Team:TU Delft/Project/rbs-characterization
From 2010.igem.org
(→Ribosome Binding Site Characterization) |
|||
| Line 4: | Line 4: | ||
Part of our project is to measure the effect of various ribosome binding site (RBS) sequences. A RBS sequence is a specific mRNA sequence that folds in such a way that it attracts the ribosome. This ribosome binds to the mRNA molecule and starts translation of the mRNA into protein. Accurate information on how well this binding occurs (The binding strength) can be very useful when designing new biological systems. | Part of our project is to measure the effect of various ribosome binding site (RBS) sequences. A RBS sequence is a specific mRNA sequence that folds in such a way that it attracts the ribosome. This ribosome binds to the mRNA molecule and starts translation of the mRNA into protein. Accurate information on how well this binding occurs (The binding strength) can be very useful when designing new biological systems. | ||
| + | ===Continue reading=== | ||
| + | * [[Team:TU_Delft/Project/rbs-characterization/parts|Proteins & Parts]] | ||
| + | * [[Team:TU_Delft/Project/rbs-characterization/characterization|Characterization]] | ||
| + | * [[Team:TU_Delft/Project/rbs-characterization/results|Results and Conclusions]] | ||
| + | |||
| + | ==Experimental Setup== | ||
We took five different RBS sequences from the [http://partsregistry.org/Ribosome_Binding_Sites/Prokaryotic/Constitutive/Anderson Anderson RBS family] (J61100, J61101, J61107, J61117, J61127). The standard RBS [http://partsregistry.org/Part:BBa_B0032 B0032] was used as a reference for our characterization. | We took five different RBS sequences from the [http://partsregistry.org/Ribosome_Binding_Sites/Prokaryotic/Constitutive/Anderson Anderson RBS family] (J61100, J61101, J61107, J61117, J61127). The standard RBS [http://partsregistry.org/Part:BBa_B0032 B0032] was used as a reference for our characterization. | ||
| Line 48: | Line 54: | ||
<html><div style='clear:both'> | <html><div style='clear:both'> | ||
</html> | </html> | ||
| - | |||
| - | |||
| - | |||
| - | |||
Revision as of 09:03, 20 October 2010
Ribosome Binding Site Characterization
Part of our project is to measure the effect of various ribosome binding site (RBS) sequences. A RBS sequence is a specific mRNA sequence that folds in such a way that it attracts the ribosome. This ribosome binds to the mRNA molecule and starts translation of the mRNA into protein. Accurate information on how well this binding occurs (The binding strength) can be very useful when designing new biological systems.
Continue reading
Experimental Setup
We took five different RBS sequences from the Anderson RBS family (J61100, J61101, J61107, J61117, J61127). The standard RBS B0032 was used as a reference for our characterization.
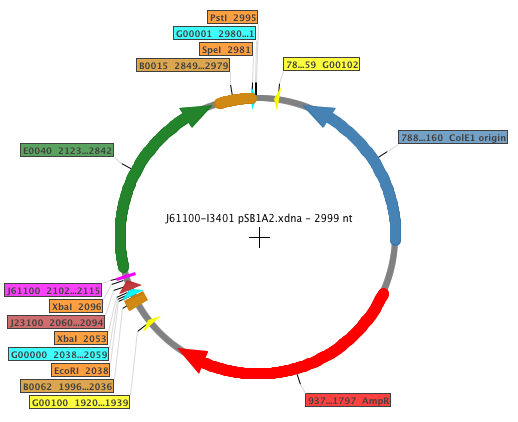
All these RBS sequences were placed in front of the standard GFP coding sequence, so expression could be measured. The Biobricks generated in order to perform the experiments were: K398500, K398501, K398502, K398503, K398504. The general map of the construction is shown below, where the RBS is displayed in fucsia.
| Feature | Function |
| AmpR | Ampicillin resistance |
| B0015 | Transcriptional (double) terminator |
| B0062 | Transcriptional terminator |
| E0040 | GFP |
| G00000 | Standard prefix |
| G00001 | Standard suffix |
| G00100 | VF2 primer binding site |
| G00101 | VR primer binding site |
| J61100 | RBS Anderson family |
| J23100 | Promoter |
 "
"