Team:Newcastle/modelling
From 2010.igem.org
(Difference between revisions)
Swoodhouse (Talk | contribs) (→Flux balance analysis) |
Swoodhouse (Talk | contribs) (→Calcium carbonate) |
||
| Line 17: | Line 17: | ||
===Flux balance analysis=== | ===Flux balance analysis=== | ||
| - | In order to inform our part design, [[Team:Newcastle/Urease#Flux_balance_analysis|we performed flux balance analysis]] to identify pathways which indirectly lead to urea hydrolysis. | + | In order to inform our calcium carbonate part design, [[Team:Newcastle/Urease#Flux_balance_analysis|we performed flux balance analysis]] to identify pathways which indirectly lead to urea hydrolysis. |
[[Image:Newcastle_Arginine_and_Ornithine_Degradation.png|600px]] | [[Image:Newcastle_Arginine_and_Ornithine_Degradation.png|600px]] | ||
| Line 29: | Line 29: | ||
*[[Media:Newcastle_FBA_urease.txt|Metabolic flux during maximum urease activity]] (reaction rxn00101, Urea amidohydrolase), showing large flux through the L-Arginine amidinohydrolase reaction (rxn00394). | *[[Media:Newcastle_FBA_urease.txt|Metabolic flux during maximum urease activity]] (reaction rxn00101, Urea amidohydrolase), showing large flux through the L-Arginine amidinohydrolase reaction (rxn00394). | ||
*[[Media:Newcastle_flux_balance_analysis.m.txt|Matlab code]] | *[[Media:Newcastle_flux_balance_analysis.m.txt|Matlab code]] | ||
| + | |||
===Biochemical network in SBML=== | ===Biochemical network in SBML=== | ||
Revision as of 20:41, 25 October 2010

| |||||||||||||
| |||||||||||||
Contents |
Modelling
In order to understand the behaviour of our parts, we computationally modelled them by encoding biochemical networks in SBML and simulating them using COPASI.
Calcium carbonate
For more details, please see the Calcium carbonate precipitation via urease expression.
Flux balance analysis
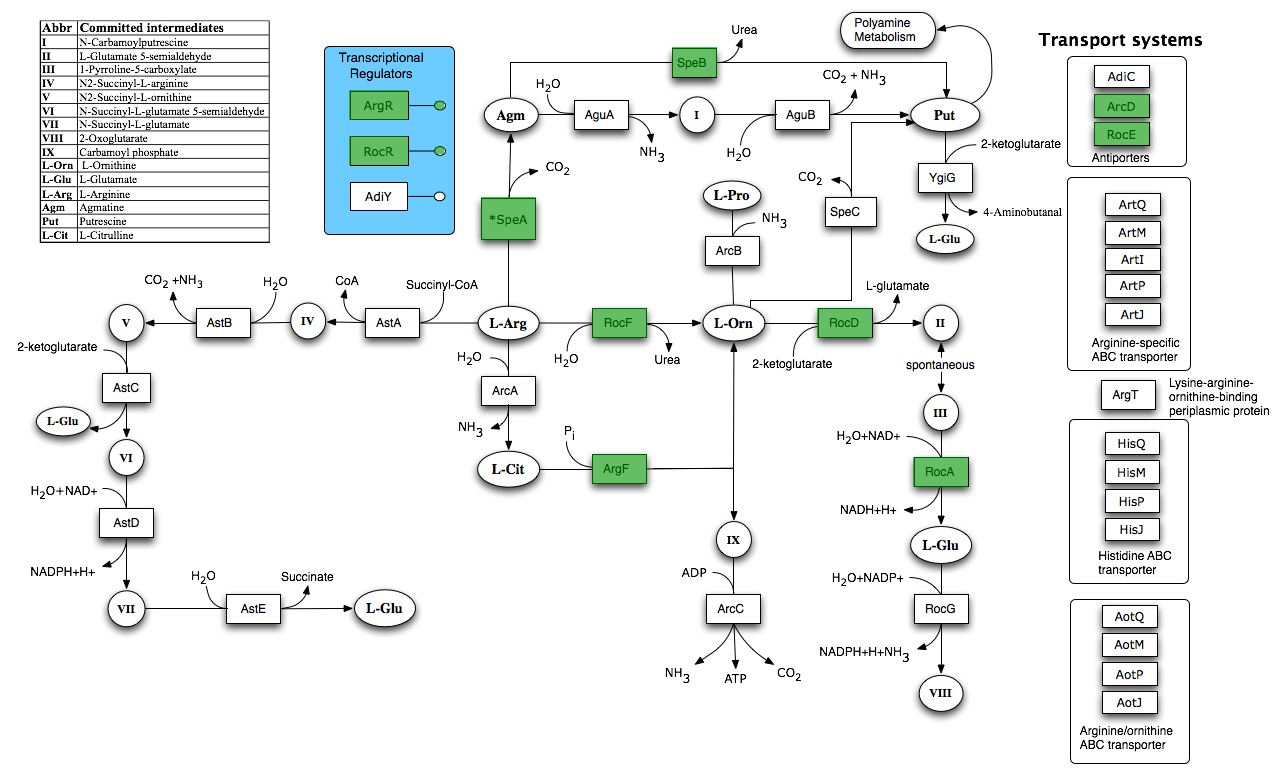
In order to inform our calcium carbonate part design, we performed flux balance analysis to identify pathways which indirectly lead to urea hydrolysis.
Taken from SEED
Downloads:
- Metabolic flux in standard conditions (maximal growth)
- Metabolic flux during maximum urease activity (reaction rxn00101, Urea amidohydrolase), showing large flux through the L-Arginine amidinohydrolase reaction (rxn00394).
- Matlab code
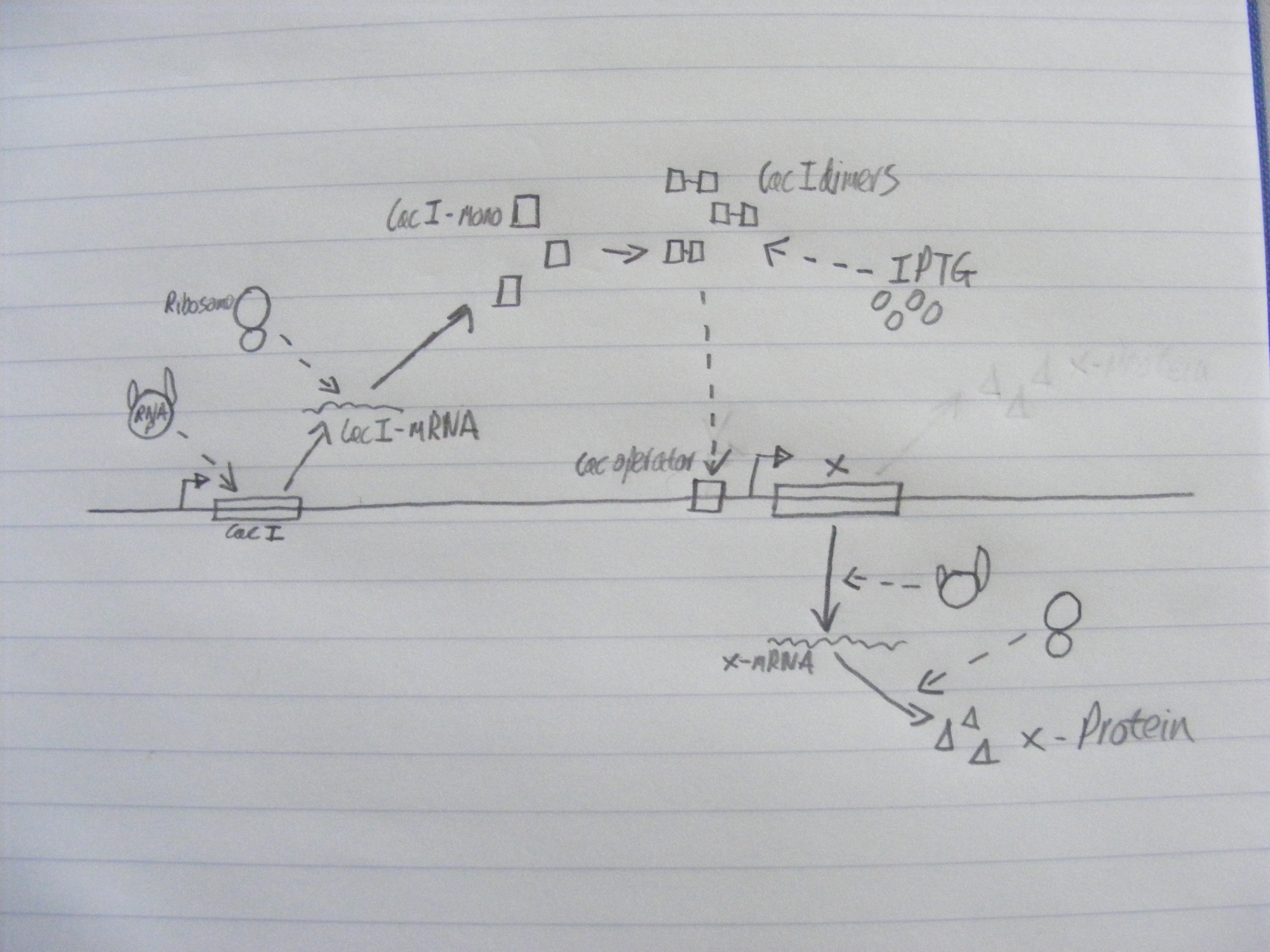
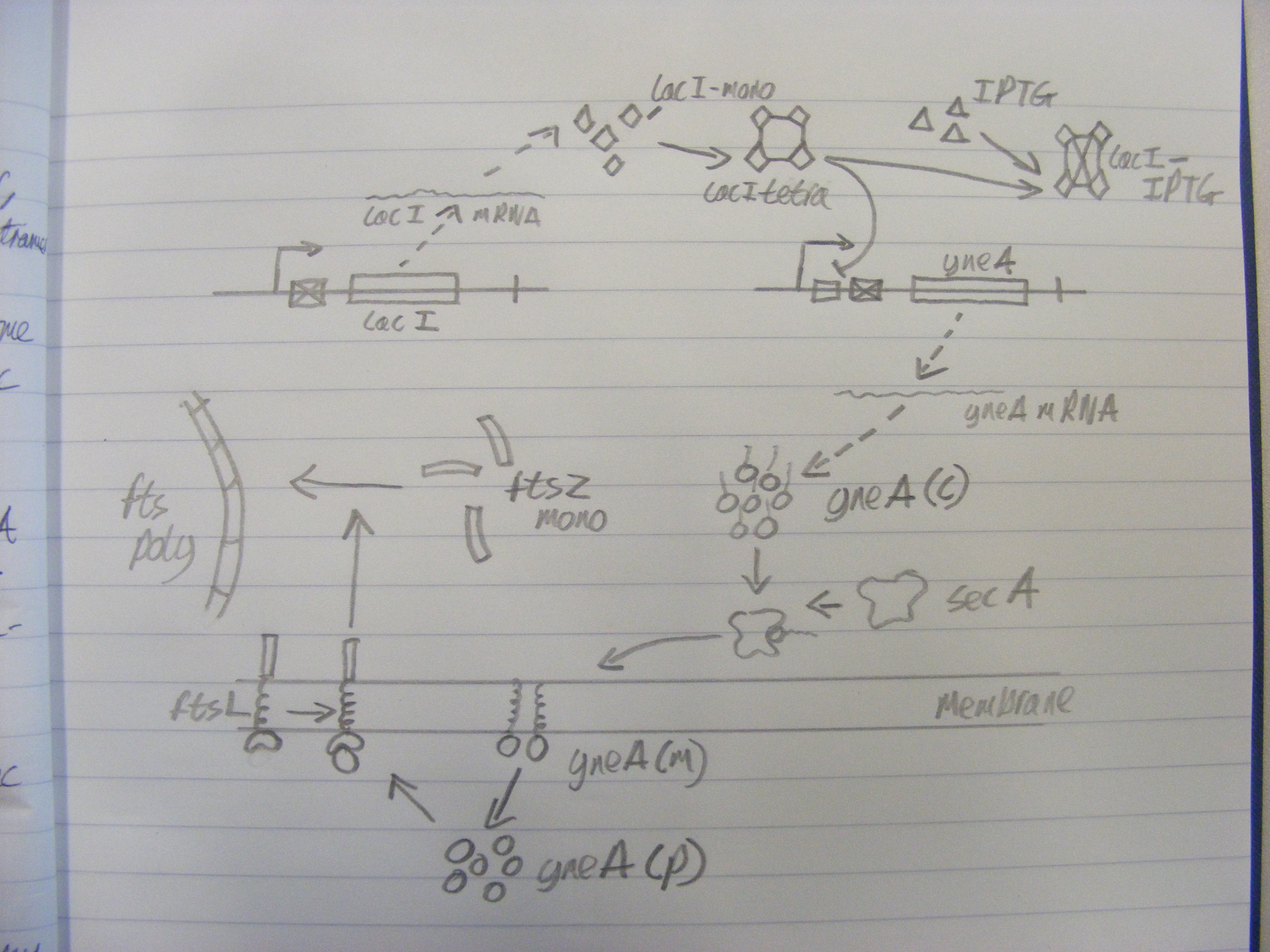
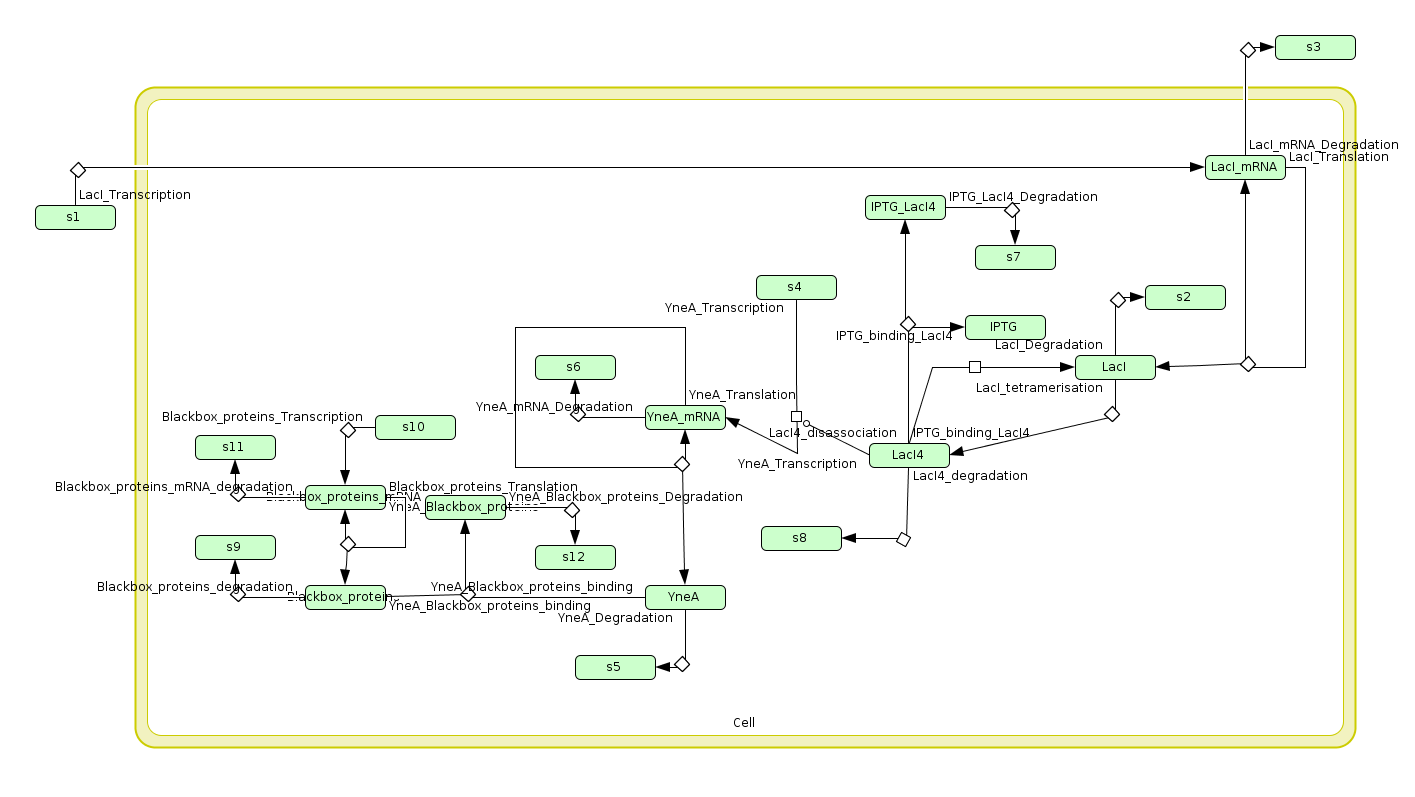
Biochemical network in SBML
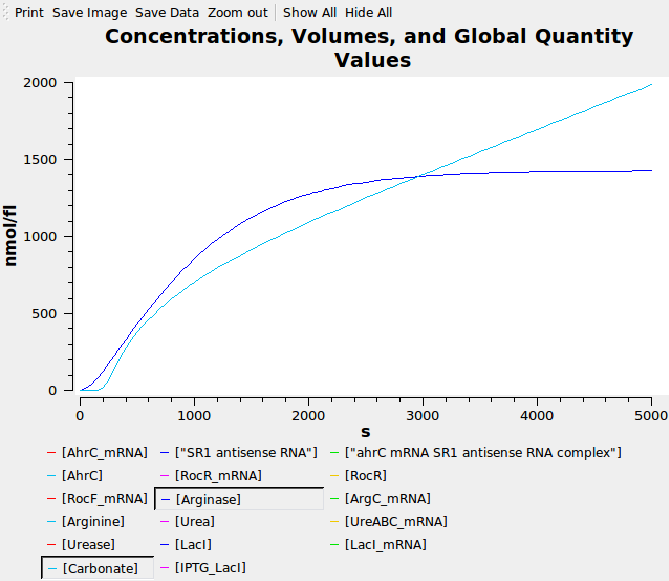
We wrote a computational model of our calcium carbonate system in SBML and simulated it in COPASI. The graph below shows that carbonate increases over time, as desired.
Downloads:
 
|
 "
"