Team:UCL London/Project Description
From 2010.igem.org
(→Project Description) |
(→Project Description) |
||
| Line 41: | Line 41: | ||
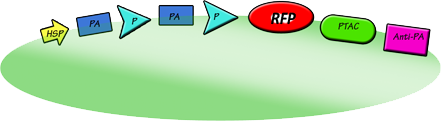
[[Image:UCL-papapa.png|150px|center]] | [[Image:UCL-papapa.png|150px|center]] | ||
| - | The promoter activator activates the promoter, which then activates another promoter activator and thus produces more of the promoter. This is essential as it increases the rate of protein expression. With this being activated, the | + | The promoter activator activates the promoter, which then activates another promoter activator and thus produces more of the promoter. This is essential as it increases the rate of protein expression. With this being activated, the RFP, which is our red dye protein, will be activated subsequently resulting in the expression of our red dye protein indicating the succesful operation of our circuit. In our circuit for the sake of proof of principle, instead of expressing a typical protein, we will introduce a genetic sequence which translates a fluorescent red protein so that once it is translated, it will secrete the red dye and hence prove that our circuit does in actual fact work and added an artistic touch to our project. |
[[Image:Ucl-PAPAP-A.png|700px|center]] | [[Image:Ucl-PAPAP-A.png|700px|center]] | ||
Latest revision as of 02:17, 28 October 2010
Project Hypoxon
Project Abstract
UCL’s Biochemical Engineering department has been at the forefront of biopharmaceutical manufacture for many years. Extraordinary advances in the life sciences have great potential to improve our quality of life through better medicines and a cleaner environment.
Our project aims to create “independent” cells, capable of self-induction into the production phase, without the introduction of any chemical into the closed system. By exploiting genetically modified E.coli to respond to hypoxia, we eliminate the need of IPTG induction. The functioning genetic circuit would be signaled by the production of red fluorescent proteins (RFPs).
While the RFP can be used as a live monitoring technique for oxygen, replacing the RFP with pharmaceutical proteins promises higher yields and more consistent batches which is a step closer to providing cheaper healthcare to everyone.
Application in Industry
Cells need constant monitoring for an "oxygen spike". This spike occurs when cells have reached their maximum growth capacity in the fermenter, and are competing for nutrients in the fermenter. At that point, an operator has to introduce a pre-determined volume of IPTG (chemical) into the fermenter to stop the cell growth and channel the energy into transcription of a certain gene. This is how the cells are manipulated to produce pharmaceuticals!
Reliability
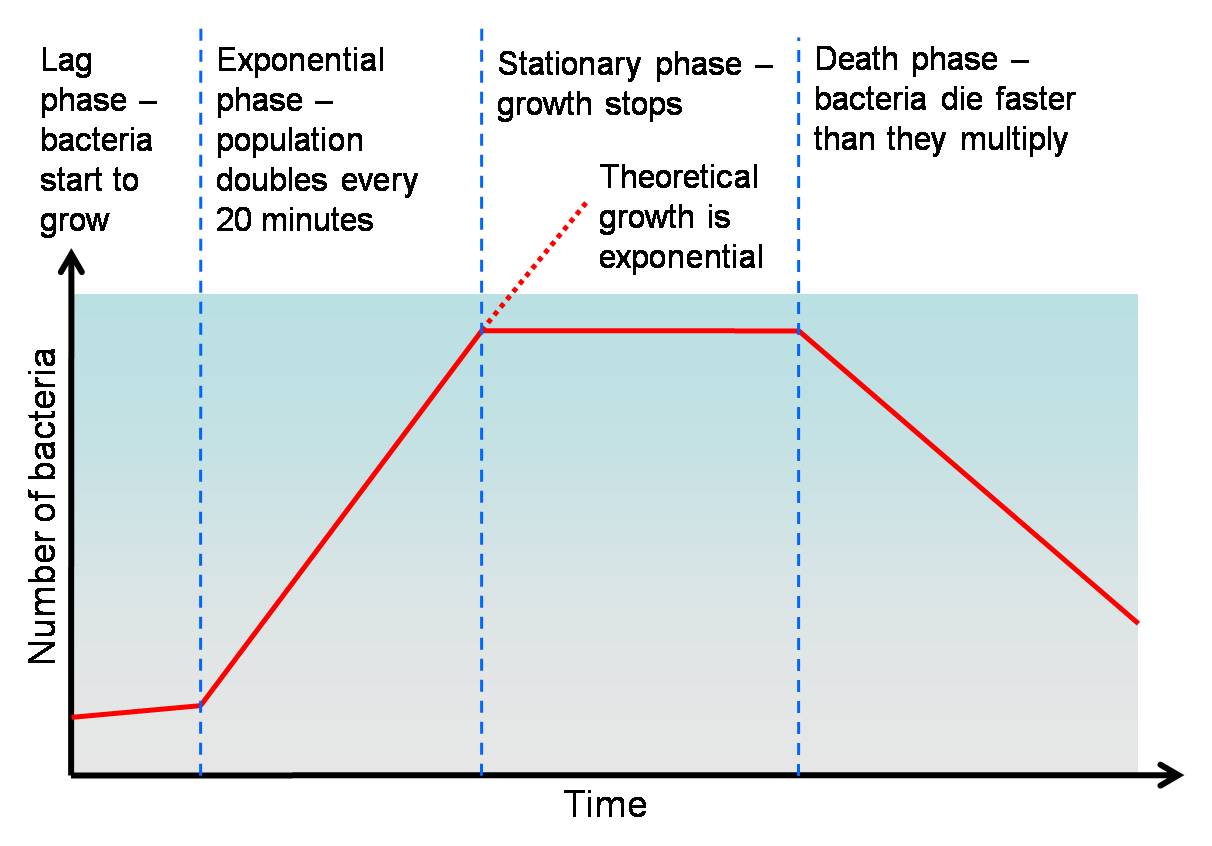
Cells go through different phases before reaching their maximum number. There is no way of telling how long the lag phase would be, or when the oxygen spike would occur. (We experienced a four day lag phase in one of our four fermentations where the cells were not dead, but not growing! This can get frustrating, and can give Steve, the guardian of the fermenters horrible ideas like killing the cells and starting again!
What if the oxygen spike happens on a weekend, or falls somewhere outside the 8-hour-shift? This is where auto-induction becomes useful. The cells would automatically realise the oxygen spike and change from growth to manufacturing automatically!
Cost
Not only does auto-induction save money on IPTG, it can eliminate the need of IPTG storage saving massive amounts of money on refrigeration in a sterile environment(IPTG must be stored at -20 degrees). Also, there is no more risks of contamination of the IPTG, or of the batch when introducing the IPTG! That means that less batches are wasted, and there is therefore a lower chance of batch failure per year. Therefore, with more batches produced per annum, the cost of the drug would decrease giving more people access to it!
Environmental
As mentioned above, IPTG needs to be stored at -20 degrees. Not using IPTG would eliminate the need for refrigerators, making the pharmaceutical industry a little greener.
Project Description
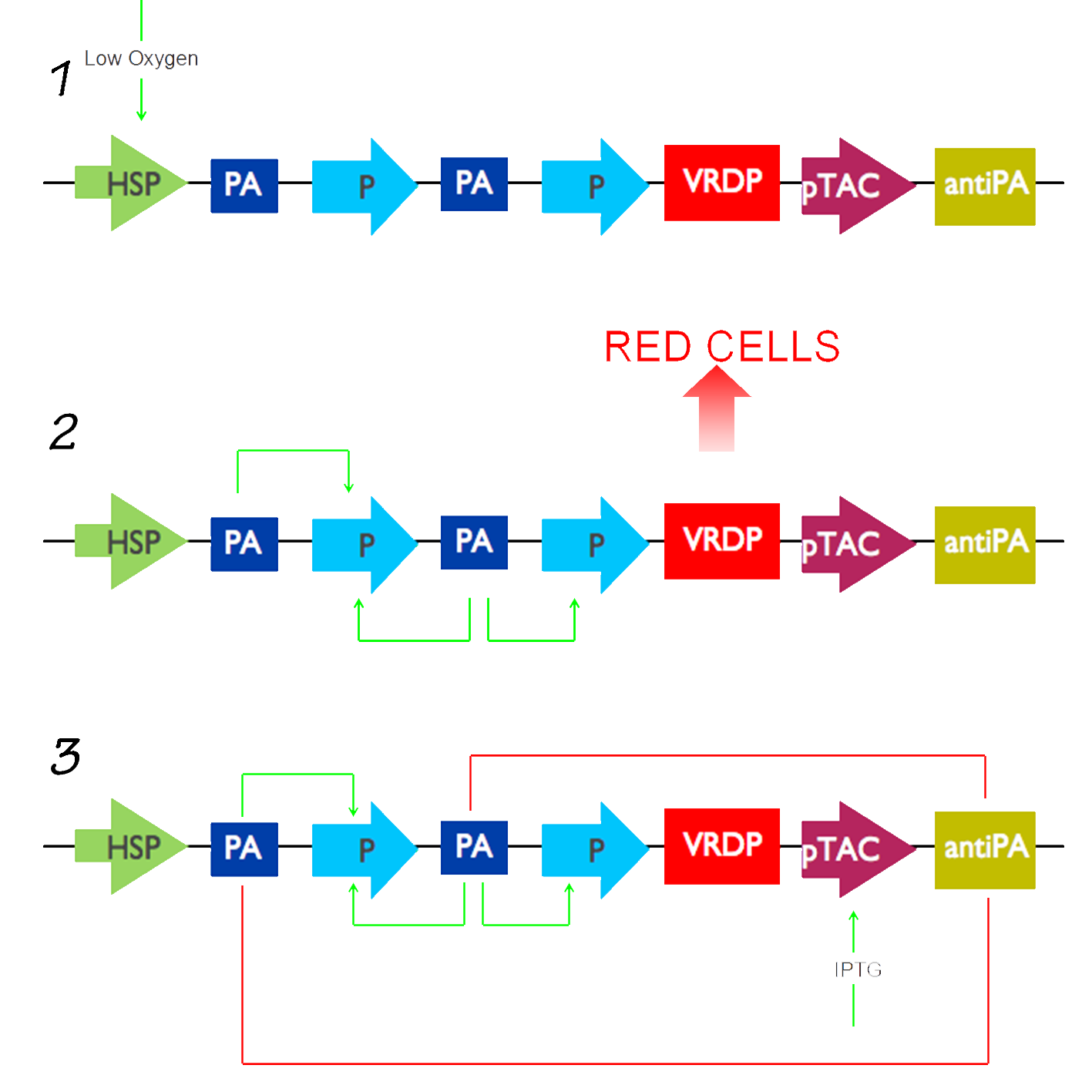
During the initial phase of the fermentation process, the recombinant E.coli cells with our genetic circuit will grow very slow at first, and this initial phase is referred to as the "Log Phase". As this happens, the DOT(Dissolved Oxygen Tension) in the fermenter decreases, at a rate which is indirectly proportional to the rate of cellular growth. The DOT will continue to fall untill it falls in the region of 15-20%. Usually during protein expression in E.coli cells lacking our genetic circuit, this is the point at which our inducer IPTG would be added, manually resulting in the LacI repressser being repressed itself and thus allowing our hybrid pTAC to be expressed. The desired protein is then expressed. However, bearing in mind the DOT, we will use this unique factor to introduce hypoxia into our circuit. Essentilly, our circuit will start of with an HSP, Hypoxia Sensitive Promoter, which will detect the change in DOT when it falls below the threshold of 20%. At that point, it will be triggered and will activate a positive feedback loop consisting of 2 promoter Activators and 2 Promoters in the following format;
The promoter activator activates the promoter, which then activates another promoter activator and thus produces more of the promoter. This is essential as it increases the rate of protein expression. With this being activated, the RFP, which is our red dye protein, will be activated subsequently resulting in the expression of our red dye protein indicating the succesful operation of our circuit. In our circuit for the sake of proof of principle, instead of expressing a typical protein, we will introduce a genetic sequence which translates a fluorescent red protein so that once it is translated, it will secrete the red dye and hence prove that our circuit does in actual fact work and added an artistic touch to our project.
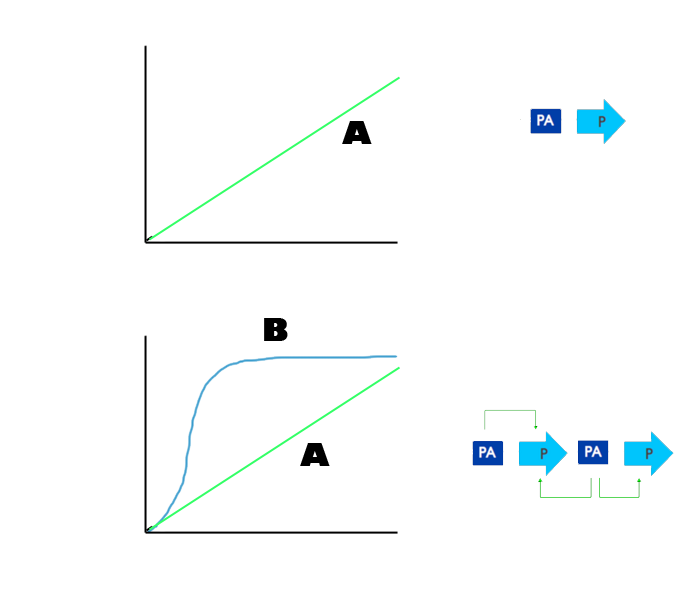
The above diagram clearly demonstrates the effect the PA-P-PA-P loop has on the production of the promoter. Line A is that of the PA-P system alone and it takes more time, nearly double that of the feedback loop, to transcribe the same amount of the promoter.
The pTAC here no longer serves the purpose of being our expression system, it now serves the purpose of commencing the switch off of our circuit. This will happen by addition of IPTG which will result in the pTAC no longer being repressed. This will allow the activation of the pTAC which will activate the consecutive antiPA gene resulting in repression of the promoter activators and hence the switching off of our genetic circuit. Now in biopharmaceutical production, this extra sequence consisting of the pTAC and the antiPA is not required since there is no need to switch the genetic circuit of and so will allow maximum protein expression. But in this project, we feel it is essential to provide a mechanism for its closure, a genuine rule of thumb in Synthetic biology.


 "
"



 Twitter
Twitter Facebook
Facebook UCL
UCL Flickr
Flickr YouTube
YouTube