Team:Imperial College London/Results
From 2010.igem.org
| Results |
Contents |
Cat-O2-lase Protein
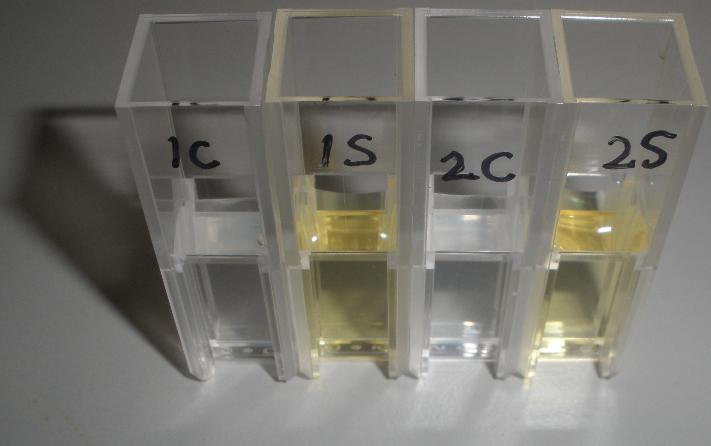
The XylE gene, originating from the TOL plasmid of Pseudomonas putida, was the choice for the output signal mediator of our system. The system is based on catechol 2,3-dioxygenase enzyme. Colonies of cells that express xylE gene product become yellow within seconds after selection plates are sprayed with catechol, a colorless substrate, that is converted by CatO2ase to the yellow product, 2-hydroxymuconic semialdehyde. This reporter system is ideal for our project as it has very fast kinetics, gives a visual output by naked eye and the substrate used is very cheap.
Characterization of CatO2lase protein
Experiment 1
The CatO2lase protein, except as an ideal output signal for our engineered bacterial detector it can also serve as a great reporter gene. Its characteristics can be exploited in a wide range of fields across biological sciences field, so it was one of the team's first candidates for further characterization. Qualitative and quantitative data were gathered and presented. All further tests involving XylE transformed cells were quantified by measuring absorbonce at 380nm wavelenth.
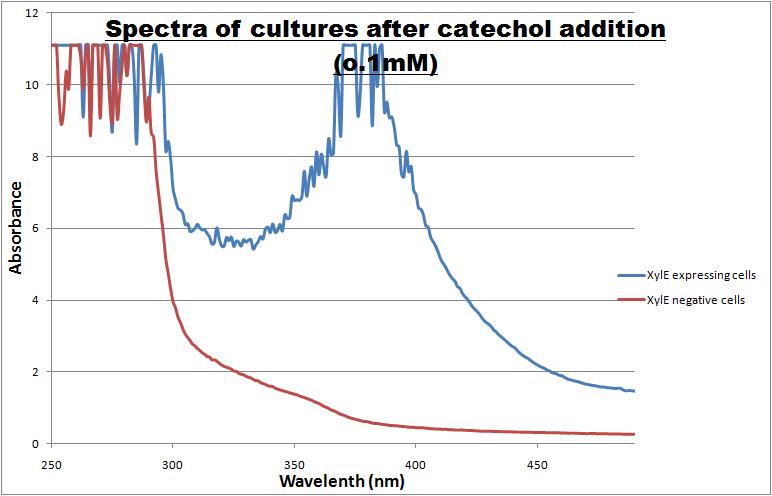
Catechol ,the initial substrate of CatO2las enzyme, is colourless. However within seconds of its addition the colonies/liquid cultures of XylE expressing cells become yellow, indicating production of product which absorbs light in the region visible spectrum. An spectrophotometric assay was prepared, where the spectra of two cultures of E.coli (one XylE gene transformed and the other not)were compared on addition of 0.1mM Catechol substrate. The spectra (figure 1) showed that in XylE transformed cells, a broad peak appears at about 380nm. The absorbance of the particular wavelengths of light by the product of the enzymatic reaction 2-hydroxymuconic semialdehyde, is what causes the yellow output.
References
- Chromogenic identification of genetic regulatory signals in Bacillus subtilis based on expression of a cloned Pseudomonas gene. Zukowski, M.M., Gaffney, D.F., Speck, D., Kauffmann, M., Findeli, A., Wisecup, A., Lecocq, J.P. Proc. Natl. Acad. Sci. U.S.A. (1983)
Experiment 2
In our Parasight project the XylE gene product, C(2,3)lase enzyme, is the means by which an output signal is generated. If the parasite is present is the water supply, the bacterial system is activated by an input signal which produces the yellow color, the positive output signal. Test assays in the lab on colonies of cells expressing XylE gene, become yellow only seconds after addition of catechol (100mM). From this it was realized our group that the kinetics of the C(2,3)lase enzyme are such that make it an attractive candidate as a reporter enzyme. So, we decided that further characterization of the kinetics of the C(2,3)lase might help identify a really efficient reporter for research centers.
In this experiment we investigated how the velocity of the reaction (catechol converted to 2-hydroxymuconic semialdehyde by C(2,3)lase) varies with increasing initial concentrations of catechol, the substrate. Overnight cultures of XylE transformed E.coli were plated on a 96 wells plate at an optical density (O.D.) of 0.5 units of absorbance. (The optimum initial O.D. of cells was determined in a previous experiment. This O.D. may vary across different labs as the sensitivity of the spectrophotometer/plate reader is a factor). Catechol concentrations ranging from 0.1-1mM were made from 100mM stock solution and added to the cells of the 96-well plate. The velocity of the reaction was monitored using the plate reader spectrophotometer as the reaction is directly proportional to the production of yellow color. The yellow output was measured at 380nm wavelength, while cell density at 600nm. The XylE gene expression was under the influence of J23101 promoter.
Experiment 3
In paper blabla.... In our construct, GFP was added to the N-terminal of C(2,3) monomer. C(2,3) enzyme is a homotetramer were all 4 subunits are required for its activity. We hoped that by adding GFP at the N-terminus of the catechol dioxygenase enzyme, we would block tetramerization of the enzyme, thus render it inactive. This modification is important as it makes our system inducible. The cleavage of the GFP by TEV protease, on arrival of input signal, means that Catechol dioxygenase becomes active only when the parasite is detected.
To investigate this, an experiment was designed where catechol was added to XylE transformed E.coli. The velocity of the reaction was monitored at various initial catechol concentrations. The experimental set up is essentially the same as described above for determining the kinetics of C(2,3)O enzyme. The only difference is that E.coli cells used are the ones transformed with GFP-XylE gene construct instead of XylE gene.
Experiment 4
threshold value of colour output
 "
"