Project/PlcR Promoter/
From 2010.igem.org

Contents |
PlcR Promoter - [http://partsregistry.org/Part:BBa_K354003 BBa_K354003]
Background
Sequence Development and Annotation
In the same way that we assembled the Lac/Ara-1 promoter from primer-primer adhesion and their subsequent extension, the PlcR promoter was assembled in the same fashion. See the Lac/Ara -1 Promoter for the full assembly details.
Promoter Assembly
The forward primer used in assembly is:
The sequence in red is the EcoR1 restriction site with that in blue being the Xba1 restriction site. The region in purple are the bases that the forward and reverse primers overlap with one another.
The reverse primer used in assembly is:
The sequence in green is a Pst1 restriction site and that in orange is the Spe1 restriction site. The region in purple is the sequence of shared overlap with the forward primer. As can be seen, there have been randomised nucleotides introduced so as to create a library of varying promoters. This was performed so as to increase the stochastic nature associated with the promoter efficacy. According to the IUPAC nomenclature, Y indicates either a C or a T while W indicates either an A or a T.
The entire PlcR promoter sequence is:
As can be seen, the 'g' in red resulted from the insertion of a C and the 't' in green from the insertion of an A.
The Package
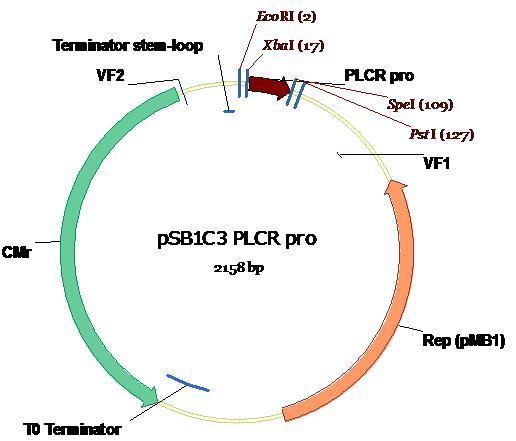
The plasmid map, annotated with appropriate restriction sites and sizes depicts the PlcR promoter ligated into PSB1C3.
The following confirmation of the successful ligation of the PlcR promoter into PSB1C3 is in a 2% (m/v) agarose gel.
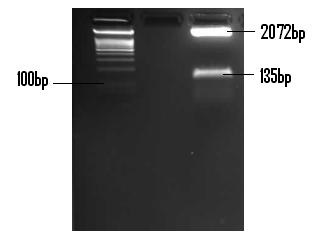
The above gel indicates the PlcR promoter (135bp) and the linearised PSB1C3 plasmid (2072bp). The fragments were isolated by digestion with EcoR1 and Pst1.
Testing and Validation
References
 "
"