Team:Heidelberg/Notebook/miMeasure/July
From 2010.igem.org
(New page: {{:Team:Heidelberg/Template}} {{:Team:Heidelberg/Pagetop|note_miMeasure}} = 05/07/2010 - 11/07/2010 = <br /> == Preperation of competent E. coli Top10 and DH5alpha == * plating of E. co...) |
|||
| (13 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | {{:Team:Heidelberg/ | + | {{:Team:Heidelberg/Single}} |
| + | {{:Team:Heidelberg/Single_Pagetop|note_miMeasure}} | ||
| + | {{:Team:Heidelberg/Side_Top}} | ||
| - | { | + | {| cellpadding="5" cellspacing="0" style="text-align: center; color:#4e93a4; border: 3px solid #4e93a4;" |
| + | |- border="0" | ||
| + | ! colspan="7" style="background:#4e93a4;" | [https://2010.igem.org/Team:Heidelberg/Notebook/miMeasure/August<font color="white">August</font>] | ||
| + | |- style="background:#4e93a4; color:white" | ||
| + | |width="20pt"|'''M'''||width="20pt"|'''T'''||width="20pt"|'''W'''||width="20pt"|'''T'''||width="20pt"|'''F'''||width="20pt"|'''S'''||width="20pt"|'''S''' | ||
| + | |- | ||
| + | |colspan="6"| ||'''1''' | ||
| + | |- | ||
| + | |'''2'''||'''3'''||'''4'''||'''5'''||'''6'''||'''7'''||'''8''' | ||
| + | |- | ||
| + | |'''9'''||'''10'''||'''11'''||'''12'''||'''13'''||'''14'''||'''15''' | ||
| + | |- | ||
| + | |'''16'''||'''17'''||'''18'''||'''19'''||'''20'''||'''21'''||'''22''' | ||
| + | |- | ||
| + | |'''23'''||'''24'''||'''25'''||'''26'''||'''27'''||'''28'''||'''29''' | ||
| + | |- | ||
| + | |'''30'''||'''31'''||colspan="5"| | ||
| + | |} | ||
| - | = 05/ | + | {| cellpadding="5" cellspacing="0" style="text-align: center; color:#4e93a4; border: 3px solid #4e93a4;" |
| - | < | + | |- border="0" |
| - | == | + | ! colspan="7" style="background:#4e93a4;" | [https://2010.igem.org/Team:Heidelberg/Notebook/miMeasure/September<font color="white">September</font>] |
| - | + | |- style="background:#4e93a4; color:white" | |
| - | + | |width="20pt"|'''M'''||width="20pt"|'''T'''||width="20pt"|'''W'''||width="20pt"|'''T'''||width="20pt"|'''F'''||width="20pt"|'''S'''||width="20pt"|'''S''' | |
| - | < | + | |- |
| - | == | + | |colspan="2"| ||'''1'''||'''2'''||'''3'''||'''4'''||'''5''' |
| - | + | |- | |
| - | : | + | |'''6'''||'''7'''||'''8'''||'''9'''||'''10'''||'''11'''||'''12''' |
| - | : | + | |- |
| - | : | + | |'''13'''||'''14'''||'''15'''||'''16'''||'''17'''||'''18'''||'''19''' |
| - | : | + | |- |
| - | < | + | |'''20'''||'''21'''||'''22'''||'''23'''||'''24'''||'''25'''||'''26''' |
| - | + | |- | |
| - | : | + | |'''27'''||'''28'''||'''29'''||'''30''' |
| - | + | | colspan="7"| <span style="color:#ffffff">-</span> | |
| - | + | |} | |
| - | + | ||
| - | ''' | + | {| cellpadding="5" cellspacing="0" style="text-align: center; color:#4e93a4; border: 3px solid #4e93a4;" |
| - | + | |- border="0" | |
| - | + | ! colspan="7" style="background:#4e93a4;" | [https://2010.igem.org/Team:Heidelberg/Notebook/miMeasure/October<font color="white">October</font>] | |
| - | + | |- style="background:#4e93a4; color:white" | |
| - | + | |width="20pt"|'''M'''||width="20pt"|'''T'''||width="20pt"|'''W'''||width="20pt"|'''T'''||width="20pt"|'''F'''||width="20pt"|'''S'''||width="20pt"|'''S''' | |
| - | : | + | |- |
| - | : | + | |colspan="4"| ||'''1'''||'''2'''||'''3''' |
| - | + | |- | |
| + | |'''4'''||'''5'''||'''6'''||'''7'''||'''8'''||'''9'''||'''10''' | ||
| + | |- | ||
| + | |'''11'''||'''12'''||'''13'''||'''14'''||'''15'''||'''16'''||'''17''' | ||
| + | |- | ||
| + | |'''18'''||'''19'''||'''20'''||'''21'''||'''22'''||'''23'''||'''24''' | ||
| + | |- | ||
| + | |'''25'''||'''26'''||'''27'''||'''28'''||'''29'''||'''30'''||'''31''' | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | {{:Team:Heidelberg/Side_Bottom}} | ||
| + | |||
| + | __NOTOC__ | ||
| + | |||
| + | ===05/07/2010 - 11/07/2010=== | ||
<br /> | <br /> | ||
| + | Preperation of competent E. coli Top10 and DH5alpha | ||
| + | Cell Culture Starting | ||
| - | = 12/07/2010 - 19/07/2010 = | + | ===12/07/2010 - 19/07/2010=== |
<br /> | <br /> | ||
| - | + | Preperation of RNA extracts for miRNA profiling | |
<br /> | <br /> | ||
*Hela, Hek and HUH were plated on p100 dishes in their according media (10E6 cells/dish) and grown to 60-70 % confluence | *Hela, Hek and HUH were plated on p100 dishes in their according media (10E6 cells/dish) and grown to 60-70 % confluence | ||
| Line 43: | Line 79: | ||
*pSB1AC3 from the registry was transformed into Top10 cells according to the standard transformation protocol; a day later, a 5 ml LB culture was inocculated and incubated for 8 hours; plasmid was extracted by applying the Qiagen MiniPrep Protocol | *pSB1AC3 from the registry was transformed into Top10 cells according to the standard transformation protocol; a day later, a 5 ml LB culture was inocculated and incubated for 8 hours; plasmid was extracted by applying the Qiagen MiniPrep Protocol | ||
<br /> | <br /> | ||
| - | + | design of diraPCR oligos | |
<br /> | <br /> | ||
* a diraPCR Designer tool (called miRACLE Designer) was programmed for enabling quick&easy design of diraPCR oligos. The program takes any microRNA guiding strand sequence of choice and constructs synthetic microRNA binding site oligos. Those oligos have the following properties: | * a diraPCR Designer tool (called miRACLE Designer) was programmed for enabling quick&easy design of diraPCR oligos. The program takes any microRNA guiding strand sequence of choice and constructs synthetic microRNA binding site oligos. Those oligos have the following properties: | ||
| Line 53: | Line 89: | ||
<br /> | <br /> | ||
| - | + | design of measurement standard | |
<br /> | <br /> | ||
* in order to design an appropriety standard respecting all the rules according to the standard of mammalian synthetic biology (RFC12), we developed a dual reporter measurment construct with the following properties: | * in order to design an appropriety standard respecting all the rules according to the standard of mammalian synthetic biology (RFC12), we developed a dual reporter measurment construct with the following properties: | ||
| Line 62: | Line 98: | ||
:* strong SV40 terminators behind both reporters | :* strong SV40 terminators behind both reporters | ||
:* vector backbone, containing an Amp resistance, Hygromycin resistance, pBR322 origin and an FRT site for stable integration | :* vector backbone, containing an Amp resistance, Hygromycin resistance, pBR322 origin and an FRT site for stable integration | ||
| - | |||
| - | = | + | ===20/07/2010=== |
<br /> | <br /> | ||
''' Dilution of raPCR-Oligos ''' | ''' Dilution of raPCR-Oligos ''' | ||
| Line 115: | Line 150: | ||
<br /> | <br /> | ||
| - | = | + | ===21/07/2010=== |
* optimization of cycle number and oligo concentration for raPCR | * optimization of cycle number and oligo concentration for raPCR | ||
| Line 187: | Line 222: | ||
<br /> | <br /> | ||
<br /> | <br /> | ||
| - | = 22/07/2010 = | + | |
| + | ===22/07/2010=== | ||
<br /> | <br /> | ||
* the PCR products from the previous days' raPCR were loaded on a 1 % agarose gel and the brightest 200, 400, 700 and 1400 bp bands were gel extracted using the Qiagen Gel Extraction Kit (elution in 32 ul of nuclease free water) | * the PCR products from the previous days' raPCR were loaded on a 1 % agarose gel and the brightest 200, 400, 700 and 1400 bp bands were gel extracted using the Qiagen Gel Extraction Kit (elution in 32 ul of nuclease free water) | ||
| Line 195: | Line 231: | ||
<br /> | <br /> | ||
<br /> | <br /> | ||
| - | = 23/07/2010 = | + | |
| + | ===23/07/2010=== | ||
<br /> | <br /> | ||
*transformation of ligation product (previous day) into E. coli Top10 cells according to the standard transformation protocol | *transformation of ligation product (previous day) into E. coli Top10 cells according to the standard transformation protocol | ||
<br /> | <br /> | ||
<br /> | <br /> | ||
| - | = 26/07/2010 = | + | |
| + | ===26/07/2010=== | ||
<br /> | <br /> | ||
* of 21 colonies, 7 of each hsa-mir-886_3p 200, 400 and 700 bp band cloning product and inocculation of Miniprep LB cultures | * of 21 colonies, 7 of each hsa-mir-886_3p 200, 400 and 700 bp band cloning product and inocculation of Miniprep LB cultures | ||
<br /> | <br /> | ||
| - | + | single binding site synthesis | |
<br /> | <br /> | ||
In order to construct a database of synthetic single binding sites, we developed and optimized a standardized single binding site synthesis protocol; | In order to construct a database of synthetic single binding sites, we developed and optimized a standardized single binding site synthesis protocol; | ||
| Line 233: | Line 271: | ||
<br /> | <br /> | ||
| - | = 27/07/2010 = | + | ===27/07/2010=== |
<br /> | <br /> | ||
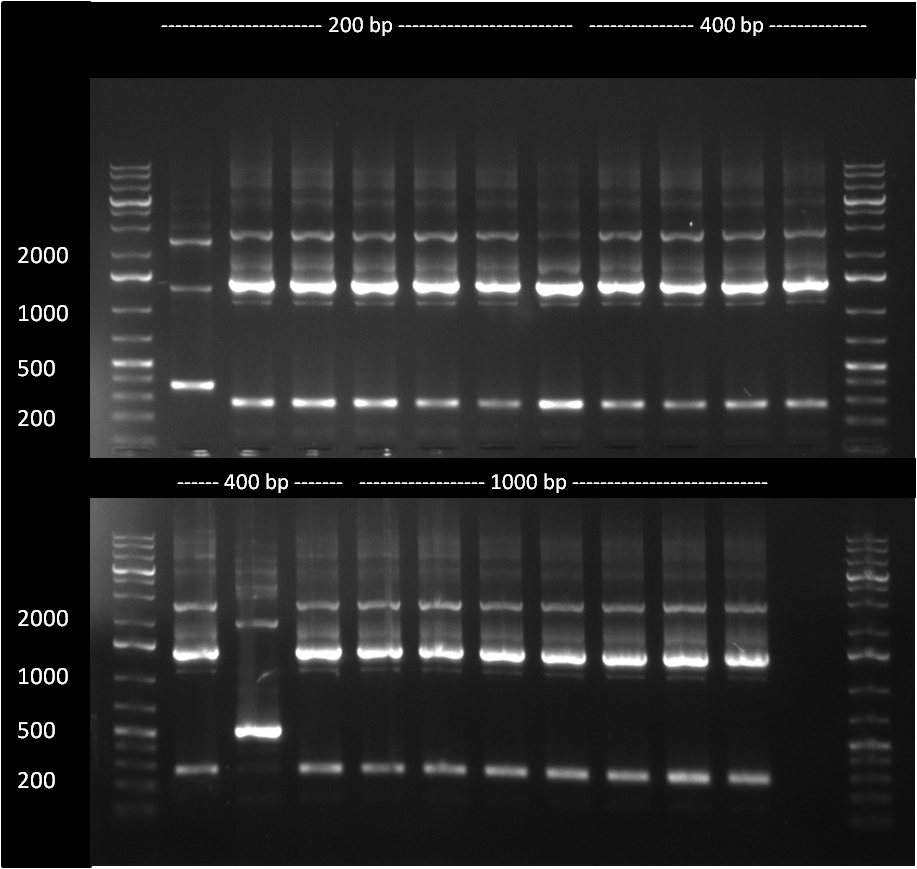
[[Image:886colonyPCR.png|thumb|500 px|right|The 200, 400 and 1000 bp cloning products were analyzed in an analytic PCR. Only The very first sample of the 200 bp product seems to be positive, as it shows a higher band (~ 300 bp) compared to the religated vector product band at ~ 200 bp]] | [[Image:886colonyPCR.png|thumb|500 px|right|The 200, 400 and 1000 bp cloning products were analyzed in an analytic PCR. Only The very first sample of the 200 bp product seems to be positive, as it shows a higher band (~ 300 bp) compared to the religated vector product band at ~ 200 bp]] | ||
| Line 264: | Line 302: | ||
| - | + | Single Binding Site Synthesis - Optimization | |
*Further optimization of the single binding site synthesis protocol | *Further optimization of the single binding site synthesis protocol | ||
[[Image:singlebs2.png|thumb|400 px|right|The unspecific or multimer side products were clearly reduced, but there is still sime unwanted, longer product in case of mir-122.]] | [[Image:singlebs2.png|thumb|400 px|right|The unspecific or multimer side products were clearly reduced, but there is still sime unwanted, longer product in case of mir-122.]] | ||
| Line 288: | Line 326: | ||
<br /> | <br /> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<br /> | <br /> | ||
| - | |||
<br /> | <br /> | ||
| - | |||
| - | |||
| - | |||
| - | |||
<br /> | <br /> | ||
| - | |||
| - | |||
| - | |||
| - | |||
<br /> | <br /> | ||
| - | |||
| - | + | ===31/07/2010=== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | = 31/07/2010 = | + | |
<br /> | <br /> | ||
* ready-to-use vector pSB1A3 from the registry (25 ng/ul) digestion with EcoRI/PstI (NEB Buffer EcoRI + BSA, volume: 40 ul) | * ready-to-use vector pSB1A3 from the registry (25 ng/ul) digestion with EcoRI/PstI (NEB Buffer EcoRI + BSA, volume: 40 ul) | ||
| Line 355: | Line 353: | ||
* no digestion product was detectable on the gel; therefor raPCR was repeated | * no digestion product was detectable on the gel; therefor raPCR was repeated | ||
<br /> | <br /> | ||
| - | + | diraPCR for constructing hsa-886-3p binding site patterns | |
<br /> | <br /> | ||
* the diraPCR was pipetted according to the following protocol in three repeats: | * the diraPCR was pipetted according to the following protocol in three repeats: | ||
| Line 377: | Line 375: | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | {{:Team:Heidelberg/ | + | {{:Team:Heidelberg/Single_Bottom}} |
Latest revision as of 00:54, 27 October 2010

05/07/2010 - 11/07/2010
12/07/2010 - 19/07/2010
Protocol
design of measurement standard
20/07/2010
................................................
................................................ (7x)
................................................
................................................
................................................ (25x)
................................................
................................................
21/07/2010
................................................
................................................ (12x)
................................................
................................................
................................................
................................................ (3x)
................................................
................................................
................................................ (25x)
................................................
................................................
22/07/2010
23/07/2010
26/07/2010
PCR protocol:
................................................
................................................ (30x)
................................................
................................................
27/07/2010
................................................
................................................ (30x)
................................................
................................................
31/07/2010
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"