Team:Lethbridge/Modeling
From 2010.igem.org
David.franz (Talk | contribs) (Added a short homology blurb) |
David.franz (Talk | contribs) (Added placeholder images) |
||
| Line 103: | Line 103: | ||
==<font color="white">Why Metabolic Modeling== | ==<font color="white">Why Metabolic Modeling== | ||
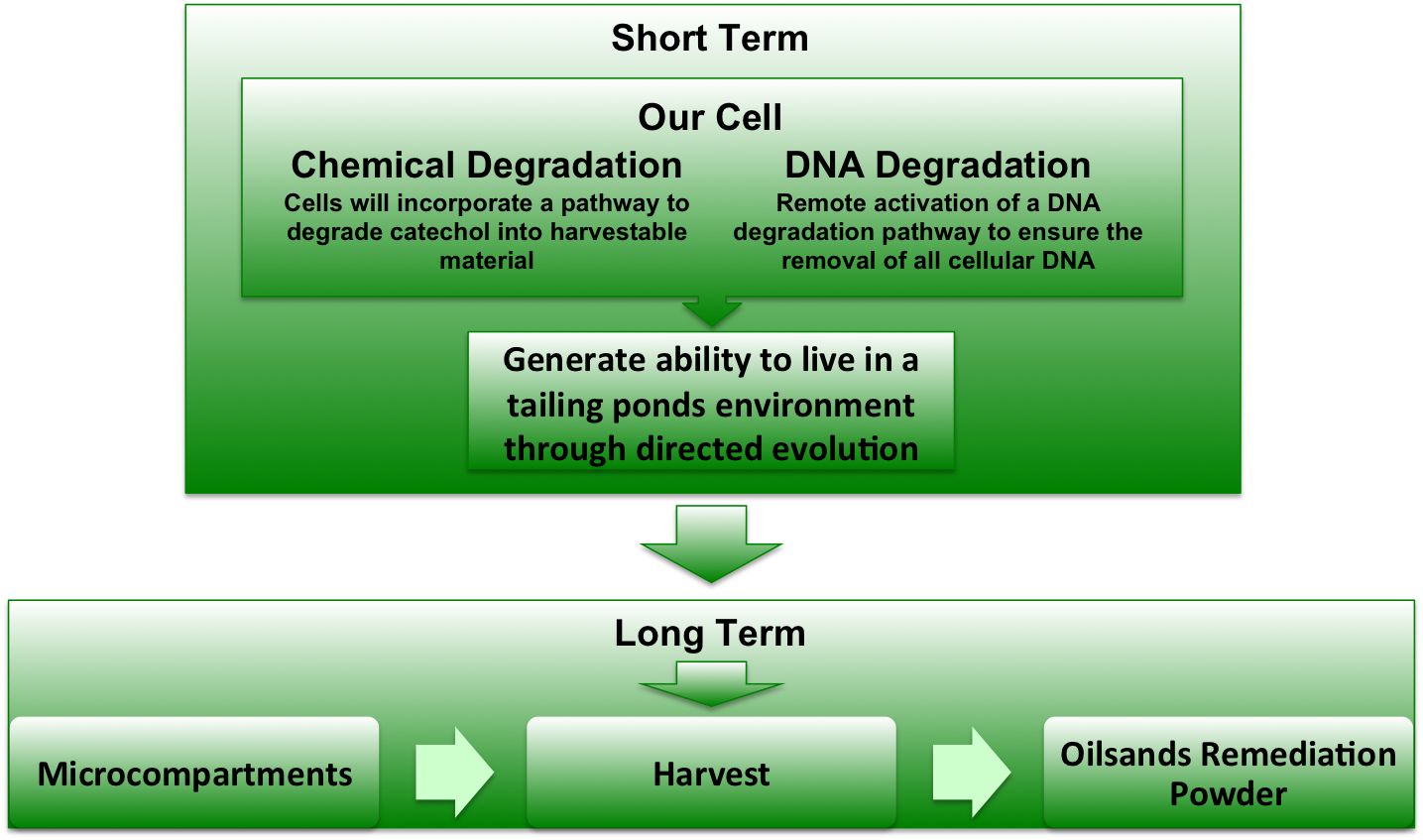
The majority of degredation pathways for oilsands contaminants convert various organic compounds into catechol. Our project this year in the incorporation of the gene <i>xylE</i> into <i>E. coli</i>. The <i>xylE</i> gene encodes catechol-2,3,-dioxygenase, which converts catechol into 2-hydroxymuconate semialdehyde. This semialdehyde can be metabilized by <i>E. coli</i> by means of the glycolysis / Krebbs cycle. | The majority of degredation pathways for oilsands contaminants convert various organic compounds into catechol. Our project this year in the incorporation of the gene <i>xylE</i> into <i>E. coli</i>. The <i>xylE</i> gene encodes catechol-2,3,-dioxygenase, which converts catechol into 2-hydroxymuconate semialdehyde. This semialdehyde can be metabilized by <i>E. coli</i> by means of the glycolysis / Krebbs cycle. | ||
| + | |||
| + | [[Image:UofLProjectOverview.jpg|x150px|right|text-top]] | ||
| + | |||
However, little is known about how the effect of increased flux of these metabolites into cell (such as intermediate concentrations and metabolite fluxes). Thus to asses the viability of specialized bioremediation and metabolic engineering it is very important to understand what is happening within the cell. By using computer modeling of metabolism pathways and using kintetic data, we can asses the robustness of the system and what we can do to improve the efficiency of the system. | However, little is known about how the effect of increased flux of these metabolites into cell (such as intermediate concentrations and metabolite fluxes). Thus to asses the viability of specialized bioremediation and metabolic engineering it is very important to understand what is happening within the cell. By using computer modeling of metabolism pathways and using kintetic data, we can asses the robustness of the system and what we can do to improve the efficiency of the system. | ||
| Line 127: | Line 130: | ||
==<font color="white">Results== | ==<font color="white">Results== | ||
| + | [[Image:UofLProjectOverview.jpg|x150px|right|text-top]] | ||
<br> | <br> | ||
<br> | <br> | ||
 "
"