Team:Washington/Tools Created/New Software
From 2010.igem.org
As the systems created by synthetic biologists get more complex, automation and computer-aided design will be needed. Computers will eventually have a central in the design, construction, and testing of new devices. To meet the emerging needs of the synthetic biology community, the Washington 2010 iGEM team has developed two software tools.
WikiDust is a plugin that can be used to export TinkerCell models to the iGEM wiki or other webpages. Users can then download the models for use in their own larger system, or to see how they respond when parameters are changed. We hope it will provide an easy way to share models, encouraging more teams and researchers to describe their parts quantitatively. This will allow successful reuse of their parts in the future.
The first thing WikiDust does is allow you to associate TinkerCell items with parts on the registry. When the TinkerCell model is uploaded to a webpage, clicking on each item will go to the correct part page--in fact, this diagram was generated using WikiDust. The same linking mechanism could also be extended in the future to download the reaction kinetics, icon, etc. of the items into TinkerCell. For now though, it's mostly a proof of concept demonstrating the use of a new semantic knowledgebase of parts.
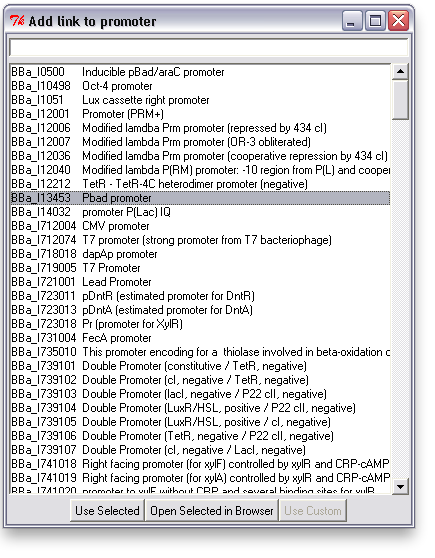
Associating a link with an item is easy: just right click on an item and choose "Add Link". This will bring up a search window with suggested parts. You can search for a phrase, a BioBrick ID, etc. You can also open parts in a browser to read more about them or confirm that you've found the right one.
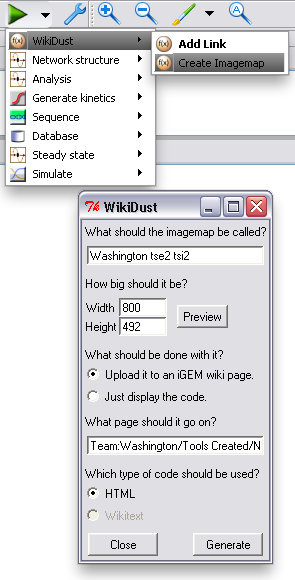
After the model of your device or system is finished, you can upload a representation of it to your page. WikiDust will automatically handle uploading to most MediaWiki sites, or it can just display code for you to copy and paste. By default, a download link for the complete model file is also included.

Download TinkerCell to start using WIkiDust today.
We have also developed a program to submit multiple parts to the Parts Registry at once. It will be useful for anyone who needs to submit a series of parts—promoters of different strengths, variations on a protein, etc.
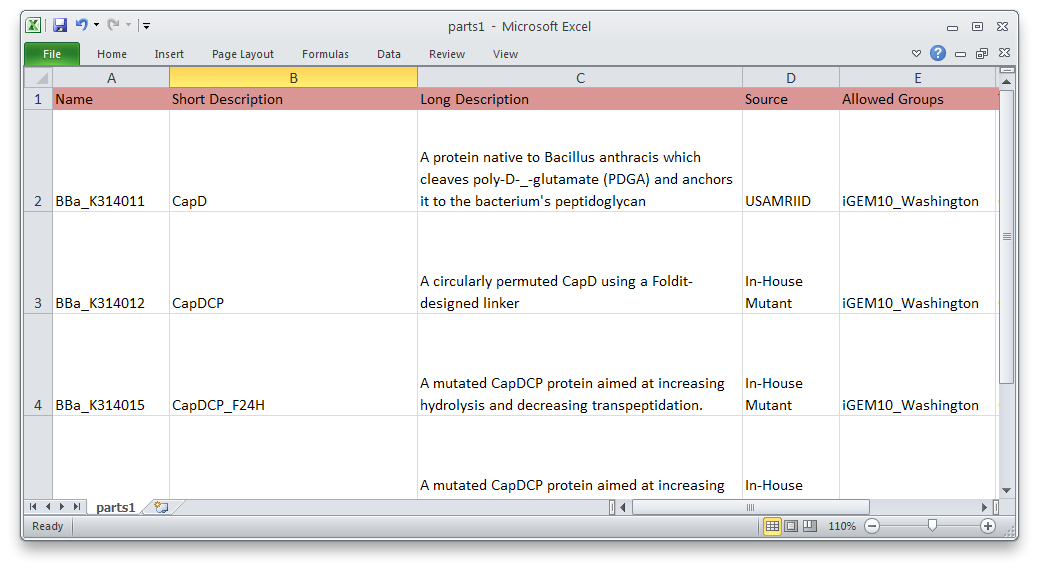
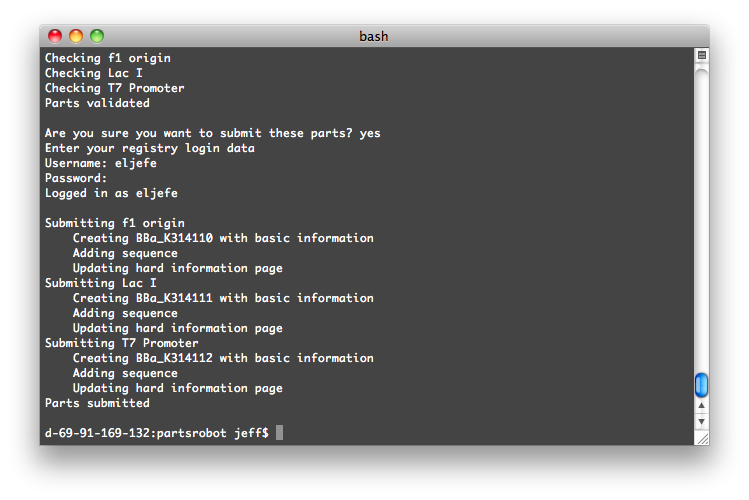
Using PartsRobot is simple. Just fill out the template spreadsheet with information about each part. Then type:
python partsrobot.py submit yourspreadsheet.csv
The program will log into the Registry and check your parts for syntax errors. Then, after you give the go-ahead, it will submit them.
Get PartsRobot from the SourceForge Page.
 "
"