Team:Alberta/Tour/biobytes
From 2010.igem.org
We continued to develop the biobytes assembly method developed by last year’s team. The method has three main components:
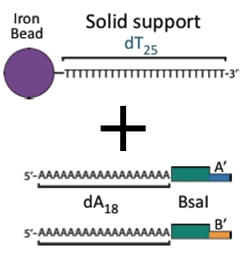
The Anchor: a ferro-magnetic bead attached to a piece of DNA. This piece serves as the initial piece from which we assemble a DNA construct. The bead allows us to manipulate the DNA with magnets making washing and subsequent attachments easier.
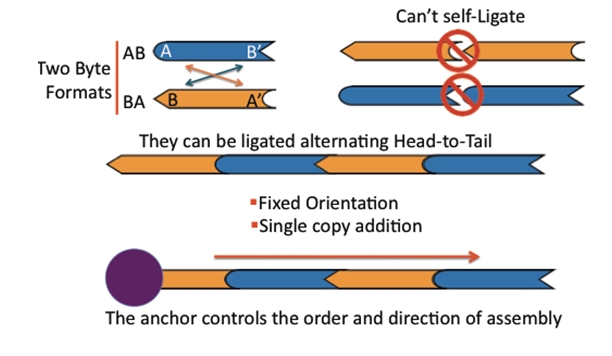
The bytes: DNA fragments that can be attached together to build up a larger construct. There are two types of pieces, AB and BA. The A end can join only with another A end and the B end can join only with another B end. As a result pieces can only be joined in a single orientation.
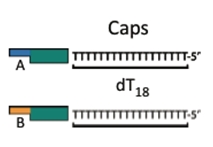
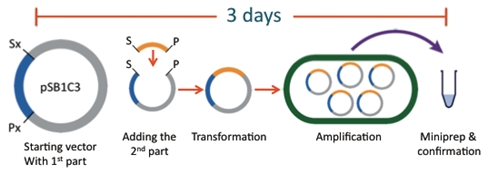
The cap: a DNA fragment that finishes off a construct and allows for circularization of the construct into a plasmid.
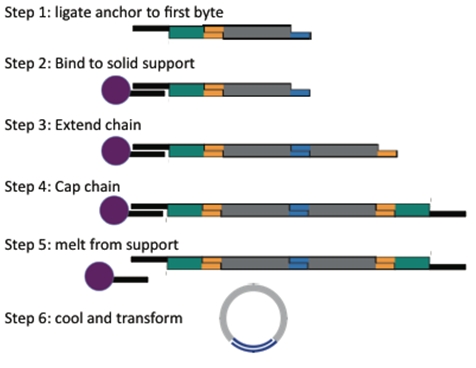
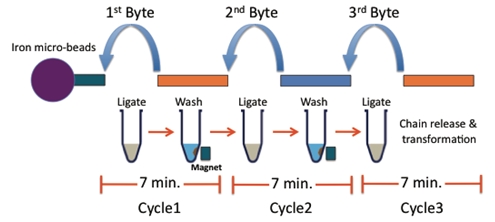
The process of building a plasmid is elegant and rapid!
Starting with an anchor, add the first piece and ligate.
Then hold the single piece construct in the tube by placing it on the magnetic rack. Now you can wash away most of the excess of piece 1.
Add the next piece and repeat until you have added all the pieces you want.
Then add the cap.
Now just heat to release the anchor and open up the cap, upon cooling the construct will circularize.
Easy!
Using this process we were able to assemble eight pieces in an afternoon!
 "
"