Team:Freiburg Bioware/NoteBook/Labjournal/October2
From 2010.igem.org
- March (labday 1)
- April (labday 2 - 5)
- May (labday 6 - 17)
- June (labday 18 - 45)
- July (labday 46 - 75)
- August part 1 (labday 76 - 92)
- August part 2 (labday 93 - 106)
- September part 1 (labday 107 - 123)
- September part 2 (labday 124 - 135)
- October part 1 (labday 136 - 145 )
- October part 2 (labday 146 - 155 )
- October part 3 (labday 156 - 166 )
- November (labday 167 - 170 )
- Cellculture
146. labday 11.10.2010
Colony PCR of Cloning VP2 Fusion and Super constructs into pSB1C3
Investigator: Achim, Hanna
Comment: Because yesterday's cloning didn't deliver a good separartion of the expected gel bands. Nevertheless ligation and trafo was performed. In order to immediately find out, whether we received successful results a colony PCR will be performed.
Two clones were picked from each plat. In addition to that a positive (pAAV_RC) and a negative control (pSB1C3_lITR) was prepared.
Used primer: 4200 rev and Cap3500 for. Expected fragment size: 885 bp.
PCR was performed following the standard protocol.
All samples match the positive control!
To do: Mini-Prep and sequencing.
Preparation of SDS-PAGE gel (10%)
Investigator: Hanna
Comment: In order to perform a Western Blot of different virus capsids (with and without capsid-motifs), 2 10% SDS polyacrylamid gels were prepared.
Resolving gel: 15 mL
- H2O: 5.9 mL
- Acryl-bisacrylamide mix (30%): 5 mL
- Tris (1.5 M, pH 8.8): 3.8 mL
- SDS (10%): 0.15 mL
- Ammonium persulfate (10%): 0.15 mL
- TEMED: 0.006 mL
Stacking gel (5%): 5 mL
- H2O: 3.4 mL
- Acryl-bisacrylamide mix (30%): 0.83 mL
- Tris (1.5 M, pH 6.8): 0.63 mL
- SDS (10%): 0.05 mL
- Ammonium persulfate (10%): 0.05 mL
- TEMED: 0.005 mL
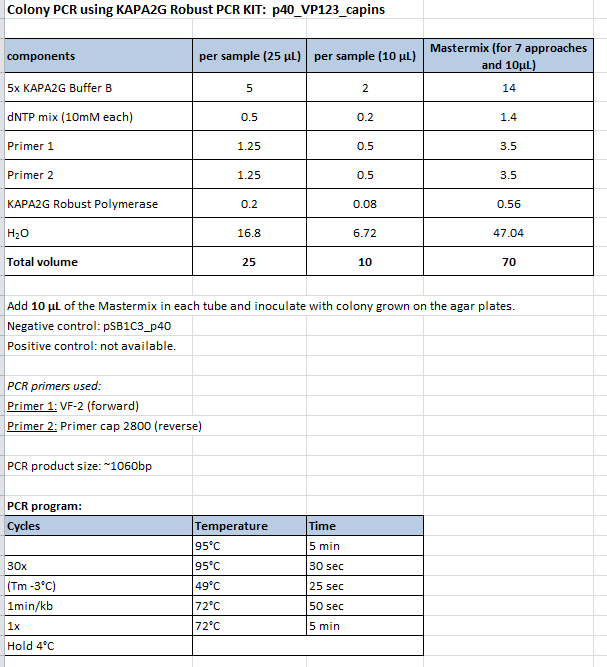
Colony PCR of p40_VP123_capins
Investigator: Bea
Comment: Since the first attempt did not work,and no cells grew on the plate and the same ligation was transformed again into BL-21 and a lot of clones grew on the plate, I decided to perform a colony PCR in order to check several colonies and to inoculate at the same day for a Midi-Prep.
Protocol:
- Primer used: O162
- Primer used: O38
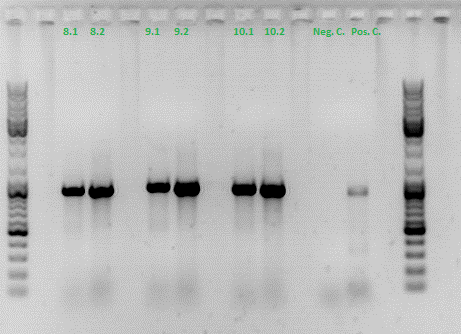
The PCR products were loaded on a 1% agarose gel. The results can be seen above in the gel picture:
Result: We can see two things: The cloning of p40 to VP123 worked quiet well AND the Robust PCR Kit which was used for the first time worked as well.
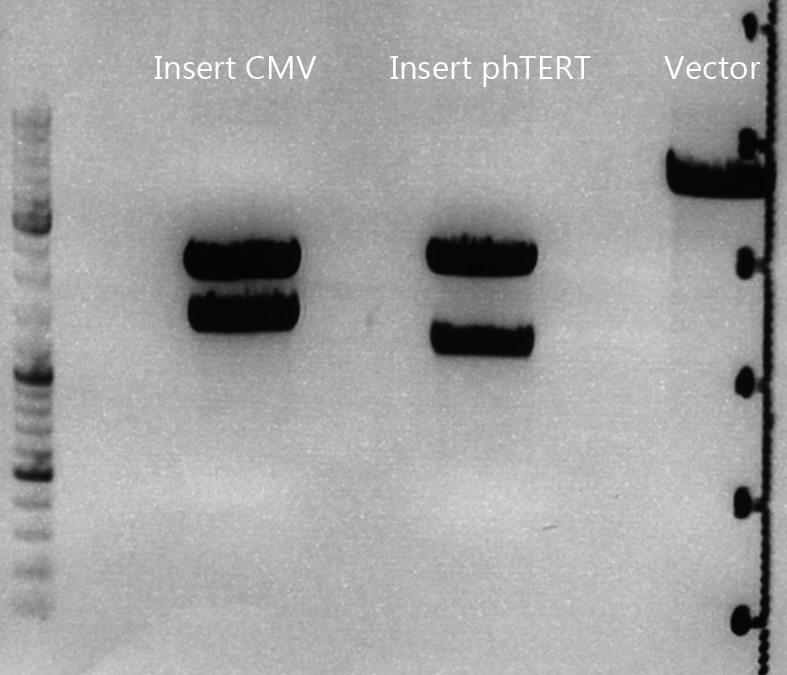
Cloning of lITR_CMV_betaglobin and lITR_phTERT_betaglobin into pSB1C3_CD
Investigator: Stefan
Comment: To produce another GOI for testing in cell culture, the cytosine deaminase needs to be assembled with lITR_promotor_betaglobin. In the next step hgH_rITR needs to be added.
Vector name:
pSB1C3_CD clone 1
pSB1C3_CD clone 2
Insert name:
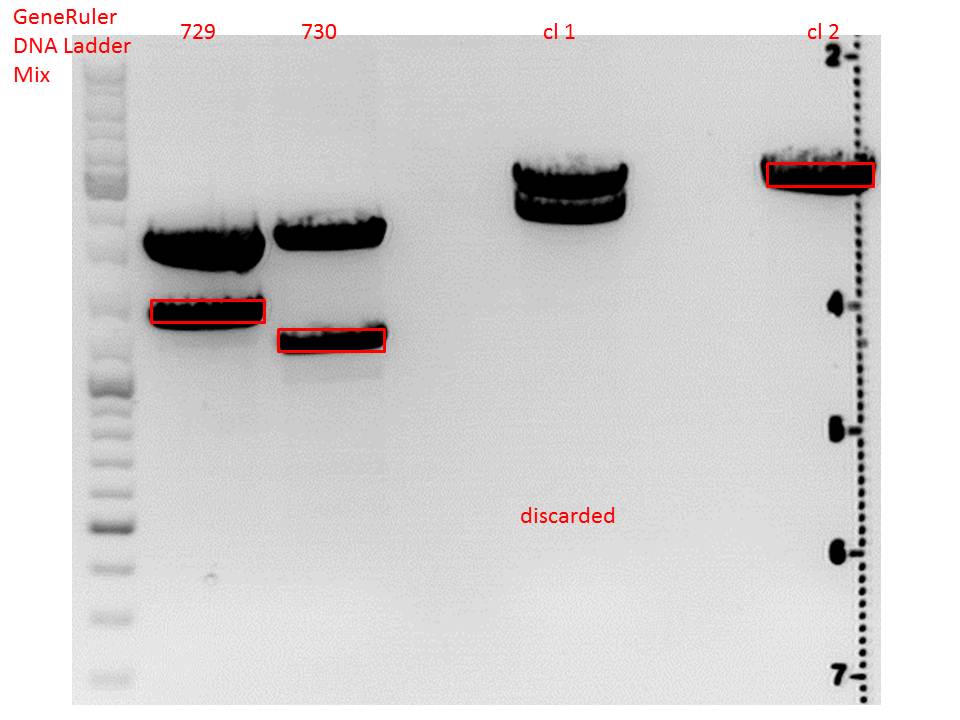
pSB1C3_lITR_CMV_beta-globin (P729)
pSB1C3_lITR_phTERT_beta-globin (P730)
Digestion:
| components | volume CD clone 1 + 2 /µl | volume P729 /µl | volume P730 /µl |
| DNA | 6 | 14 | 6 |
| BSA (10x) | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 |
| Enzyme EcoI | 1 | 1 | 1 |
| Enzyme XbaI | 1 | - | - |
| Enzyme SpeI | - | 1 | 1 |
| H2O | 8 | - | 8 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 | 20 |
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
CD clone 1 yielded to bands around 2500 bp to 3000 bp. Since the vector was cut only using EcoRI and SpeI, it was expected to be linearized, not to be cut into two fragments this size. Therefore, this sample was discarded and cloning was continued using CD clone 2.
Gel extraction:
Was performed according to protocol.
T4 Ligation:
| ligation name | 729 + CD cl2 | 730 + CD cl2 |
| volume of vector | 3,67 | 2,82 |
| volume of insert | 4,33 | 5,18 |
| T4 ligase buffer (10x) | 1 | 1 |
| T4 ligase | 1 | 1 |
Transformation:
Was performed according to standard protocol using BL21 cells.
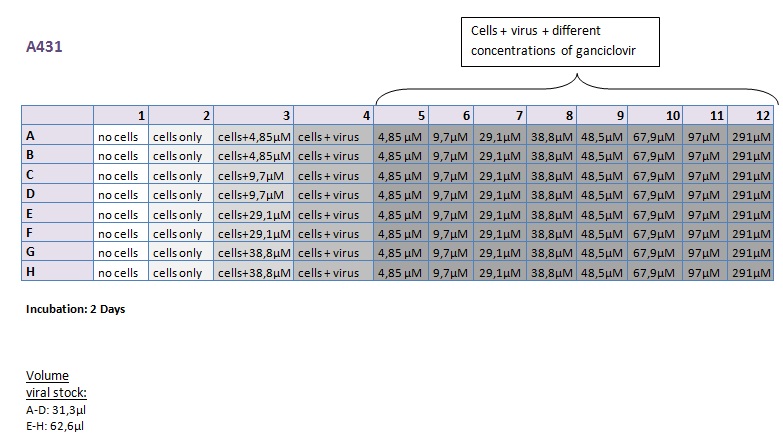
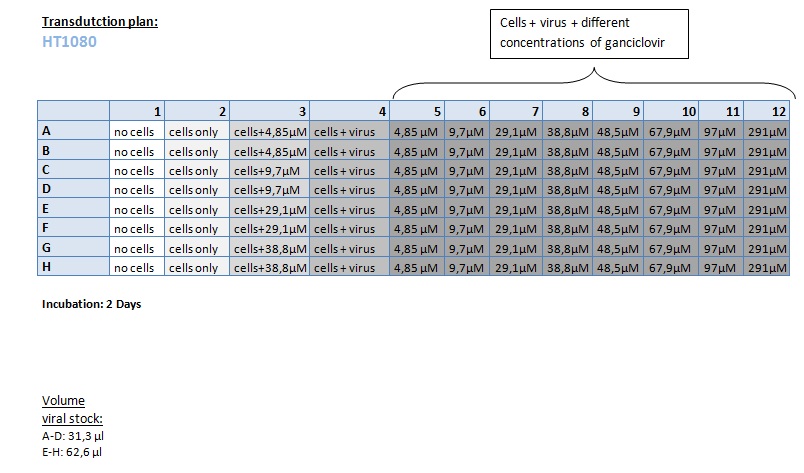
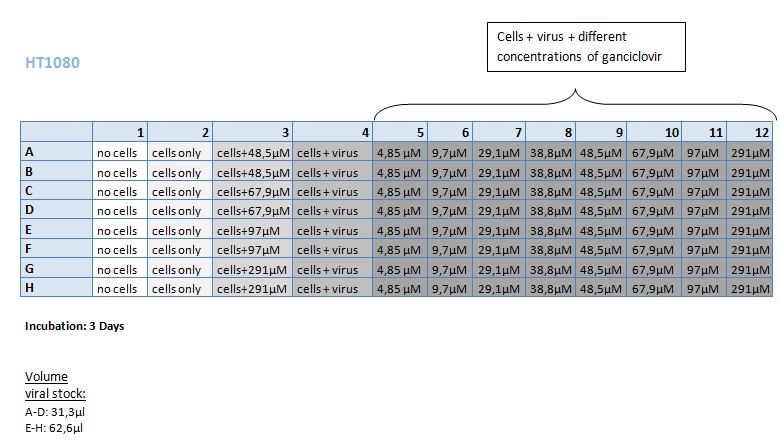
Seeding HT1080 and A431 for testing different concentrations of ganciclovir by MTT-Assay
Investigator: Kerstin, Anissa
- Seeding 4x 96-well plates: 2x HT1080 and 2x A431
FACS-Analysis
Investigator: Kerstin
...
Preparation of the ELISA
Investigator: Volker
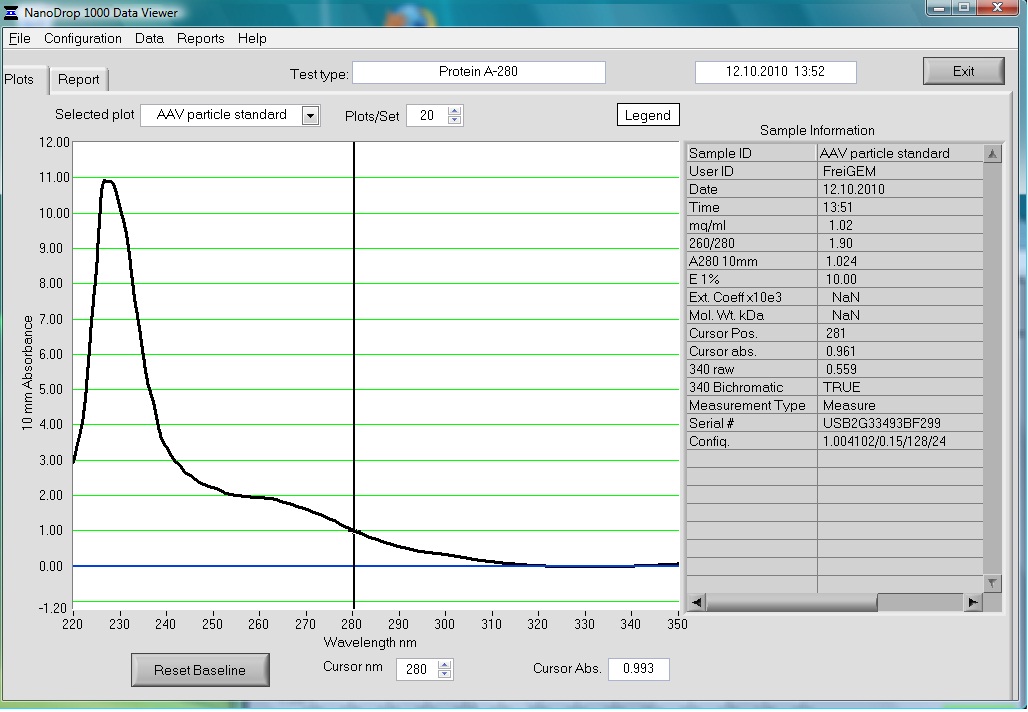
The AAV particle standard that contains 3.6x10^9 viral particles was dissolved in 500µl as described in the protocol of the Progen AAV Titration ELISA and a absorption spectrum was measured.
These purified and concentrated viral particles could be used for biophysical measurements, there for the possibility to detect the viral particles by absorption was interesting for us.
The Spectrum measured in the NanoDrop is the following:
Trafo evaluation of Rep52
Investigator:Kira
T4 ligation seems to worked out. The plate contained many colonies.
147. labday 12.10.2010
Sequencing analysis: pSB1C3_mGMK_TK30 and pSB1C3_CD
Investigator: Stefan
Test digestion of pSB1C3_CD and pSB1C3_CD_SDM-PstI_new
Investigator: Stefan
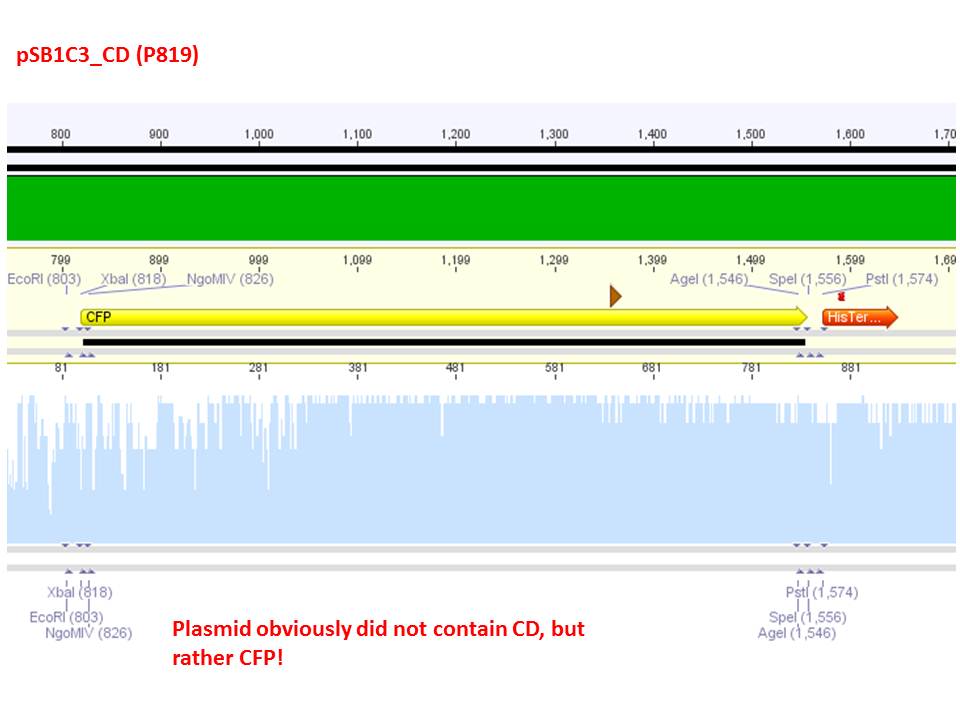
Comment: Sequencing results revealed that there is CFP rather than CD in P819. In order to verify this and exclude the possibility that the wrong plasmid was sent for sequencing test digestions were performed. Additionally the original vector at which the SDM to remove the PstI was performed was digested as well to verify its insert.
| Components | digest 1 / µl | digest 2 / µl |
| DNA | 1 | 7 |
| Buffer 4 | 1 | 1 |
| BSA (10x) | 1 | 1 |
| Enzyme I | SpeI 0,3 | PstI 0,3 |
| Enzyme II | XbaI 0,3 | XbaI 0,3 |
| H2O | 6,4 | 0,4 |
| Total volume | 10 | 10 |
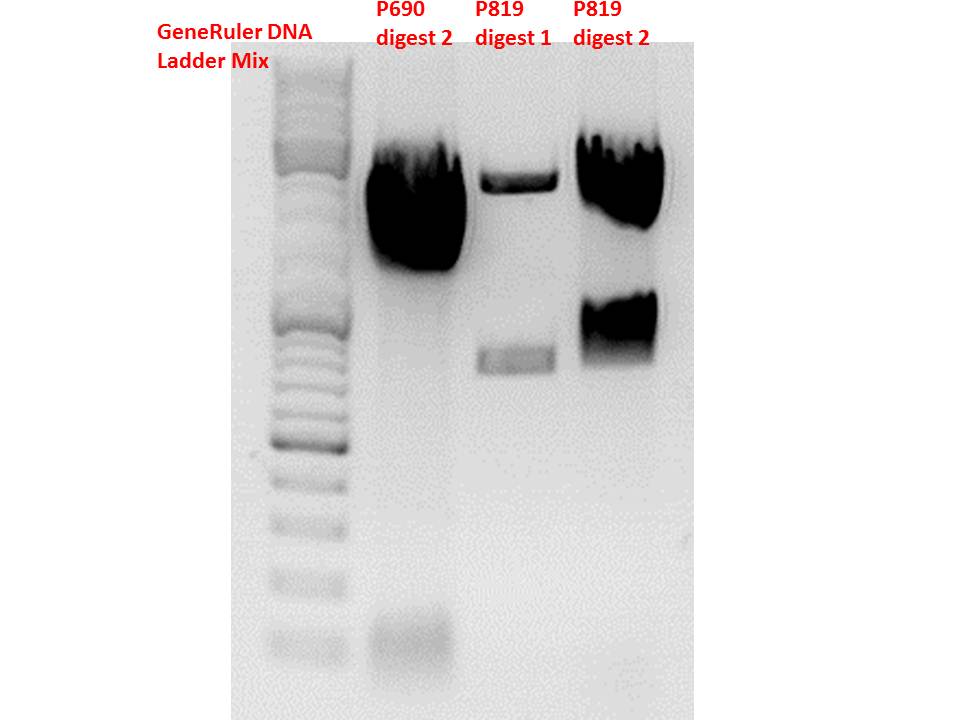
Comment: Using these digestion approaches, it was expected for CFP to deliver always bands at ~750 bp. For the CD it was expected to show bands at 90 bp, 1200 bp and 2000 bp if the CD still contains the PstI restictiton site, 1300 bp and 2000 bp if it was removed.
Gel:
0,5g agarose, 50 ml TAE (1%), 3 µl GELRED, 115 Volt, running time ~50 minutes
Comment: As it can be seen, P819 does contain CFP, therefore a mistakenly sequenced plasmid can be excluded. Also it can be seen that P690 contains the CD. Therefore an additonal SDM of P690 to remove the PstI restriction site is necessary.
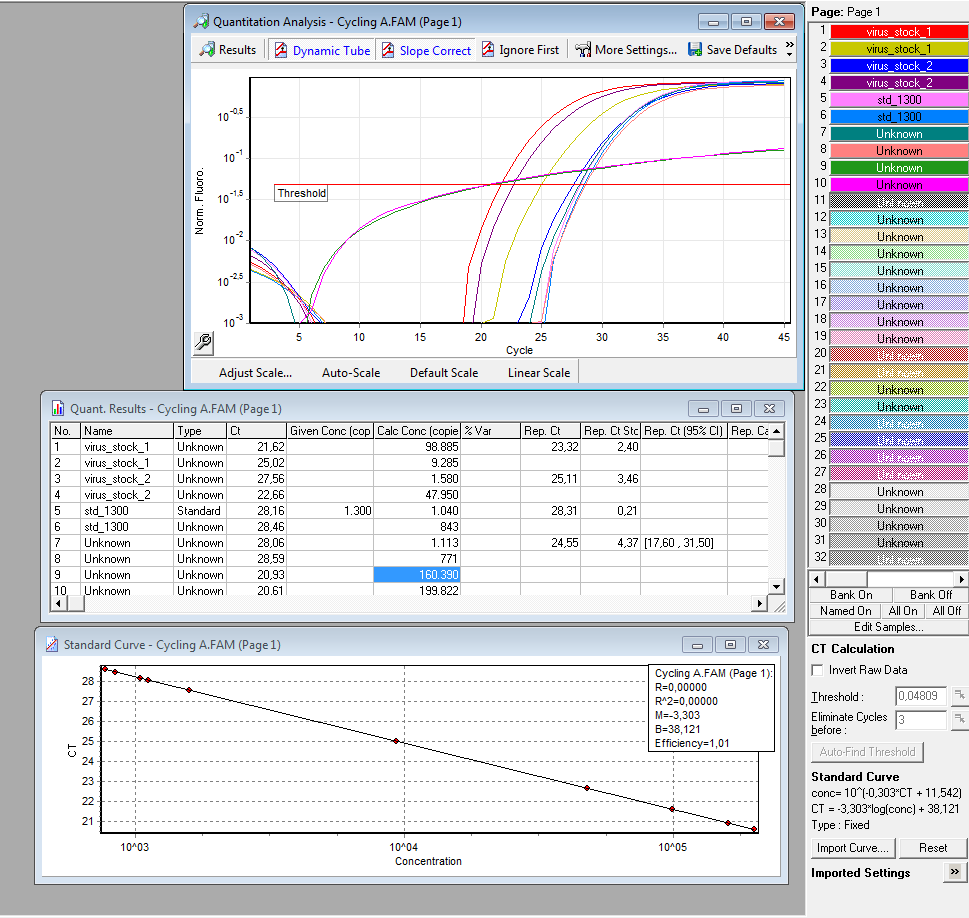
qPCR of virus stock: P816/ TKGMk-Standard; R/C: P468 from 5.10. (VLP harvested by Adrian)
Investigator: Achim
Protocol for quantitative real-time PCR of virus particles
Date: 12.10.
Achim
Virus Stock: A.F.; P816; TKGMK-Standard; R/C: P468; 5.10; consists of supernatant from pelleted cell fragments.
Following the Protocol used by (Rohr et al., 2002).
I digested 5 µl virus dilution with 7,5 µl DNAse I (7,5 units) and 25 µl 50 mM MgCl2 (end concentration 25 mM) in a final volume of 50 µl at 37°C for 30 min.
è DNAse should be heat inactivated at 65°C for 10 min. I forgot that step. The enzyme should however be inactivated in the initial denaturation step.
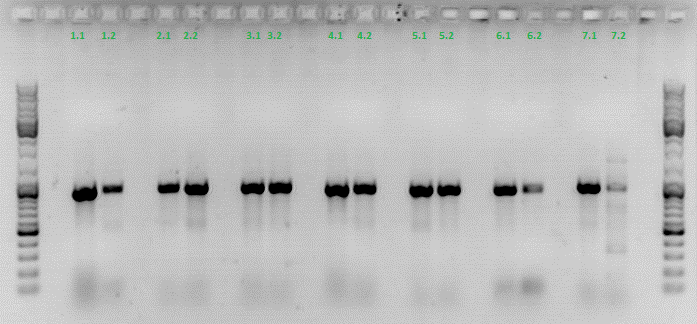
I prepared the following PCR reactions:
|
Sample |
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
|
9 |
10 |
|
|
1.1 |
1.2 |
2.1 |
2.2 |
S.1 |
S.2 |
NP1 |
NP2 |
MM for samples 1 -8 |
NH1 |
NH2 |
|
PCRmix |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
112.5 |
12.5 |
12.5 |
|
Primer for |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
9 |
- |
- |
|
Primer rev |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
9 |
- |
- |
|
Template |
2 |
2 |
2 |
2 |
5 |
5 |
- |
- |
- |
- |
- |
|
H2O |
8.5 |
8.5 |
8.5 |
8.5 |
5.5 |
5.5 |
10.5 |
10.5 |
- |
12.5 |
12.5 |
|
Total |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
|
25 |
25 |
|
Step |
Time |
Temp |
|
Initial denaturation |
7’ |
95°C |
|
|
|
|
|
Denaturation |
10’’ |
95°C |
|
Annealing/Extension |
30’’ |
60°C |
|
Number of Cycles |
45 |
|
Results:
è Both negative controls containing primers were contaminated with DNA.
è Negative controls containing only H2O were negative.
è Values for plasmids/reaction vary; from 1 – 100 for the same virus stock
è Dilution of the virus solution: 1:10; 2µl PCR sample: 3.2 * 10^5 virus particles/ml
è Because of the contaminated negative controls and the high deviation, I will repeat the experiment tomorrow to verify the results.
References:
Rohr, U., Wulf, M., Stahn, S., Steidl, U., Haas, R., Kronenwett, R., et al. (2002). Fast and reliable titration of recombinant adeno-associated virus type-2 using quantitative real-time PCR. Journal of virological methods, 106(1), 81-8. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/12367732.
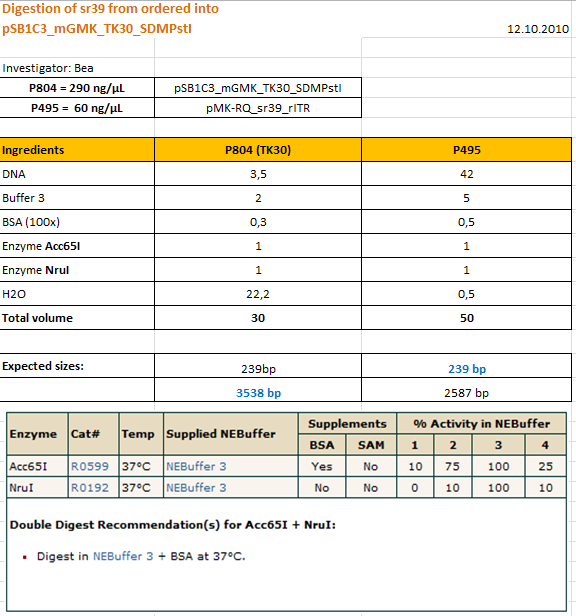
Cloning sr39 into mGMK_TK30
Investigator: Bea
Comment: We do not only want to submit the tk30, a mutant version of the thymidine kinase, but as well another mutant protein which is named sr39. SR39 seems to have a better activity than the tk30 without being fused to the guanylate kinase (mgmk), but the sr39 also works quite well as a fusion protein. We ordered one part of the sr39 and we are now subcloning it into the mgmnk_tk30 construct in order to obtain mgmk_sr39.
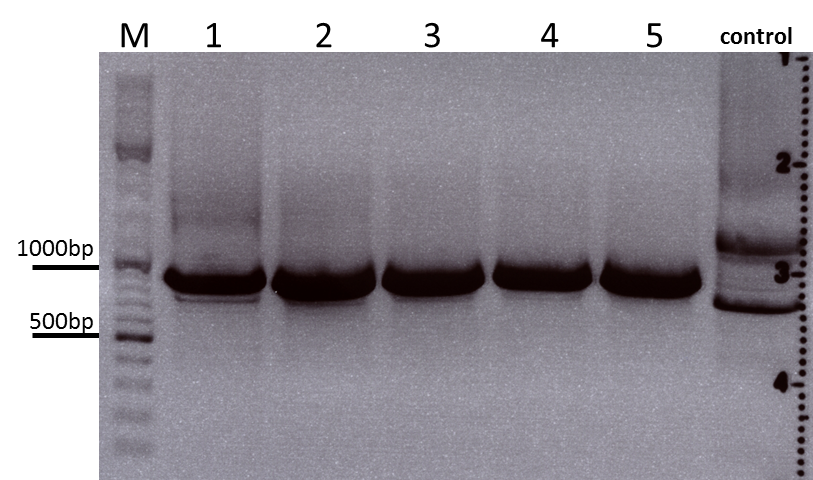
Protocol:
The digested fragments were loaded on a 1.2% agarose gel. The results can be seen above in the gel picture: I cut out the small visible band in the right lane, and the upper intensive band of the left lane.
Result:
After gel extraction has been performed, I ligated the two fragments and transformed BL-21 cells, plated them on cm agar plates and incubated them over night at 37°C.
Concentration of virus stock for Western Blot analysis
Investigator: Hanna
For the first Western Blot try, we investigate whether is is enough to concentrate the virus stock (from cell supernatant) and then to simply load it onto a SDS-PAGE gel and perfom immuno-staining.
For this purpose 8 mL virus stock (pHelper, pAAV_RC, GOI=YFP) were concentrated via amicon filtration.
- First, amicon filter was washed with 4 mL PBS.
- Then 4 mL virus stock was loaded onto the filter and centrifuged at 3000 rpm for 15 minutes.
- Flow-through was discarded and additional virus stock was loaded, centrifuged at 3000 rpm for 25 minutes.
- Flow-through was discarded, remaining solution was re-suspended and additional virus stock was added. Amicon filter was centrifuged at 3000 rpm for 50 minutes.
- Flow-through was discarded. Residual protein solution: 500 µL. Filter was washed with 3 mL PBS, centrifuging 35 minutes two times.
- 2 x 40 µL protein solution (16x) was taken and mixed with Laemmli buffer and stored over night @ 4°C.
To do: SDS-PAGE and Western Blot tomorrow.
Perfomance of another try by harvesting viruses of the cell pellet with RIPA buffer & freeze/ thaw.
Western Blot of unmodified, and modified capsids (N-terminal fusion and VP1 insertion with CFP ref. mVenus) --> size shift should be detectable.
Transduction of HT1080 and A431
Investigator: Kerstin
Mini-Prep and test digestion of several constructs
Investigator: Stefan
Glycerol stocks were prepared:
- B651 = pSB1C3_P40_VP123
- B652 = pSB1C3_lITR_phTERT_ß-globin_mGMK_TK30_PstI_SDM_hgH_rITR
- B653 = pSB1C3_lITR_phTERT_ß-globin_mGMK_TK30_PstI_SDM_hgH_rITR
- B654 = pSB1C3_lITR_CMV_ß-globin_mGMK_TK30_hgH_rITR
- B655 = pSB1C3_lITR_CMV_ß-globin_mGMK_TK30_hgH_rITR
- B656 = pSB1C3_CMV_VP123_453_Z34C_HSPG-KO
- B657 = pSB1C3_CMV_VP123_453_Z34C_HSPG-KO
- B658 = pSB1C3_CMV_VP123_453_RGD_HSPG-KO
- B659 = pSB1C3_CMV_VP123_453_RGD_HSPG-KO
- B660 = pSB1C3_lITR_phTERT_ß-globin_CFP_hgH-rITR
- B661 = pSB1C3_lITR_phTERT_ß-globin_CFP_hgH-rITR
Mini-Prep was performed according to standard protocol:
- P821 = pSB1C3_P40_VP123 c = 327,4 ng/µl
- P822 = pSB1C3_lITR_phTERT_ß-globin_mGMK_TK30_pSTI_SDM_hgH_rITR c = 342,9 ng/µl
- P823 = pSB1C3_lITR_phTERT_ß-globin_mGMK_TK30_pSTI_SDM_hgH_rITR c = 324,0 ng/µl
- P824 = pSB1C3_lITR_CMV_ß-globin_mGMK_TK30_hgH_rITR c = 81,6 ng/µl
- P825 = pSB1C3_lITR_CMV_ß-globin_mGMK_TK30_hgH_rITR c = 148,4 ng/µl
- P826 = pSB1C3_CMV_VP123_453_Z34C_HSPG-KO c = 105,8 ng/µl
- P827 = pSB1C3_CMV_VP123_453_Z34C_HSPG-KO c = 121,1 ng/µl
- P828 = pSB1C3_CMV_VP123_453_RGD_HSPG-KO c = 188,4 ng/µl
- P829 = pSB1C3_CMV_VP123_453_RGD_HSPG-KO c = 140,4 ng/µl
- P830 = pSB1C3_lITR_phTERT_ß-globin_CFP_hgH-rITR c = 197,2 ng/µl
- P831 = pSB1C3_lITR_phTERT_ß-globin_CFP_hgH-rITR c = 207,3 ng/µl
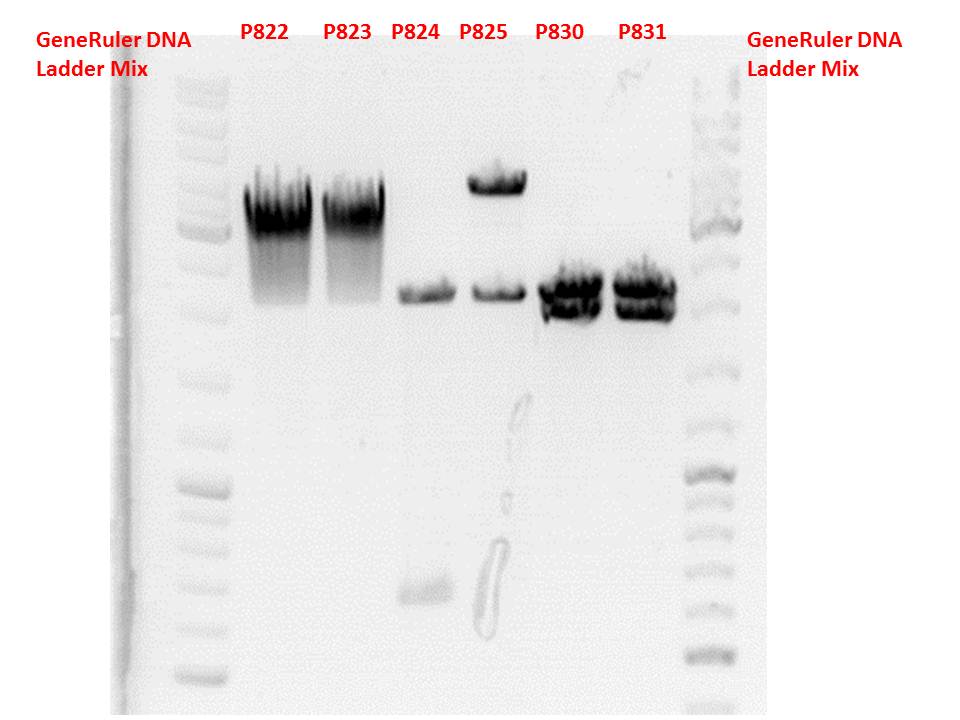
Test digestion:
| Components | P822 + P823 / µl | P824 + P825 / µl | P830 + P831 / µl |
| DNA | 1,5 | 1,5 | 2 |
| Buffer 4 | 1 | 1 | 1 |
| BSA (10x) | 1 | 1 | 1 |
| PstI | 0,3 | - | - |
| XbaI | 0,3 | 0,3 | 0,3 |
| SpeI | - | 0,3 | 0,3 |
| H2O | 5,9 | 5,9 | 5,4 |
| Total volume | 10 | 10 | 10 |
Gel:
0,5g agarose, 50 ml TAE (1%), 3 µl GELRED, 115 Volt, running time ~50 minutes
Comment: P822 and P823 smear a lot, therefore evaluation is difficult. Another test digestion would be reasonable.
P824 contains hgH_rITR, therefore it will be discarded. P825 looks well and was sent for sequencing.
P830 and P831 look well and will be sent for sequencing tomorrow.
Repetiton: Site-directed mutagenesis in pSB1C3_CD construct
Investigator: Stefan
Comment: Sequencing revealed that there is no CD in our pSB1C3_CD but rather CFP. So, anonther approach of the SDM needs to be performed.
Protocol using the QuickChange Lightning Kit from Stratagene:
Ingredients:
| Ingredients | Volume / µl |
| 10x reaction buffer | 2,5 |
| dNTP | 0,5 |
| forward primer: O191 | 0,56 |
| reverse primer: O192 | 0,57 |
| DNA Template | 0,5 (1:10 dilution) |
| QuikSolution Reagent | 0,75 |
| QuikChangeLightning Enzyme | 0,5 |
| H2O | 14,35 |
| Total volume | 25 |
The following PCR program was used:
- 95 °C 120 s
- 95 °C 20 s
- 60 °C 60 s
- 68 °C 105 s
- Goto step 2 repeat 17 times
- 68 °C 300 s
Digestion using DpnI:
To digest methylated and hemi-methylated DNA, 1 µl of DpnI was added and the mixture was incubated for 10 minutes at 37 °C.
Transformation:
Trafo was performed according to protocol, but using XL1b cells instead of XL-10 Gold cells.
148. labday 13.10.2010
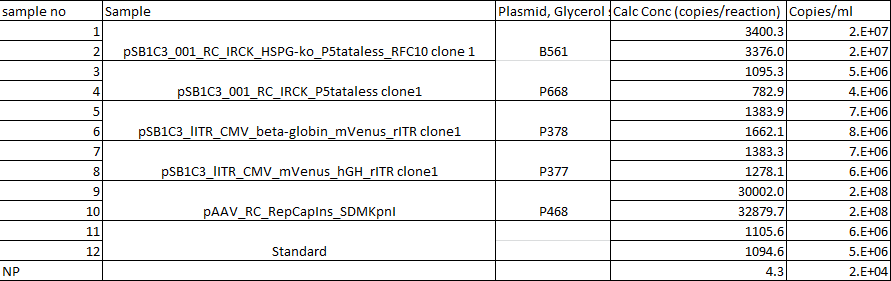
qPCRs of virus stocks
Investigator: Achim
|
Sample: |
Plasmid, Glycerol stock: |
Sample No. |
|
pSB1C3_001_RC_IRCK_HSPG-ko_P5tataless_RFC10 clone 1 |
B561 |
1 |
|
pSB1C3_001_RC_IRCK_P5tataless clone1 |
P668 |
2 |
|
pSB1C3_lITR_CMV_beta-globin_mVenus_rITR clone1 |
P378 |
3 |
|
pSB1C3_lITR_CMV_mVenus_hGH_rITR clone1 |
P377 |
4 |
|
pAAV_RC_RepCapIns_SDMKpnI |
P468 |
5 |
Digestion: 37°C, 30 min
|
|
1 |
2 |
3 |
4 |
5 |
MM x6 |
|
Virus solution |
5 |
5 |
5 |
5 |
5 |
|
|
DNAseI (7,5 U) |
7,5 |
7,5 |
7,5 |
7,5 |
7,5 |
45 |
|
MgCl2 50 mM |
5 |
5 |
5 |
5 |
5 |
30 |
|
H2O |
32.5 |
32.5 |
32.5 |
32.5 |
32.5 |
195 |
|
Total |
50 |
50 |
50 |
50 |
50 |
300 |
PCR reaction:
|
Sample |
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
11 |
12 |
13 |
14 |
|
15 |
16 |
|
|
1.1 |
1.2 |
2.1 |
2.2 |
3.1 |
3.2 |
4.1 |
4.2 |
5.1 |
5.2 |
S.1 |
S.2 |
NP1 |
NP2 |
MM for samples 1 -14 (x15) |
NH1 |
NH2 |
|
PCRmix |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
12.5 |
187,5 |
12.5 |
12.5 |
|
Primer for 1:10 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
15 |
- |
- |
|
Primer rev 1:10 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
15 |
- |
- |
|
Template |
2 |
2 |
2 |
2 |
2 |
2 |
2 |
2 |
2 |
2 |
5 |
5 |
- |
- |
- |
- |
- |
|
H2O |
8.5 |
8.5 |
8.5 |
8.5 |
8.5 |
8.5 |
8.5 |
8.5 |
8.5 |
8.5 |
5.5 |
5.5 |
10.5 |
10.5 |
- |
12.5 |
12.5 |
|
Total |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
25 |
|
25 |
25 |
Results:
SDS Page of unmodified AAV2 capsids
Investigator: Hanna
Comment: Yesterday a 8x concentration of AAV2 supernatant-stock was performed. 10 µL Laemmli-buffer was added to 40 µL AAV stock (8x) and stored over night at 4°C.
Today a SDS-PAGE of these samples will be performed.
Samples were incubated for 10 minutes at 95°C for denaturation.
Loading plan:
| 5 µL PageRuler Prestained Protein Ladder | 10 µL Sample | 15 µL Sample | 25 µL Sample |
After running front reached the end of the gel, PAGE was stopped.
Western Blot:
Three layers of transfer buffer soaked whatman paper were placed onto the blotter. A polyvinylidene difluoride membrane was activated by methanol (5 seconds) and then incubated for 5 minutes in transfer buffer. Membrane was then placed onto the whatman paper, gel was carefully layed on top. Air bubbles were removed, ensuring elictricity flow. After coverage by 3 further layers of whatman, blotting was performed at 200 mA for 1 hour.
Blocking:
In order to minimize nonspecific antibody interactions, membrane was incubated in blocking buffer (TBS (pH 7.5), 5% milk powder) for one hour, shaking.
Primary antibody incubation:
A 1:10 dilution of the B1 antibody in TBS + 1% milk was prepared. Membrane was incubated in 50 mL falcon over night at 4°C, rotating.
Midi-Prep
Investigator: Chris W.
Midi-Prep of:
pSB1C3_CMV_VP123_453_Z34C =P832 =B656
pSB1C3_CMV_VP123_453_RGD_HSPG-KO =P833 =B658
pSB1C3_001_RC_IRCK_HSPG-ko_P5tataless_RFC10 clone 1 =P834 =B561
pSB1C3_001_RC_IRCK_P5tataless clone 1 =P835 =B516
pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone1 =P836 =B200
pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone1 =P837 =B200
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | P832 | P833 | P834 | P835 | P836 | P837 |
| concentration (ng/µl) | 3615,8 | 1504,7 | x | x | x | x |
Comment: P834-837 given to sven
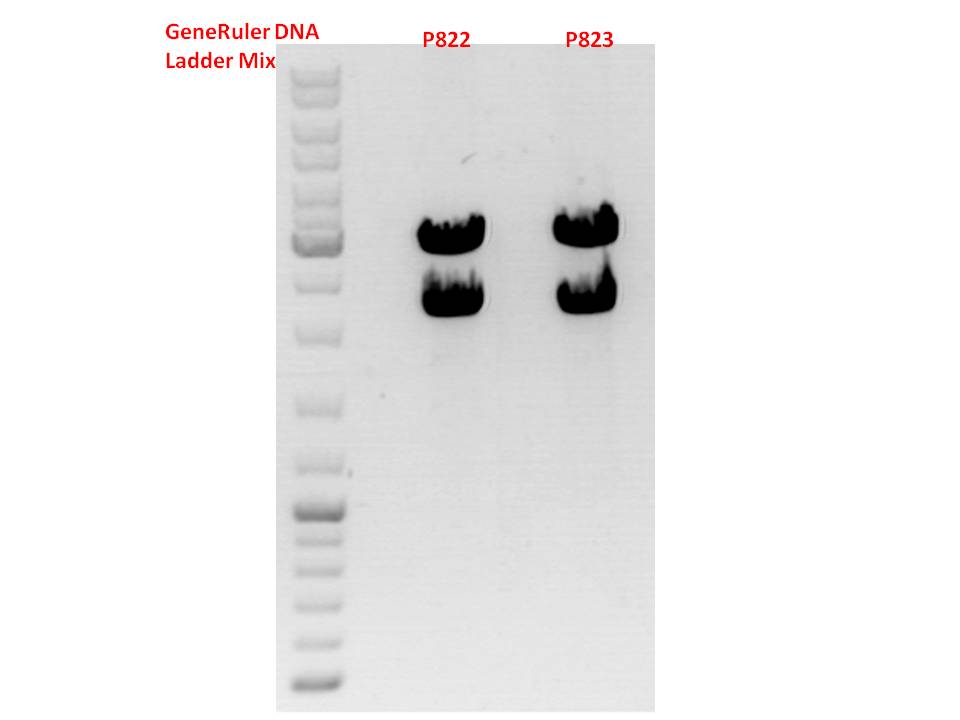
Repetiton: Test digestion of pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_SDM-PstI_hgH_rITR
Investigator: Stefan
Comment: Repetiton of yesterday's test digestion which yielded no clear band at 2000 bp as it was expected.
| Components | P822/P823 / µl |
| DNA | 1,5 |
| Buffer 4 | 1 |
| BSA (10x) | 1 |
| SpeI | 0,3 |
| XbaI | 0,3 |
| H2O | 5,9 |
| Total volume | 10 |
Gel:
0,45g agarose, 50 ml TAE (0,9%), 3 µl GELRED, 90 Volt, running time ~110 minutes
Comment: This time both samples look well. P822 will be sent for sequencing.
Sequencing analysis: pSB1C3_leftITR_pCMV_betaglobin_mGMK_TK30_hgH_rITR (P825)
Investigator: Stefan
For sequencing the following primers were used_
- qPCR primer CMV for (O20)
- GOI primer for (O30)
- seq. primer TK GMK for (O23)
- TK30 SDM Pst for (O191)
Sequencing comfirmed that the pysical sequence fits our theoretical. Next step will be to remove the remaining PstI restriction site within TK30.
Repetition of cloning of Rep into Rep78
Investigator: Kira
Comment: The last 2 approaches to digest Rep78 and to clone failed because Rep78 was almost gone after PCR purification and was not detectable on the gel, even if I used the double amount of DNA. Thus, new over night culture was grown in order to perform the experiment again
Regarding the activity conditions of the enzymes, digestion was performed in 2 steps.
| Components | Rep78 |
| DNA | 3 µl |
| BSA (100x) | 2 µl |
| Buffer no. 3 | 2,0 µl |
| Enzyme SwaI | 1,0 µl |
| H2O | 12 µl |
| Total volume | 20 |
incubation @ 25 C for approx. 2 h
Dephosphorylation was performed because of the blunt cleavage. PCR phosphorylation followed in order to get rid off the buffer because Buffer 3 is not appropriate for further digestion.
| Components | Rep78 |
| DNA | 30 µl |
| BSA (100x) | 0 µl |
| Buffer no. 2 | 4,5 µl |
| Enzyme HindIII | 1,0 µl |
| H2O | 9,5 µl |
| Total volume | 45 |
incubation for 2h at 37C
Ligation with T4 ligase
v(insert)=4,4 ul
v(vector)= 3,6 ul
T4 buffer= 1 ul
T4 ligase= 1 ul
incubation at RT for 45 min
Transformation according to the standard protocol with BL21
Midi-Prep
Investigator: Chris W.
Midi-Prep of:
pSB1C3_CMV_VP123_capins =P862 =B474
pSB1C3_P40_VP123 =P863 =B651
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | P862 | P863 |
| concentration (ng/µl) | 3310,5 | 2209,2 |
149. labday 14.10.2010
Mini-Prep and test digestion of several constructs
Investigator: Stefan
Glycerol stocks were prepared:
- B658 = pSB1C3_CD_SDM-PstI_III clone 1
- B659 = pSB1C3_CD_SDM-PstI_III clone 2
- B660 = pSB1C3_mGMK_sr39 clone 1
- B661 = pSB1C3_mGMK_sr39 clone 2
Mini-Prep was performed according to standard protocol:
- P858 = pSB1C3_CD_SDM-PstI_III clone 1 c = 241,7 ng/µl
- P859 = pSB1C3_CD_SDM-PstI_III clone 2 c = 212,1 ng/µl
- P860 = pSB1C3_mGMK_sr39 clone 1 c = 231,8 ng/µl
- P861 = pSB1C3_mGMK_sr39 clone 2 c = 265,2 ng/µl
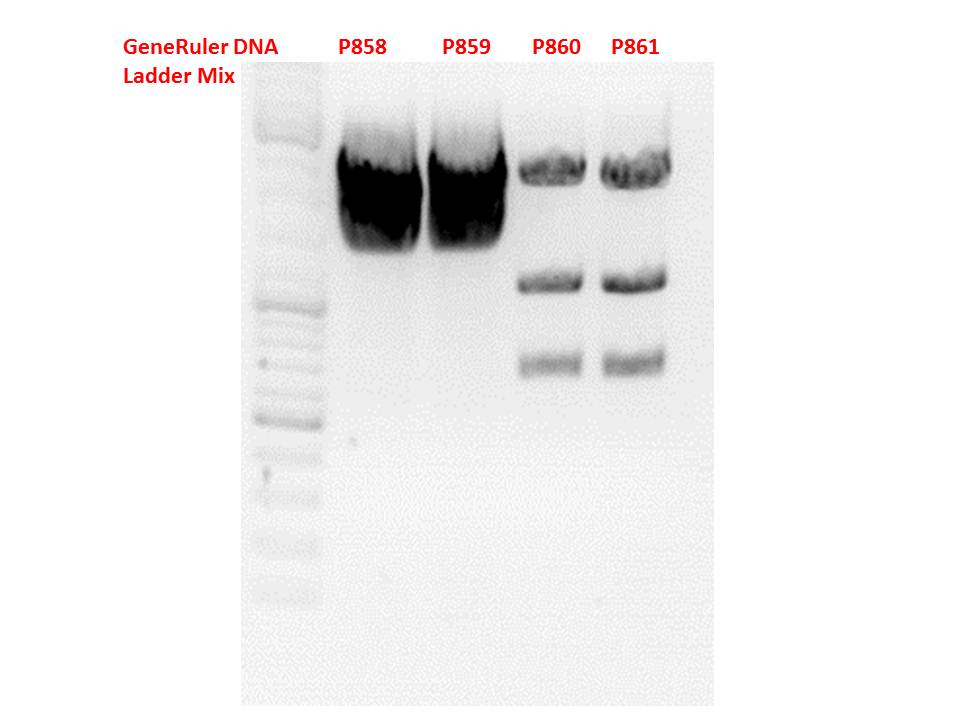
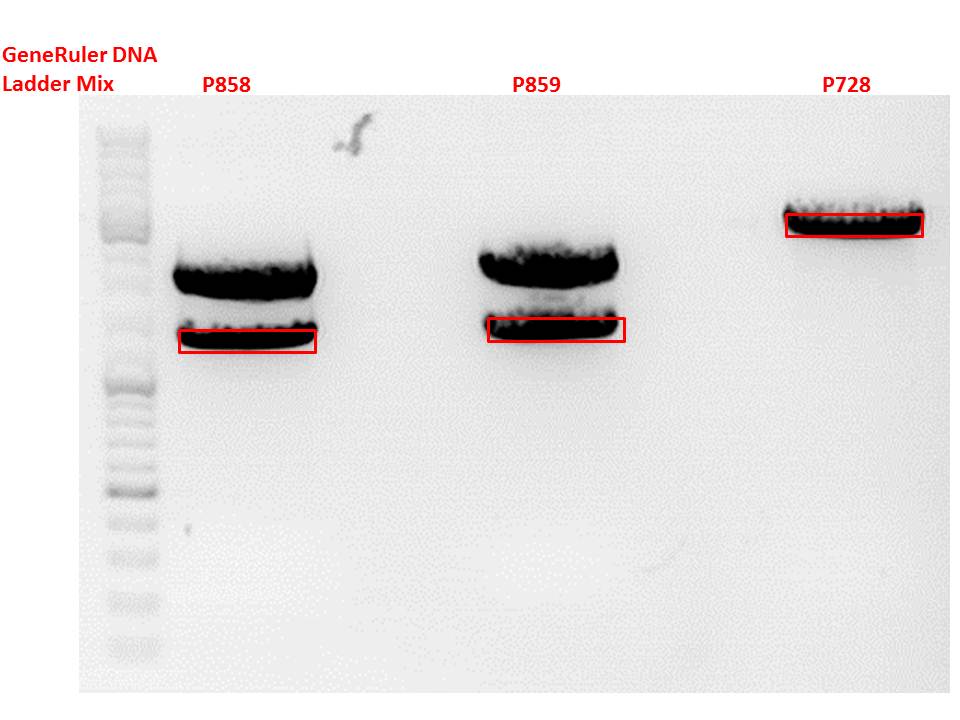
Test digestion:
| Components | P858 + P859 / µl | P860 + P861 / µl |
| DNA | 7 | 1,5 |
| Buffer 4 | 1 | 1 |
| BSA (10x) | 1 | 1 |
| Enzyme I | PstI 0,3 | BstXI 0,3 |
| Enzyme II | EcoRI 0,3 | XbaI 0,3 |
| Enzyme III | - | SpeI 0,3 |
| H2O | 0,4 | 5 |
| Total volume | 10 | 10 |
Gel:
0,5g agarose, 50 ml TAE (1%), 3 µl GELRED, 115 Volt, running time ~50 minutes
Comment: .
Continuation of Western Blot of unmodified AAV2 capsids
Investigator: Hanna
Comment: Yesterday, SDS PAGE, Western Blot and primary B1 mouse antibody incubation was performed.
Today, secondary antibody incubation and ECL analysis will be done.
- Membrane was washed 3 x 7 minutes in TBST (0.2% Tween).
- A 1:5000 dilution of secondary goat anti-mouse HRP (sc2005) coupled antibody in TBS was prepared.
- Membrane was incubated in secondary antiobody for 1 hour.
- Membrane was washed 3 x 10 minutes.
- Pierce ECL Western Blotting Substrate was prepared: 350 µL Luminol Enhancer Solution + 350 µL Peroxidase Solution
- Analysis of the membrane revealed that the samples didn't run well. Much unspecific antibody binding was detectable.
Because of these dissatisfying results SDS PAGE will be repeated. Instead of performing Western Blot, gel wioll be stained by Coomassie in order to optimize SDS-PAGE conditions.
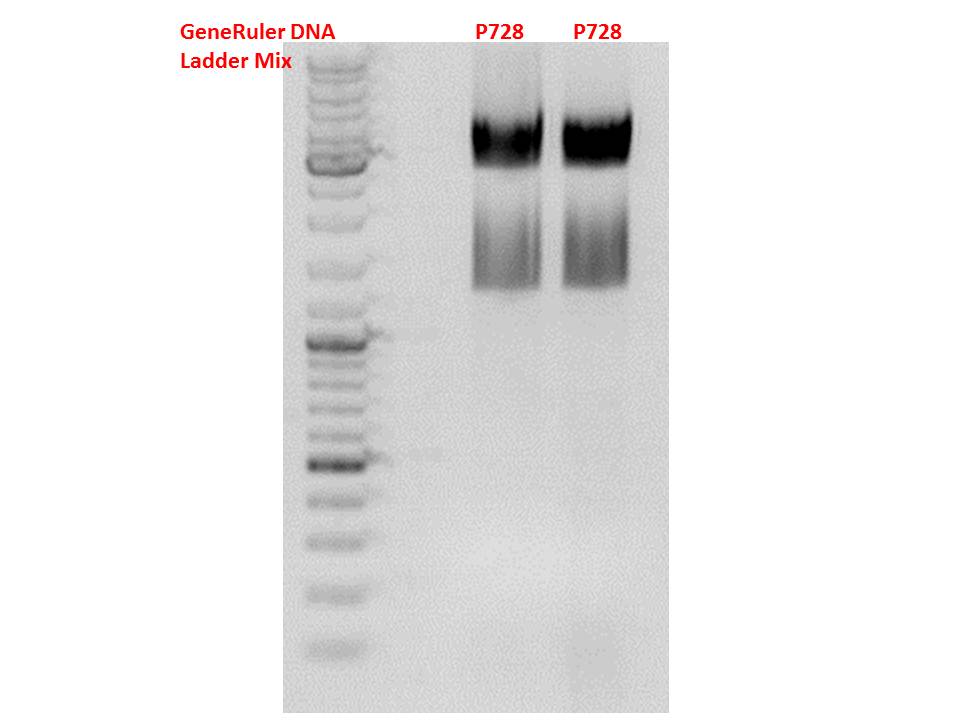
Cloning hGH_rITR into (form P728) into pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_SDM-PstI (P822)
Investigator Patrick
Digestions, 1 h 40 minutes, 37 °C:
- P728: 1 µl XbaI, 1 µl PstI, 2 µl BSA, 2 µl Buffer 4, 6 µl DNA, 8 µl H2O
- P822: 1 µl SpeI, 1 µl PstI, 2 µl BSA, 2 µl Buffer 4, 6 µl DNA, 8 µl H2O
Expected results for the 1% agarose gel:
- P728: about 2100 and 650 bp
- P822: about about 4900 and 10 bp (wont be visible on the gel)

As usual the PSB1C3 bands indicate a higher bp amount than they actually got.
The gelextraction ...
- P728: 9 ng/µl
- P822: 67,5 ng/µl
... and following ligation (2 µl Insert, 6 µl vector, 1 µl T4 DNA Ligase, 1 µl T4 DNA Ligase Buffer, 40 minutes, RT) were performed according to the standard protocol. After the transformation (with BL21) the cells were plated and put into the 37°C room.
Cloning of the ViralBricks 453-empty and 587 empty
These ViralBricks were not ordered as hybridized oligos, but can be cloned from the Cap-Gene-Synthesis into the ViralBrick MCS.
| components | volume of p320 (ViralBrick-BLA) /µl | volume of p320 (ViralBrick-BLA) /µl | volume of p668 (pSB1C3_RepCap_IRCK) /µl | volume of p668 (pSB1C3_RepCap_IRCK) /µl |
| DNA | 2µl | 2µl | 7µl | 7µl |
| BSA (100x) | 2.5µl | 2.5µl | 2.5µl | 2.5µl |
| Buffer 4 (10x) | 2.5µl | 2.5µl | 2.5µl | 2.5µl |
| SspI | 1µl | 0 | 1µl | 0 |
| SalI | 1µl | 0 | 1µl | 0 |
| BamHI | 0 | 1µl | 0 | 1µl |
| PvuII | 0 | 1µl | 0 | 1µl |
| H2O | 16µl | 16µl | 11µl | 11µl |
| Total volume | 25µl | 25µl | 25µl | 25µl |
| Sample Marker | Sample ViralBrick_BLA 453 cut | Sample ViralBrick_BLA 587 cut | Sample pSB1C3_001_RepCap_IRCK 453 cut | Sample pSB1C3_001_RepCap_IRCK 587 cut | |
|---|---|---|---|---|---|
| Lane | 7µl | 30µl | 30µl | 30µl | 30µl |
SDS PAGE of viral capsids
Investigator: Hanna
Comment: In order to optimize SDS PAGE with the viral capsid proteins, I didn't perform Western Blot, but Coomassie staining.
1.: Preparation of 10% polyacrylamide gel:
Following standard protocol.
2.: Preparation of virus samples:
Two different virus stocks with unmodified capsid were taken. 2 samples were prepared of each: One sample was untreated, the other one was centrifuged in order to sediment cell fragments.
10 µL Laemmli buffer (5x) was added to 40 µL virus sample and incubated at 95°C for 5 minutes.
3.: SDS PAGE
Loading plan:
| 10 µL PageRuler Prestained Protein Ladder | 10 µL Sample 1 (not centrifuged) | 20 µL Sample 1 (not centrifuged) | 10 µL Sample 1 (centrifuged) | 20 µL Sample 1 (centrifuged) | 10 µL Sample 2 (not centrifuged) | 20 µL Sample 2 (not centrifuged) | 10 µL Sample 2 (centrifuged) | 20 µL Sample 2 (centrifuged) |
Gel was run 1.5 hours at 10 mA.
4.: Coomassie staining
Gel was put into coomassie bath, heated in microwave and incubated for 1 hour, shaking.
In order to remove background coomassie staining, gel was incubated in acetic acid (20%) for 2 hours, shaking.
Results revealed that samples ran well (vertically :) ) - 20 µL seems to be too much. Interestingly centrifuging the virus stocks in order to remove cell debris didn't lead to better results.
To do: Perform Western Blot again - with 10 µL Virus stock, 0.5% instead of just 0.2% TBS-milk and longer washing steps.
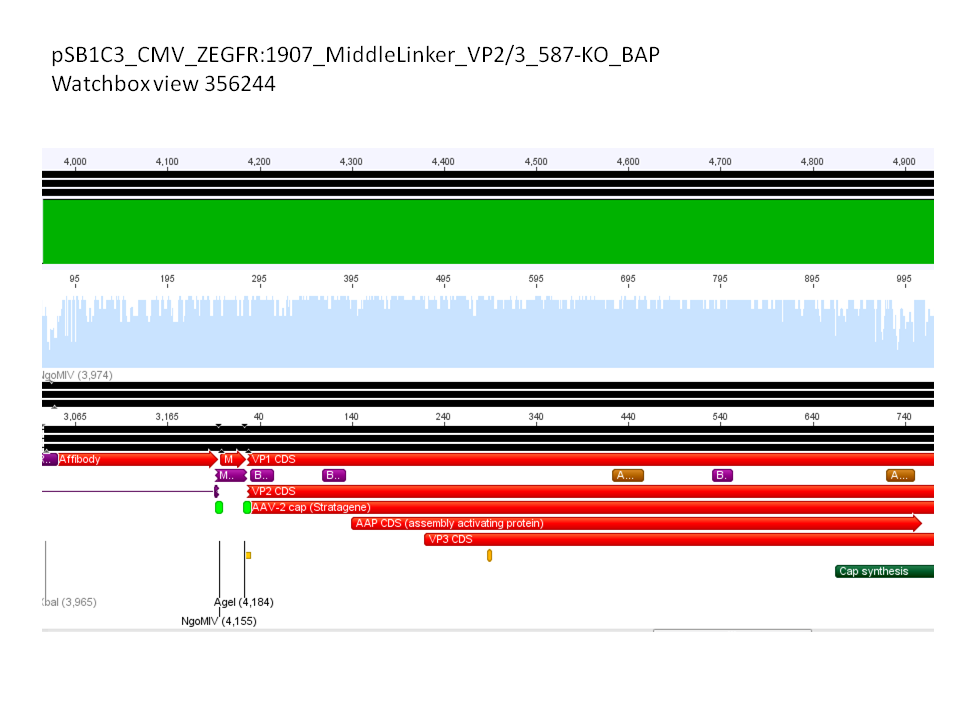
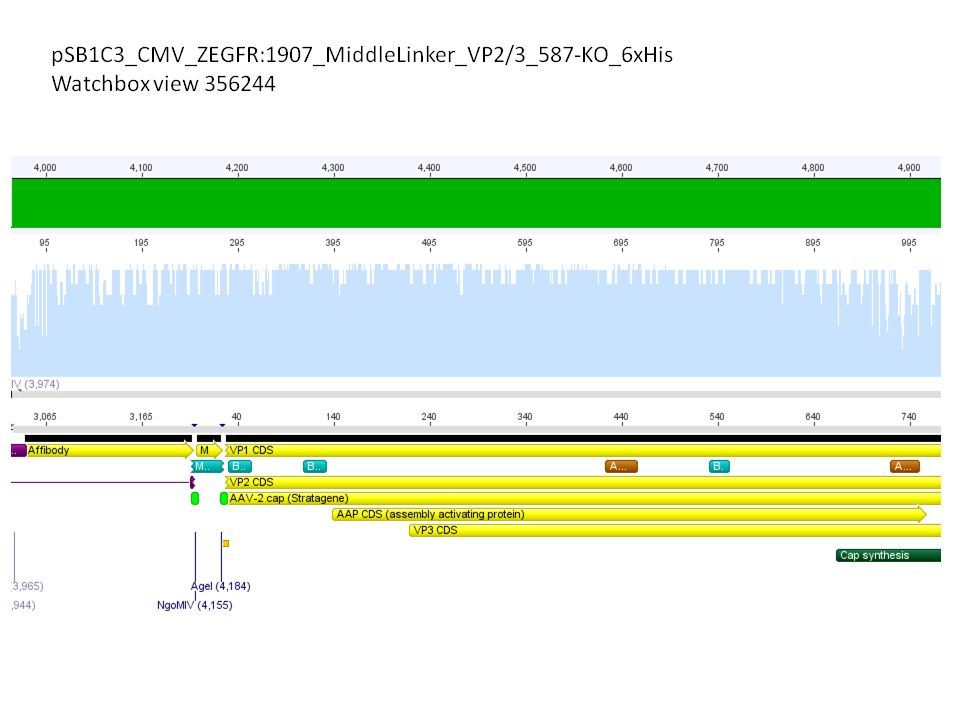
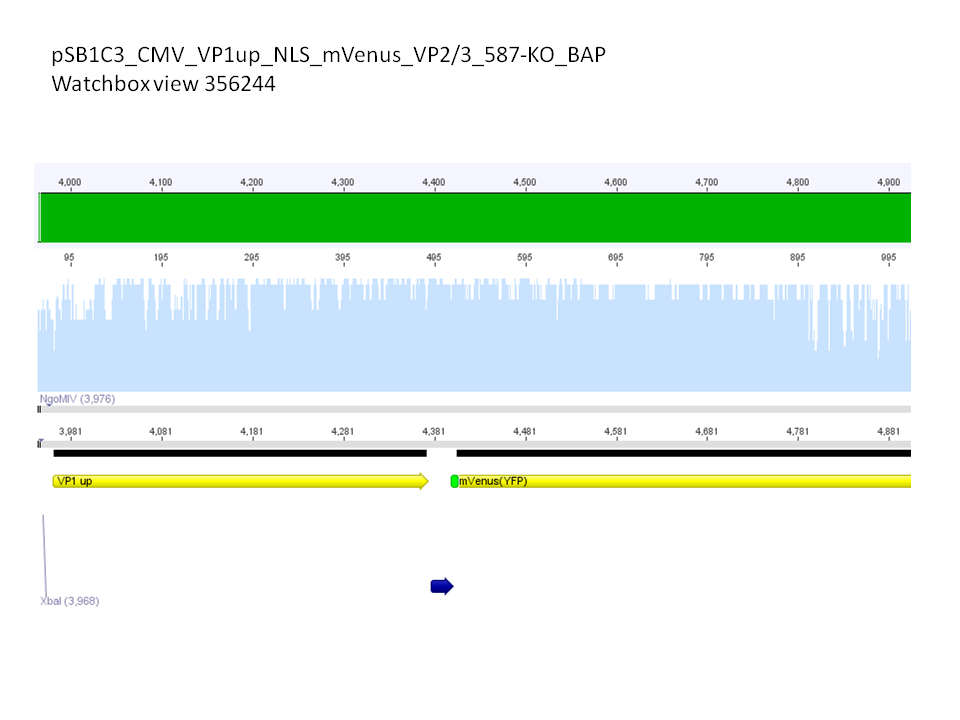
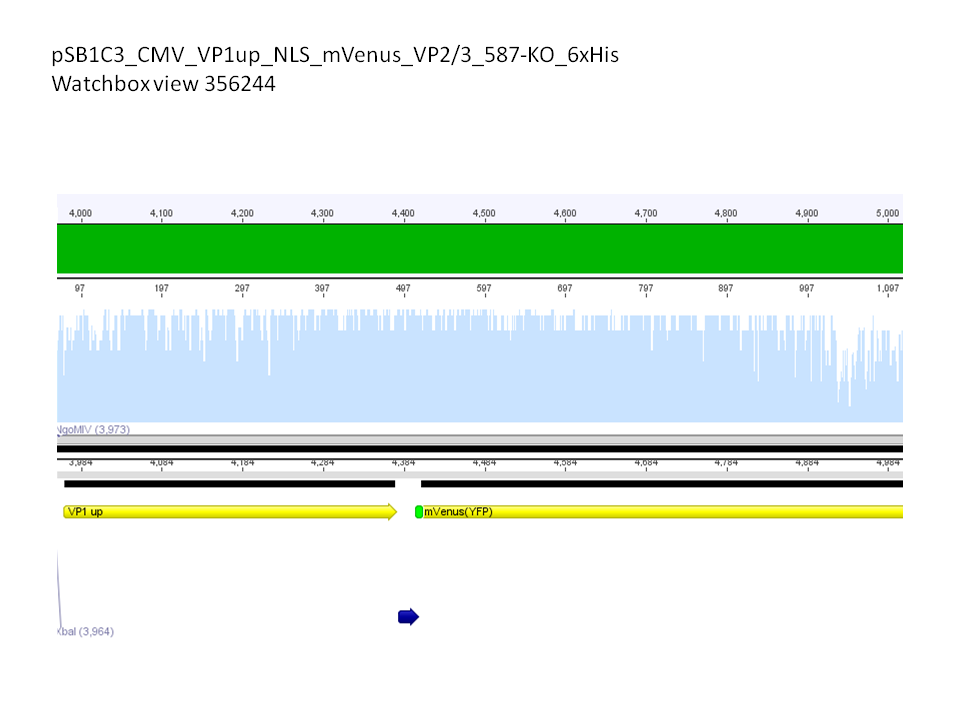
Sequence analysis of super constructs
Investigator: Hanna
Comment: We assembled targeting super constructs which specifically bind the EGF receptor with VP2 N-terminal fused ZEGFR:1907 Affibody and can either be purified via a loop inserted His-Tag or visualized via Streptavidin coupled fluorescent proteins.
Further on these loop motifs were vloned to the "mVenus VP1 insertion".
Sequencing results showed that insertion of these constructs into pSB1C3_CMV worked!
Next steps: Put sample sinto BioBrick Box.
Midi-Prep + Transfection.
Transfection with pCMV, p40, pAAV_RC
Investigator: Kira
Comment: Transfection was performed in order to evaluate the promoter activities.
Yesterday, the cells were seeded in the 6 well plate by Sven. Transformation was performed according to the manufacturer protocol (Qiagen).
Trafo evaluation of Rep78
Investigator: Kira
Transformation was successful. 2 clones will be picked for mini prep tomorrow.
Picking clones
Investigator: Kira
Rep78 and Rep52 were picked (2 clones from each plate), in order to continue with mini prep and PCR amplification.
Cloning of CD into pSB1C3_hGH_rITR
Investigator: Stefan
.
Vector name:
pSB1C3_hGH_rITR (P728)
Insert name:
pSB1C3_CD_SDM-PstI_III clone 1 (P858)
pSB1C3_CD_SDM-PstI_III clone 2 (P859)
Digestion:
| components | volume P858 + P859 /µl | volume P728 /µl |
| DNA | 6 | 8 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme EcoI | 1 | 1 |
| Enzyme XbaI | - | 1 |
| Enzyme SpeI | 1 | - |
| H2O | 8 | 6 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 |
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
Gel extraction:
Was performed according to protocol.
T4 Ligation:
| ligation name | 728 + 858 | 728 + 859 |
| volume of vector | 3,14 | 3,14 |
| volume of insert | 4,86 | 4,86 |
| T4 ligase buffer (10x) | 1 | 1 |
| T4 ligase | 1 | 1 |
Transformation:
Was performed according to standard protocol using BL21 cells.
150. labday 15.10.2010
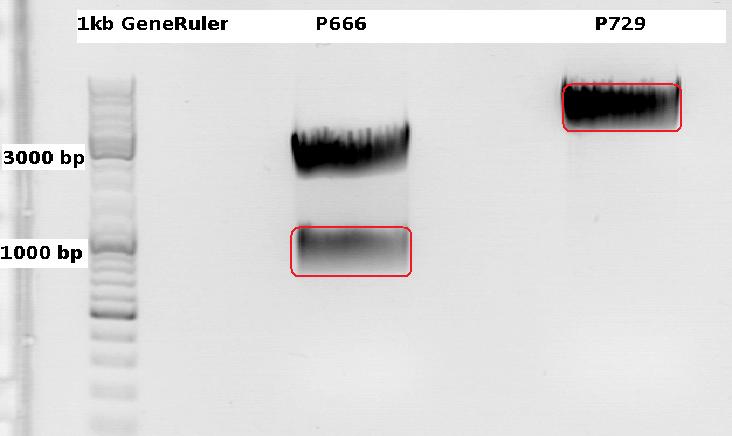
Cloning CFP (from P666: PSB1C3_CFP) into pSB1C3_leftITR_CMV_beta-globin (P729)
Investigator Patrick
Digestions, 2 h 10 minutes, 37 °C:
- P666: 5 µl DNA, 2 µl BSA, 2 µl Buffer 4 (10x), 1 µl Xba, 1 µl PstI, 9 µl H2O
- P729: 4 µl DNA, 2 µl BSA, 2 µl Buffer 4 (10x), 1 µl SpeI, 1 µl PstI, 10 µl H2O
Expected results for the 1% agarose gel:
- P666: about 2100 and 750 bp
- P729: about 3300 and 20 bp
P728: 11,8 ng/µl
P822: 34,6 ng/µl
... and following ligation (2,5 µl Insert, 5,5 µl vector, 1 µl T4 DNA Ligase, 1 µl T4 DNA Ligase Buffer (10x), 40 minutes, RT) were performed according to the standard protocol. After the transformation (with XL1B) the cells were plated and put into the 37°C room.
Two additional transformations were performed with ligations from Volker labeled: "ligation viral brick 453 empty" & "viral brick 587 empty".
The following day the plates were checked for clones. Unfortunately there grew no clones on these two plates contrary to my plate with a a lot of clones.
Midi-Prep
Investigator: Chris W.
Midi-Prep of:
pSB1C3_001_RC_IRCK_P5tataless clone 1 =P866 =B516
pSB1C3_001_CMV_VP123_587-KO_Z34C_spacer clone2 =P867 =B526
pSB1C3_001_CMV_VP123_587-KO_Z34C clone2 =P868 =B529
pSB1C3_CMV_Zegfr:1907_MiddleLinker_VP2/3_587-KO_BAP clone 1 =P869 =B680
pSB1C3_CMV_Zegfr:1907_MiddleLinker_VP2/3_587-KO_6xHis clone 1 =P870 =B200
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | P866 | P867 | P868 | P869 | P870 |
| concentration (ng/µl) | 899,80 | 954,46 | 406,97 | 1642,76 | 1585,12 |
mini prep of several constructs
Investigator: Kira
c(rep52_1)=299,04 ng/ul
c(rep52_2)=290,07 ng/ul
c(rep78_1)=142,32 ng/ul
c(rep78_2)=175,36 ng/ul
Cell culture
Investigator: Kira
The cells are still alive. Medium was exchanged.--> RNA will be harvested tomorrow
Mini-Prep and test digestion of pSB1C3_CD_SDM-PstI_hGH_rITR
Investigator: Stefan
Glycerol stocks were prepared:
- B694 = pSB1C3_CD_SDM-PstI_hGH_rITR clone 1
- B695 = pSB1C3_CD_SDM-PstI_hGH_rITR clone 2
Mini-Prep was performed according to standard protocol:
- P875 = pSB1C3_CD_SDM-PstI_hGH_rITR clone 1 c = 73,1 ng/µl
- P876 = pSB1C3_CD_SDM-PstI_hGH_rITR clone 2 c = 78,3 ng/µl
Test digestion:
| Components | P875 + P876 / µl |
| DNA | 4 |
| Buffer 4 | 1 |
| BSA (10x) | 1 |
| XbaI | 0,3 |
| AgeI | 0,3 |
| H2O | 3,4 |
| Total volume | 10 |
Gel:
0,5g agarose, 50 ml TAE (1%), 3 µl GELRED, 115 Volt, running time ~50 minutes
Comment: Test digestion looks allright, cloning will be continued using P876.
151. labday 16.10.2010
Biobrick assembly: pSB1C3_lITR_CMV_ß-globin_CD_hGH_rITR and pSB1C3_lITR_phTERT_ß-globin_CD_hGH_rITR
Investigator: Achim
Plasmids:
- P729: pSB1C3_lITR_CMV_ß-Globin
- c= 243.4 ng/µl
- P730: pSB1C3_lITR_pHTERT_ß-Globin
- c= 81.1 ng/µl
- P876: pSB1C3_CD_SDM-PstI_hGH_rITR
- c= 78.3 ng/µl
Digestion:
| components | I1 (P729) | I2 (P730) | V (P876) |
| DNA | 6 | 14 | 14 |
| BSA (10x) | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 |
| Enzyme EcoI | 1 | 1 | 1 |
| Enzyme XbaI | - | - | 1 |
| Enzyme SpeI | 1 | 1 | - |
| H2O | 8 | - | - |
| Total | 20 | 20 | 20 |
Digestion: 2h, 37°C
Prep. gel:
- 0,8%, run for 45 min
- Corresponding bands were cut out
Gel ex.
- Nanodrop concentrations:
- I1: 37.54 ng/µl
- I2: 25.38 ng/µl
- V: 26.47 ng/µl
Ligation:
| ligation name | I1 + V | I2 + V |
| volume of vector | 4.69 | 4.23 |
| volume of insert | 3.31 | 3.77 |
| T4 ligase buffer (10x) | 1 | 1 |
| T4 ligase | 1 | 1 |
- Ligation @ RT for 40 min
Trafo:
- Done by Kira
Mini-prep of mutual pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI
Investigator Patrick
Yielded concentrations & given numbers:
- pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI clone 1: 208,4 ng/µl , P877 / B696
- pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI clone 2: 251,6 ng/µl , P878 / B696
- pSB1C3_lITR_phTERT_betaglobin_mGMK_TK30_hGH_rITR_SDM-PstI clone 3: 212,6 ng/µl , P889 / B696
Test-digestion: 0,5 µl SpeI, 0,5 µl PstI, 3 µl DNA, 1 µl Buffer 4, 1 µl BSA, 4 µl H2O, 40 minutes, 37°C
Expected results: fragments with about 650 and 4900 bp
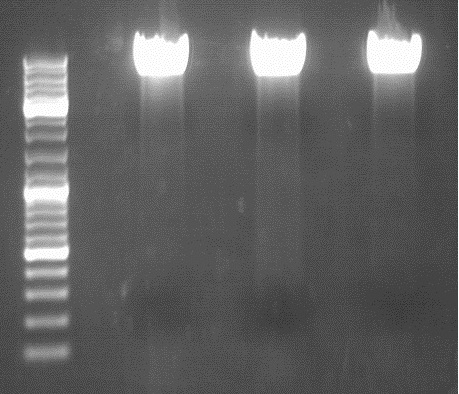
Obviously, this test digestion has to be repeated.
Preparations for tomorrow:
- Mini-prep of 3 mutual pSB1C3_leftITR_CMV_beta-globin_CFP clones (have to be picked from the plate)
- Midi-prep of pHelper
- Midi-prep of B689:pSB1C3_lITR_CMV_betaglobin_mGMK_TK30_SDM-PstI_hGH_rITR clone 2
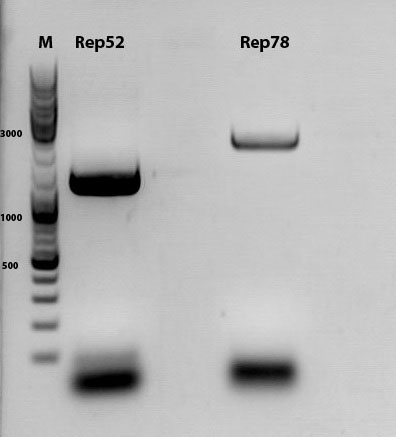
Biobrick assembly of Rep78 and Rep52
Investigator: Kira
Comment: After replacing the mutated Rep parts by the ordered Rep parts, PCR amplification has to be done in order to produce a biobrick.
PCR program:
c(Rep52)=299 ng/ul
c(Rep78)=175 ng/ul
Rep52: praefix 094 & suffix 097
Rep78: praefix 093 & suffix 097
| components | volume in µl |
| 5x Phusion HF buffer | 10 |
| 10 mM dNTP mix | 1 |
| primer_for (1:10 dilution) | 2,5 |
| primer_rev (1:10 dilution) | 2,5 |
| DNA template (1:100) | 0,5 |
| DMSO | 0,5 |
| Phusion polymerase | 0,5 |
| H2O | 32,5 |
| Total volume (e.g. 50 µl) | 50 |
| Cycles | Temperature | Time |
| 98°C | 30 sec | |
| 10x | 98°C | 15 sec |
| 63°C | 25 sec | |
| 72°C | 32 sec | |
| 20x | 98°C | 15 sec |
| 66°C | 25 sec | |
| 72°C | 32 sec | |
| 1x | 72°C | 5 min |
| Hold 4°C |
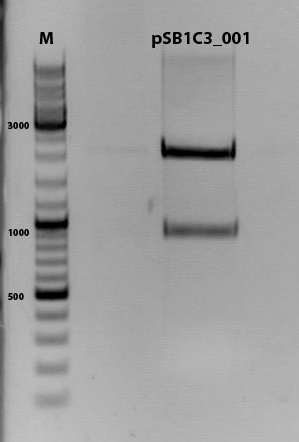
Digestion of plasmid backbone:
pSB1C3_001 is used as backbone
| Components | <b>vector Volume/µL |
| DNA | 3,5 µl |
| BSA (10x) | 2 µl |
| Buffer no. 4 (10x) | 2,0 µl |
| Enzyme 1 EcoRI-HF | 0,5 µl |
| Enzyme 2 SpeI | 1,0 µl |
| H2O | 15 µl |
| Total volume | 25 |
incubation @ 37 C for approx. 2 h
Digestion of PCR product:
| Components | PCR product Volume/µL |
| DNA | 35,0 µl |
| BSA (100x) | 0,45 µl |
| Buffer no. 4 | 4,5 µl |
| Enzyme 1 EcoRI-HF | 1,5 µl |
| Enzyme 2 SpeI | 2,0 µl |
| H2O | 1,5 µl |
| Total volume | 45 |
incubation @ 37 C for approx. 2 h
T4 ligation for 40 min
Transformation according to the standard protocol
RNA harvesting
Investigator: Kira
After transfection, the cells were incubated for 48 hours. Today, the cells will be harvested and RNA extracted, in order to perform RT-PCR and an additional PCR for evaluation of promoter activity.
The transfected cells were trypsinised and centrifuged for 2 min. The supernatant was discarded and pellet washed 2x with PBS. RNeasy Kit [Qiagen] was used for RNA extraction according to the manufacturer protocol.
c(CMV)= 335,69 ng/ul
c(P40)= 857,92 ng/ul
c(AAV_RC)= 760,21 ng/ul
152. labday 17.10.2010
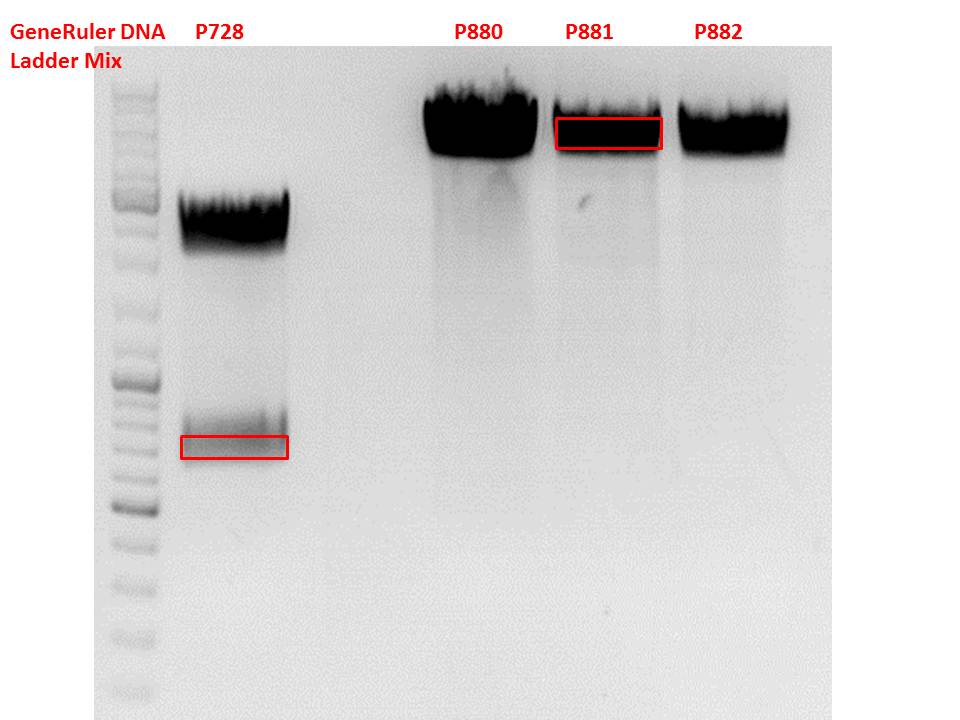
Test digestion of pSB1C3_lITR_CMV_ß-globin_CFP
Investigator: Anna
Vector name:
pSB1C3_lITR_CMV_betaglobin_CFP_cl1 (P880): c = 452,67 ng/µl
pSB1C3_lITR_CMV_betaglobin_CFP_cl2 (P881): c = 288,88 ng/µl
pSB1C3_lITR_CMV_betaglobin_CFP_cl3 (P882): c = 288,36 ng/µl
Test Digestion:
| components | volume P880 - P882 /µl | volume P434 /µl |
| DNA | 2 | 2 |
| BSA (10x) | 1 | 1 |
| Buffer 4 (10x) | 1 | 1 |
| Enzyme NgoMIV | 0,3 | 0,3 |
| Enzyme AgeI | 0,3 | 0,3 |
| H2O | 5,4 | 5,4 |
| Total volume | 10 | 10 |
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
File:Freiburg Test digestion of pSB1C3 lITR CMV ß-globin CFP.jpg
Cloning of hGH_rITR into pSB1C3_lITR_CMV_betaglobin_CFP
Investigator: Stefan
Cloning of our last GOI!
Vector name:
pSB1C3_lITR_CMV_betaglobin_CFP cl 1-3 (P880-P882)
Insert name:
pSB1C3_hGH_rITR (P728)
Digestion:
| components | volume P880 - P882 /µl | volume P728 /µl |
| DNA | 3 | 8 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme PstI | 1 | 1 |
| Enzyme XbaI | - | 1 |
| Enzyme SpeI | 1 | - |
| H2O | 11 | 4 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 |
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
Test digestion of all constructs looked alright, therefore, cloning was continued using P881 only.
Gel extraction:
Was performed according to protocol.
T4 Ligation:
| ligation name | 728 + 881 |
| volume of vector | 2,68 |
| volume of insert | 5,32 |
| T4 ligase buffer (10x) | 1 |
| T4 ligase | 1 |
Transformation:
Transformation was performed according to standard protocol using BL21 cells.
RT-PCR
Investigator: Kira
For further experiments, RNA has to be translated into cDNA. The PCR was performed according to the manufacturer protocol.
153. labday 18.10.2010
Mini-Prep and test digestion of several constructs
Investigator: Jessica
Glycerol stocks were prepared:
- B702 = pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR clone 1
- B703 = pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR clone 2
- B704 = pSB1C3_001_VP3 clone 1
- B705 = pSB1C3_001_VP3 clone 2
- B706 = pSB1C3_001_VP2 clone 1
- B707 = pSB1C3_001_VP2 clone 2
- B708 = pSB1C3_001_Rep78 clone 1
- B709 = pSB1C3_001_Rep78 clone 2
- B710 = pSB1C3_001_Rep52 clone 1
- B711 = pSB1C3_001_Rep52 clone 2
- B712 = pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR clone 1
- B713 = pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR clone 2
- B714 = pSB1C3_001_VP1 clone 1
- B715 = pSB1C3_001_VP1 clone 2
Mini-Prep was performed according to standard protocol:
- P886 = pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR clone 1 c= 232,2ng/µl
- P887 = pSB1C3_lITR_pTERT_ßglobin_CD_hGH_rITR clone 2 c= 186,2ng/µl
- P888 = pSB1C3_001_VP3 clone 1 c= 300,8ng/µl
- P889 = pSB1C3_001_VP3 clone 2 c= 284,4ng/µl
- P890 = pSB1C3_001_VP2 clone 1 c= 298,3ng/µl
- P891 = pSB1C3_001_VP2 clone 2 c= 299,9ng/µl
- P892 = pSB1C3_001_Rep78 clone 1 c= 143,9ng/µl
- P893 = pSB1C3_001_Rep78 clone 2 c= 163,4ng/µl
- P894 = pSB1C3_001_Rep52 clone 1 c= 166,6ng/µl
- P895 = pSB1C3_001_Rep52 clone 2 c= 181,6ng/µl
- P896 = pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR clone 1 c= 250,5ng/µl
- P897 = pSB1C3_lITR_CMV_ßglobin_CD_hGH_rITR clone 2 c= 173,4ng/µl
- P898 = pSB1C3_001_VP1 clone 1 c= 272,7ng/µl
- P899 = pSB1C3_001_VP1 clone 2 c= 294,8ng/µl
- P900 = pSB1C3_hGH_rITR (from B160) c= 136,7ng/µl
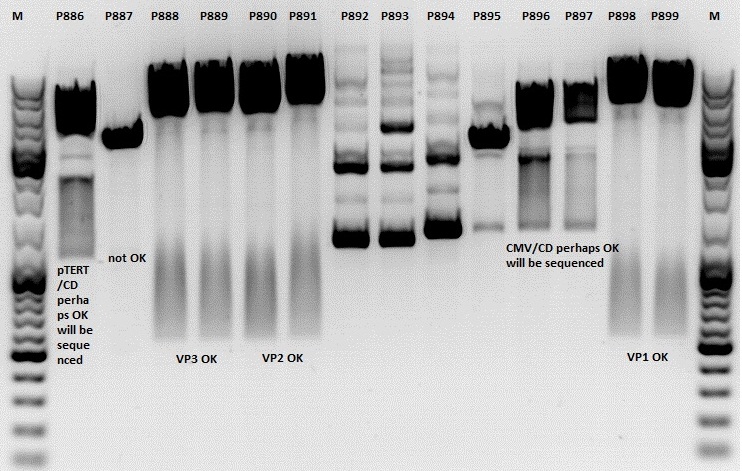
Test digestion:
| Components | P886,887,892,893,894,895,896,897 / µl | P888,889,890,891898,899 / µl |
| DNA | 1,5 | 1,5 |
| Buffer | (4) 1 | (2) 1 |
| BSA (10x) | 1 | 1 |
| NgoMIV | 0,4 | - |
| XbaI | 0,4 | - |
| PstI | - | 0,6 |
| XcmI | - | 0,4 |
| H2O | 4,5 | 4,5 |
| Total volume | 10 | 10 |
Gel:
1,0g agarose, 100 ml TAE (1%), 6 µl GELRED, Volt, running time minutes
Comment:
154. labday 19.10.2010
155. labday 20.10.2010
 "
"