Team:TU Munich/Project
From 2010.igem.org
(→Concept) |
|||
| Line 32: | Line 32: | ||
= Work Progress = | = Work Progress = | ||
Every network starts with a basic unit. While our declared aim is to enable networks allowing fine-tuning of gene expression beyond the regular on/off, exploring such an on/off switch/signal pair is the first step towards a functional network. We constructed several units and tested their efficiency, robustness and reproducibility ''in vivo'', ''in vitro'' and ''in silico''. Furthermore we developed a software which allows easy constructions of networks delivering a ready network. Conclusive elaboration of a few first RNA-based logic units is the major contribution of our iGEM team. | Every network starts with a basic unit. While our declared aim is to enable networks allowing fine-tuning of gene expression beyond the regular on/off, exploring such an on/off switch/signal pair is the first step towards a functional network. We constructed several units and tested their efficiency, robustness and reproducibility ''in vivo'', ''in vitro'' and ''in silico''. Furthermore we developed a software which allows easy constructions of networks delivering a ready network. Conclusive elaboration of a few first RNA-based logic units is the major contribution of our iGEM team. | ||
| + | |||
| + | =Results= | ||
<!-- The idea behind our project is to change the way BioBricks have been used up to now. Over the years, many receptors and signals have been constructed as BioBricks during the annual iGEM competition, but still it is not possible to interconnect these Bricks in a complex biological network resuting in a cell, that is able to respond to its environment giving differenciated responses depending on the input signals. (Beispiel: cambridge hat das gemacht, xx dies, aber eine zelle kann nicht beides...<br> | <!-- The idea behind our project is to change the way BioBricks have been used up to now. Over the years, many receptors and signals have been constructed as BioBricks during the annual iGEM competition, but still it is not possible to interconnect these Bricks in a complex biological network resuting in a cell, that is able to respond to its environment giving differenciated responses depending on the input signals. (Beispiel: cambridge hat das gemacht, xx dies, aber eine zelle kann nicht beides...<br> | ||
Revision as of 15:48, 10 October 2010
|
||||||||
|
|
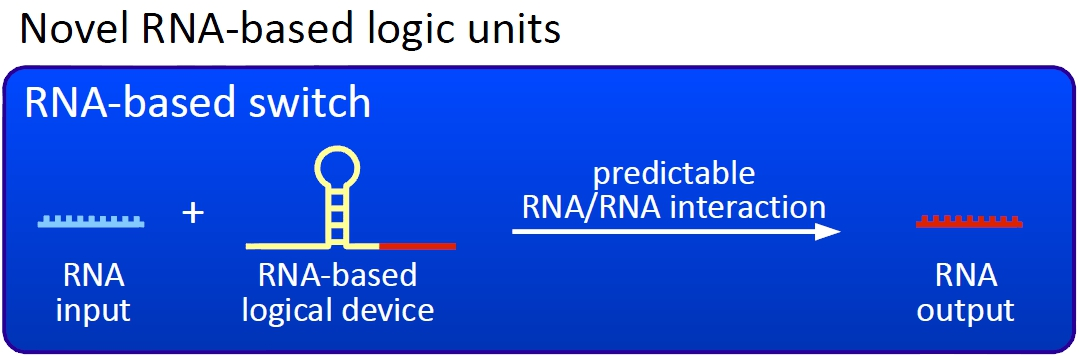
VisionAlthough classical molecular biology and genetic engineering equipped the science community with many functional proteins and possible applications, using and linking different parts together to a working network still requires highly complicated strategies. The switches established in molecular biology, for example the lac operon, are highly limited, as most of them rely on interactions including metabolites and proteins providing only one on/off-signal. Thus, it is hardly possible to build up a logic network inside a cell without interferences between different switches. Our approach is to change this by developing a new and more robust way to control E. coli cells using RNA-RNA-interaction based switches, which we call bioLOGICS. ConceptThe basic principle of our switches are short RNA sequences, the scientific idea shares similiarities with the principle of antitermination but also inherits a completely new way of RNA based transcription regulation. We used three-dimensional structure predictions and thermodynamic calculation to develop a set of switches - about 50 nucleotides - and signals - about 20 nucleotides. The switch forms a stem loop causing transcription termination, which can be resolved upon binding of a signal. On/off switching can therefore be easily controlled by signal avaiability and provides a new concept to control gene transcription.
Our switches are based on the principle of transcriptional antitermination. Transcription can be cancelled by the formation of a RNA stem loop in the nascent RNA-chain, which then causes the RNA polymerase to stop transcription and fall off the DNA. We use sequences as transcriptional switches, that are calculated to be capable of stem loop formation. We plan to control the termination at these switches with a small RNA molecule (our RNA "signal") that is complementary to a part of the stem loop forming sequence. This small functional RNA inhibits the stem loop formation by complementary base-pairing and hence avoids transcription termination. The signal is composed of two parts: While the first part provides specifity (recognition site), the second part causing stem loop disintegration (functional core) can in principle be the same for all bioLOGICS. Therefore variation of the first part allows the construction of an endless number of switches. The functional core causing stem loop disintegration is based on a working system (see attenuation) established by nature. Different stem loops were tested in this effort: Regulatory parts from the E. coli trp-operon, his-operon and one based on previous iGEM-work.
Network constructionDesigning complex biological networks based on either traditional protein engineering or our new bioLOGICS is still a complex task. We developed a software which allows the fast construction of a bioLOGICS based networks. Work ProgressEvery network starts with a basic unit. While our declared aim is to enable networks allowing fine-tuning of gene expression beyond the regular on/off, exploring such an on/off switch/signal pair is the first step towards a functional network. We constructed several units and tested their efficiency, robustness and reproducibility in vivo, in vitro and in silico. Furthermore we developed a software which allows easy constructions of networks delivering a ready network. Conclusive elaboration of a few first RNA-based logic units is the major contribution of our iGEM team. Results |
|||||||
 "
"