Team:BCCS-Bristol/Wetlab/Part Design/BioBricks/PyeaR
From 2010.igem.org
iGEM 2010
Our Biobrick : BBa_K381001
We've focussed on building one key new BioBrick; BBa_K381001; encoding expression of GFP in response to Nitrates or Nitrites. During the project we both made and well characterised this part, details of which can be found in our Experiment section, whilst information on its sequence and how to get hold of it can be found at the BBa_K381001 page on the parts registry website.To find out more about each of the component parts, including why we chose them and background information, use the navigation bar above.
Part Design
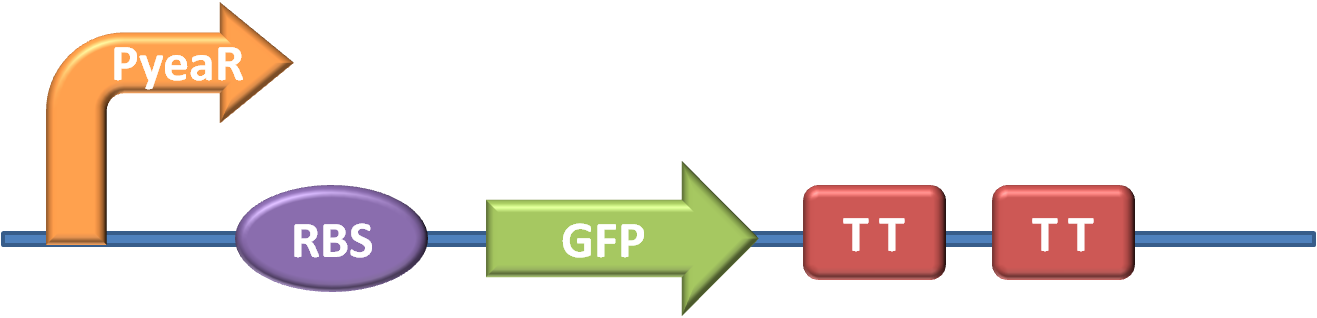
BBa_K381001 is a composite part of two pre-existing BioBricks taken from the parts registry; a Nitrate sensitive promoter and a GFP generator, along with strong RBS and transcription terminators.
The promoter is BBa_K216005, originally submitted by Edinburgh in 2009, whilst the GFP generator, along with strong RBS and transcription terminators is BBa_E0840, provided in well 12O, kit plate 1 of the 2010 distribution.
How does it work?
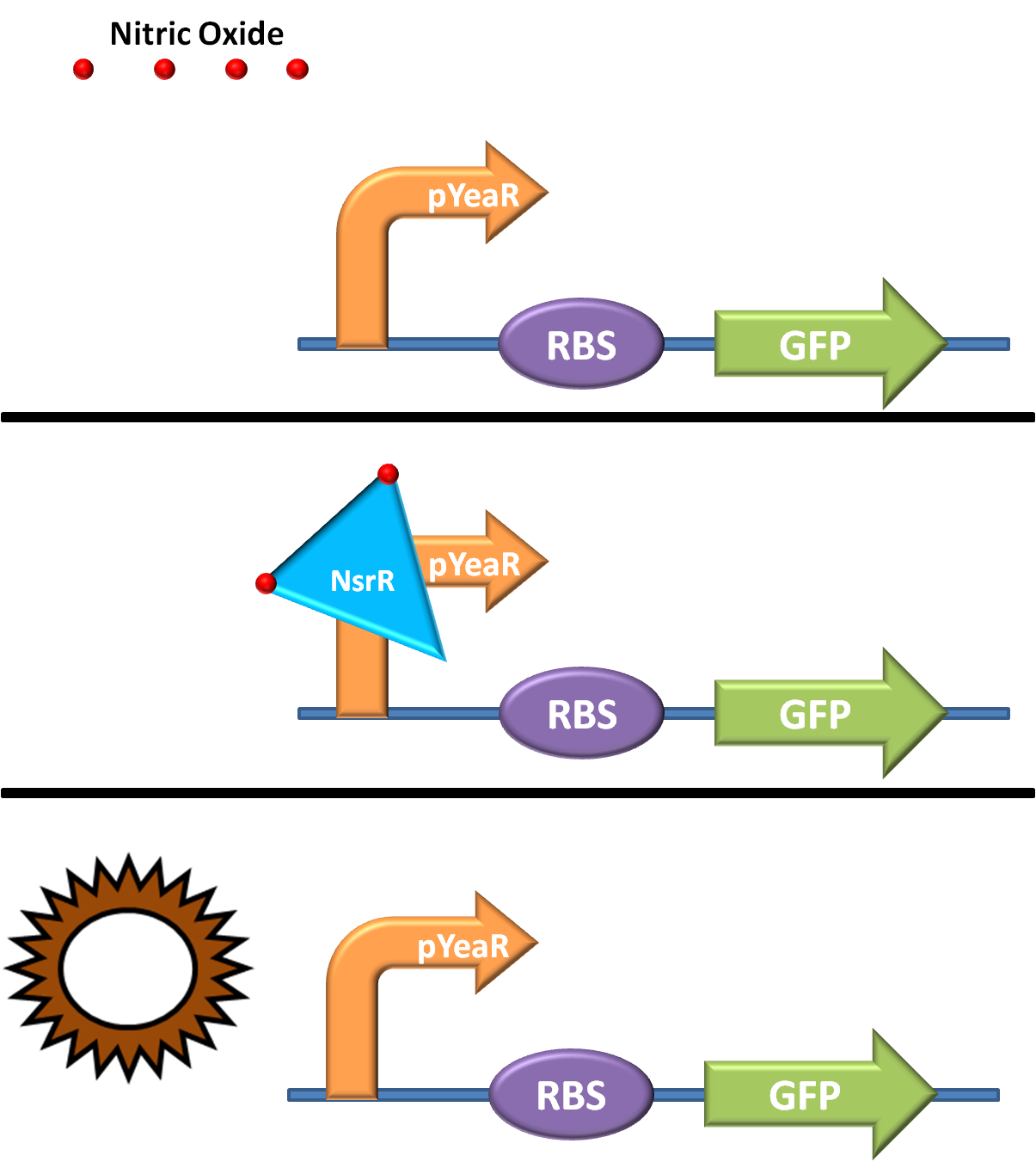
The BBa_K381001 BioBrick is normally repressed by proteins in the cell native to E. coli. However, when cells containing the BioBrick are exposed to Nitrates, these will diffuse into the cell, and be converted to Nitric Oxide.
This in turn interacts with the proteins blocking BioBrick expression and allows the production of GFP. For more information on how this works, refer to the detailed promoter page here
Performance
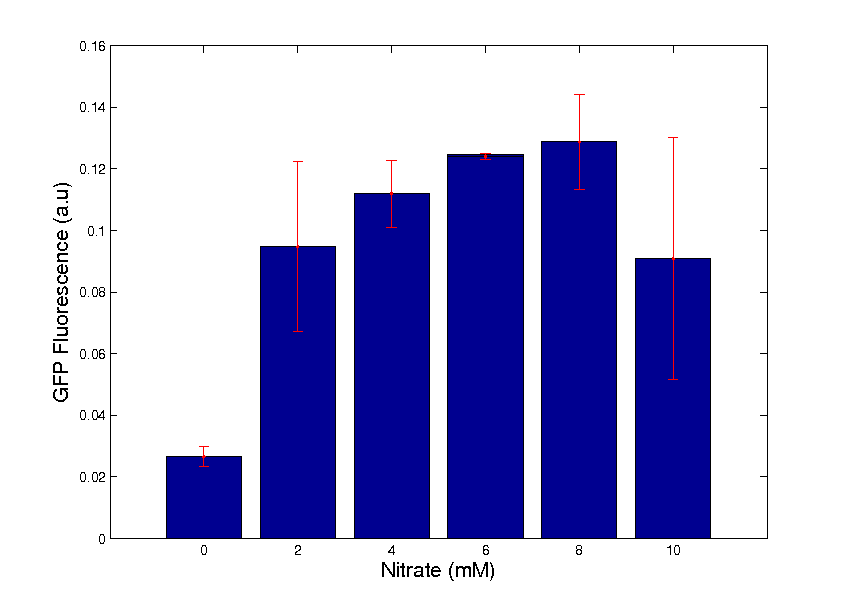
Performance of the promoter specifically was assessed using a Miller Assay. Whilst performance of the BioBrick as a whole was characterised with an assay measuring change in GFP expression with changes in Potassium Nitrate levels. Graph 1 shows a GFP assay of the BioBrick between 0 - 10mM, indicating a clear trend of increasing fluorescence with Nitrate levels. For more in depth information on our characterisation, click here.
 "
"