Team:SDU-Denmark/project-p
From 2010.igem.org
Contents |
Our Parts
<groupparts>iGEM010 SDU-Denmark</groupparts>
Characterization of parts
K343007
The part K343003 (from now on shortly called PS) is a generator for the SopII-HtrII photosensor from Natronomonas Pharaonis coupled to E.Colis chemotaxis pathway via the Salmonella protein Tar. This part's effect on the system is to make E.Coli phototactic, so that it becomes aware of different light conditions. So for characterization of this part we made examination of motility and motility patterns our first priority and plasmid stability and growth of the cells our second.
There is a wide range of motility assays for chemotaxis in bacteria, which meant that we had a broad spectrum of experiments to choose from, which just had to be tweaked for making them suited for the analysis of phototaxis. The two experiments we chose for analysing the effect of this part (PS), were growth of the bacterial cultures in semi-solid agar and computer analysis of swimming motility through video microscopy:
1. Growth of bacterial culture on semi-solid agar plates (Experiment 1):
The bacteria and plates were prepared after our own protocol, which you can find here: [1]
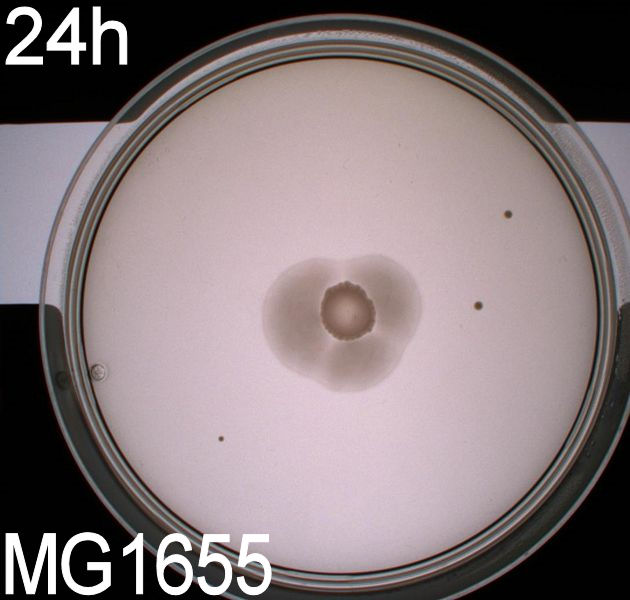
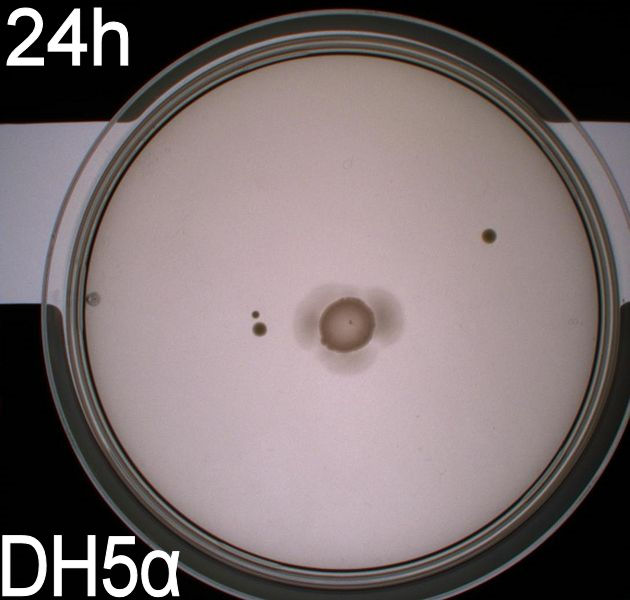
Since exposure to blue light should decrease the phototaxic bacterias tumbling frequency, the expected result was that the colony which was placed between the light and dark half of the plate would spread out in the darkness and would not move further when it reached the light. This is counter-intuitive, since decreased tumbling should lead to a longer distance traveled. What happens at the microscopic scale in semisolid agar is that the agar creates a matrix like structure where there are channels through the agar, which the bacteria can swim through. The decrease in tumbling frequency of the bacteria will make it harder for them to find the channels in the agar to swim through, which leads to them being trapped where they were placed. The result is that a colony which shows an increased run time, will look as if it it was non-motile on these plates. Our results showed exactly this; the bacterial culture had spread out to on the dark half of the plate and did not get nearly as far on the half exposed to light. This experiment was done with a normal wildtype MG1655 and a non-motile strain of E.coli, DH5alpha, as controls. As expected these cells did not show anything like the behavior described above, which indicates that the effect stems from the modification to our photosensor bacteria. These results were useable, but not fully conclusive, since there were some non-optimal conditions present in this experiment. We used ambient light instead of pure blue light, and the exposure to light for the multiple samples was not exactly even. Therefore we had to improve our experiment setup and see if we could reproduce these results with a more reliable setup.
2. Growth of bacterial culture on semi-solid agar plates (Experiment 2):
The experiment described above was repeated in a more controlled environment. This means that there were no changes as to how the plates or bacterial cultures were prepared, so we refer again to [2] in regards to how this was done. The difference lies in the setup of the light-controlled environment. What we did this time was that the plates was illuminated from above by a single light source in an otherwise completely dark environment, we prepared a cut-out so that the light would only hit one half of our plates and the other half would remain in the dark.
Another deviation from the protocol PS1.1 is that the culture was inoculated at the center of the plate, instead one colony each was inoculated in the center of the light exposed and dark half respectively. This would give us more information on how the bacteria would behave when directly exposed to either the dark or light surroundings, instead of the gradient that was present in the first experiment.
From the results of the first experiment we expected the culture containing BBa_K343007 to spread out in the dark and not to spread when exposed to light. The wildtype bacteria should spread out evenly no matter if exposed to light or not and the non-motile strain (DH5alpha) should not move regardless of the light conditions. Our expectations from the first experiment were fulfilled, as the bacteria behaved exactly as expected. This made it possible to conclude that the photosensor has an effect on the bacteria's tumbling frequency, but if it does in fact reduce the tumbling is not possible to say, since both an increased and reduced tumbling frequency will look alike.
After this we could conclude that the part had an impact on the bacterial motility, but we had to find out which. This lead us to our next experiment, which was intended for determining what happened to the tumbling frequency:
2. Videomicroscopy and computer analysis:
To get an exact idea of what is happening to the bacteria when adding the photosensor we decided on doing video microscopy of motile bacteria. The bacterial cultures for this analysis were incubated at 37° C, 180 RPM overnight, afterwards the optical density at 550nm was measured and the cultures were diluted to an OD550 of 0,02. They were then again incubated at 37 degrees and 180 RPM until their optical density reached 0,5. At this point we added retinal to the culture at a concentration of 5 uM and incubated the cells in the same conditions for another two hours. On top of phototactic bacteria we cultivated MG1655 (wildtype) and DH5alpha (non-motile) as controls. These were grown until an OD550 of 0,5 with the same methods as described for the modified culture, at this point these strains were ready for microscopy.
The actual microscopy was done on a (insert name) microscope with an optical magnification of 1000x.
To be able to observe an effect in the modified bacteria we had to expose the cultures to light, which was done through the microscope's integrated light source, which could switch between blue, green and ambient light. To further make sure that the blue light was at the correct wavelength the same optical filter that was used for the plate experiments was inserted between the lightsource and the microscopy slide. Since microscopy is impossible without light, we had to use red light as a replacement for darkness, the source of the red light was ambient light from the microscope's light source filtered through a red optical filter (about 580nm). This should not have any effect on the phototactic bacteria according to Jung et. al (insert reference!). The microscopic slides were prepared by adding 5 ul of bacterial culture in the center of a glass microscopy slide, dimensions 7,5cm x 1,5cm, at last a cover slip was used to cover the culture. This was then sealed with ordinary nail polish to eliminate any flow in the system, which could be mistaken for bacterial motility. The room where the experiment was carried out was kept as dark as possible, as to eliminate the chance of the phototactic bacteria being unwantedly exposed to light. The actual experiment was carried out in the following manner:
1. Expose bacteria to red light for 1 minute, while recording video.
2. Move to another spot on the slide, repeat 1. (repeated until 5 videos were recorded)
3. Expose bacteria to blue light for 1 minute, while recording video.
4. Wait for 30 seconds, for the bacteria to reset themselves.
5. Move the focus to another spot on the slide, repeat 3. (x5)
This was done with all three strains of bacteria.
An example of the results:
When played at real time speed these videos seem slow and not showing any tendency, though when played at 4 times the normal speed, the photosensor will seem to show slightly increased motility when exposed to blue light. Even though the bacteria displayed disappointing motility, we went on and did computer analysis on the videos by the help of the open source software "CellTrack" (Link!). The paths that were mapped showed a longer distance traveling for the photosensor exposed to blue light, when compared with the rest of the samples. This could indicated a decreased tumbling frequency, but since the bacteria's motility was very low, the results were stamped as unreliable and a new protocol had to be devised.
Computerized analysis of the bacterial motility with the Thor prototype by Unisensor A/S and the Unify software:
For this experiment we changed our protocol for cultivating swimming bacteria.
After a series of experiments we settled on a pipette tip from a freezing culture inoculated in 5 ml LB media. The culture was then incubated for 12 hours at 22° C and 160 RPM. Afterwards the unmodified cultures are ready for microscopy. At this point retinal is added to the strain containing the photosensor and it continues to grow for another two hours in darkness, at which point these bacteria also are ready for analysis.
The machine and software used for the analysis are prototypes and still under heavy development. Because of trade secrets it is impossible for us to explain how the machine works or give a detailed explanation of it's mechanism until the Thor has reached production status. Simplistically said it is a very advanced video microscope, with which it is possible to analyse liquids and the particles inside in both 2D and 3D, while also tracking them over time.
The data collection and preparation of microscopy slides, was basically done like explained above, except that instead of light filters we used diodes that already sent out light at the correct wavelengths.
The results look like these:
(YOUTUBE)
The data analysis is still ongoing.
K343004
The FlhDC operon is the master regulator of flagella synthesis. A more detailed description of the operon can be found here
In our system the purpose of the composite part is to hyper flagellate our cells so that a grater force can be generated in the microtubes. The FlhDC operon is naturally found in the E. coli strain MG1655 genome. We extracted the operon and inserted a silent mutation (T to C) at position 822 in the operon because without the mutation this site is a Pst1 digestion site and it would therefor constitute problems when assembling the composit part.
We have made three FlhDC parts:
[http://partsregistry.org/Part:BBa_K343100 K343100] is the coding sequence of the native FlhDC operon with the Pst1 digestion site
[http://partsregistry.org/Part:BBa_K343000 K343000] is the coding sequence of the mutated FlhDC operon
[http://partsregistry.org/Part:BBa_K343004 K343004] is the composite part containing the TetR repressable promoter (constitutive when no TetR is pressent) + RBS (J13002), the K343000 part and the double terminator (B0015).
The composite part is caracterized firstly by using a motility assay and secondly by measuring plasmid stability and cell growth.
1. Motility assay (Experiment 1 with WT and negative control):
Growth of bacterial culture on semi-solid agar plates
The pictures below show the negative control DH5alpha in the upper left picture, the wild type MG1655 in the upper right picture, MG1655 transformed with pSB1C3-K343004 in the lower left picture and MG1655 transformed with pSB3K3-K343004 in the lower right picture.


From the pictures above we can definately see that the bacteria containing our part is much more motile than the wild type. We assume thise is caused by overexpression of the FlhDC master flagella operon which leads to hyperflagellation of the cells.
The two buttom pictures show that bacteria with pSB1C3-K343004 have not moved as far as the bacteria containing pSB3K3-K343004. pSB1C3 is a high copy plasmid while pSB3K3 is a low-medium copy plasmid. The promoter in K343004 is a constitutive promoter (tetR repressable promoter). Bacteria containing a high copy plasmid with a constitutive promoter are more metabolically challanged than bacteria containing a low- or medium-copy plasmid with a constitutive promoter because of the higher number of plasmids per the cell. Therefore the high copy plasmid bacteria are less motile than low- or medium-copy plasmid bacteria.
2. Motility assay (doublicate with WT, negative- and positive-control):
Growth of bacterial culture on semi-solid agar plates
3. Stability assay
4. Growth measurment
 "
"