminescence/Luciferase
From 2010.igem.org

Bioluminescence: Luciferases
Mutant luciferase (P. pyralis)
- Mutant Enhanced Photon Initiating Complex (EPIC) Luciferase (P.pyralis) Sequence
- Site-directed mutagenesis of firefly luciferase active site amino acids: a proposed model for bioluminescence color
- Improved practical usefulness of firefly luciferase by gene chimerization and random mutagenesis
Other Insect luciferases
- For colours we could use the click beetle luciferase red or green. They're only at the 'planning' stage though.
Bacterial Bioluminescence
- Vibrio Fischeri
- Vibrio Harveyi
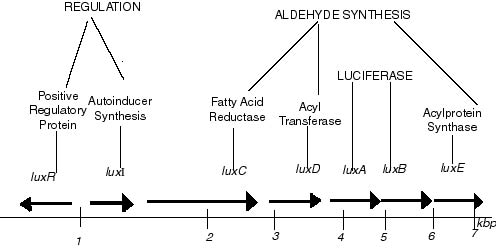
- Genetics of luxCDABE (adapted from wikipedia)
- 2006 attempts at BioBricking the lux operon
- Notes
- The lux wiki from parts registry
- NCBI Lux operon sequence
- parts registry lux operon -currently waiting on an email to see how useful/available this part is
- IMPORTANT INFO
- The Photorhabdus luminescens luxCDABE operon in the registry. It's not well characterised
Plasmid experiment
- Vibrio plasmid experiment -possible source of lux operon
- [1] - Transformation protocol for students from basics - includes several plasmids
- Plan
- set plasmid 555 into a BioBrick
- replace the promoter? sensitivity tuners?
- references
Transformation Experiment to test Firefly luciferase in registry
- Firefly luciferase is in the registry as BBa_I712019, and have been sent to us as part of the spring 2010 distribution.
- We need to find a new vector with a bacterial ribosyme binding site, initiator, and a stop codon. Having searched via the catalog in the plasmids section, then in expression plasmids, we have selected a constitutive plasmid:
- The plasmid selected is BBa_J13002, which is a TetR repressed POPS/RIPS generator, again part of the spring 2010 distribution - kit plate 1, well 13B, plasmid pSB1A2. It was proven to work by UMN iGEM 2009.
Wavelengths
548 nm http://pubs.acs.org/doi/pdf/10.1021/bi7015052
613 nm http://www.promega.com/pnotes/85/10904_11/10904_11.pdf
| Luciferase | Colour(at pH7.8) | λmax(nm)(pH 7.8) | (pH 6.0) | Base change | Amino acid change | |
|---|---|---|---|---|---|---|
| Genjii | yellow-green | 562 | 609 | |||
| C-M-1 | orange | 607 | 614 | G857-->A | Ser286-->Asn | |
| C-M-2 | red | 609 | 611 | G976-->A | Gly326-->Ser | |
| C-M-3 | red | 612 | 612 | C1297-->T | His433-->Tyr | |
| C-M-4 | yellow-orange | 595 | 609 | C1354-->T | Pro452-->Ser | |
| C-M-6 | green | 558 | 558 | G715-->A | Val239-->Ile | |
| C-M-11 | yellow | 565 | 612 |
 "
"