Team:Washington/Gram Positive/Test
From 2010.igem.org
| Line 58: | Line 58: | ||
| - | =''Mutant | + | ='Designing CapD_CP to be a better Hydrolase'= |
| + | ==Screening Mutant Libraries== | ||
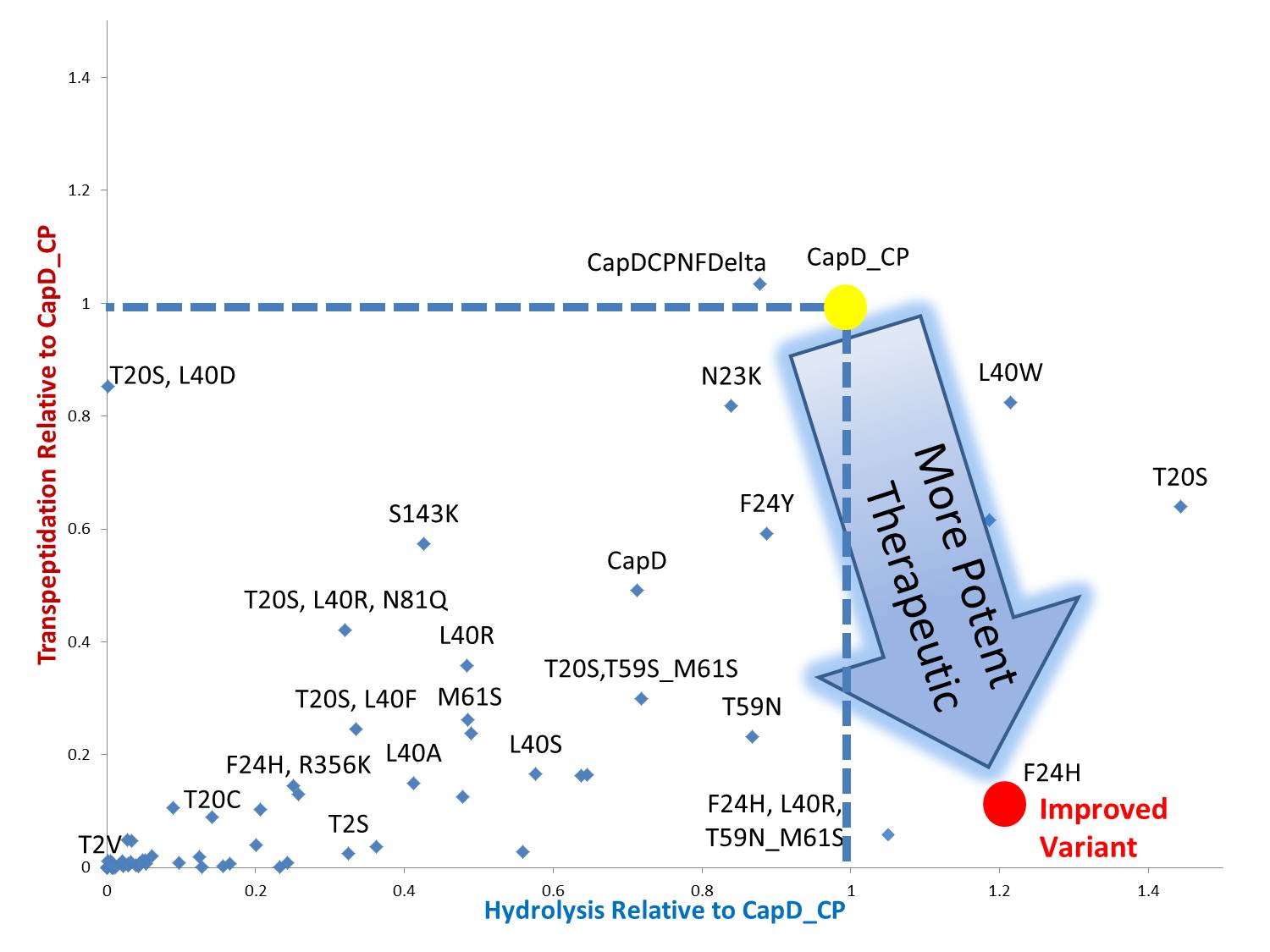
By standardizing the activity slope of each design relative to CapD_CP, a scatterplot easily portrays the qualities of each mutant. Several designs show negative catalytic curves similar to the catalytic knockouts. Some immediately show a negative activity curve meaning decrease in transpeptidation, hydrolysis, or both. T20S is a promising mutant hydrolase design. | By standardizing the activity slope of each design relative to CapD_CP, a scatterplot easily portrays the qualities of each mutant. Several designs show negative catalytic curves similar to the catalytic knockouts. Some immediately show a negative activity curve meaning decrease in transpeptidation, hydrolysis, or both. T20S is a promising mutant hydrolase design. | ||
[[Image:Washington_Mutant_design.jpg|thumb|400px|right|]] | [[Image:Washington_Mutant_design.jpg|thumb|400px|right|]] | ||
| + | <br style="clear: both" /> | ||
| + | |||
| + | ==Characterization of best hits== | ||
| + | blah blah blah | ||
| + | [[Image:Temp.jpg|thumb|400px|right|]] | ||
| + | <br style="clear: both" /> | ||
| + | |||
<!---------------------------------------PAGE CONTENT GOES ABOVE THIS----------------------------------------> | <!---------------------------------------PAGE CONTENT GOES ABOVE THIS----------------------------------------> | ||
Revision as of 19:10, 6 October 2010
More descriptive titles, verification that your CapD_CP was a monomer ( gel) should go before testing catalytic ability
Contents |
Construction of CapD_CP
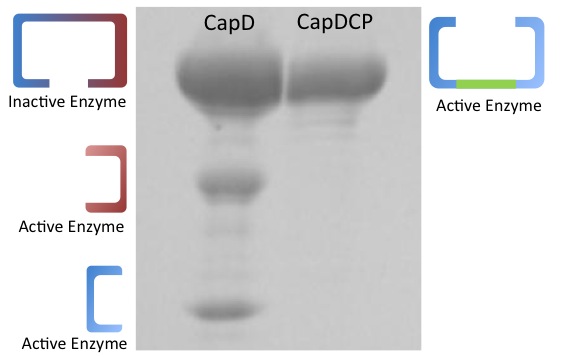
CapD_CP runs on a gel as one clean band
Depicted in Figure X is a gel comparing purified capD and capD_cp. This demonstrates one of the major advantages of CapD_CP, in addition to the fact that it is easy to express, it comes out in 1 clean band and is easy to quantify. CapD shows three bands, while the lower two bands are the expected pieces that occur after the enzyme undergoes self cleavage, the upper band is of unclear origin. It's mass corresponds to what would either be the unprocessed, inactive, monomeric form of the enzymes or the dimeric form that didn't denature. This ambiguity makes the amount of active enzyme from capD difficult to quantify.
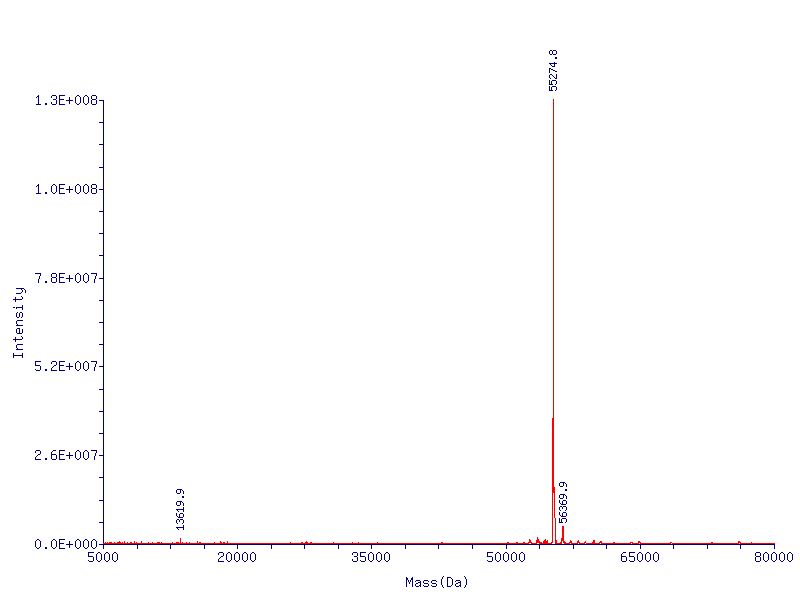
CapD_CP is produced without the N-term methionine as expected
In Figure X we show the mass spec conformation of the molecular wieght of capD_cp. As expected the N-terminal methionine is cleaved off by the native E. coli methionine aminopeptidase, resulting in a high level of pure active enzyme.
The circular permutation didn't result in a loss of activity
In Table X we show the kinetic paramters for capD and capD_cp as determined using our enzyme assay (link). While the kcat is difficult to quantify for capD the data suggests that the kinetic parameters are within error of eachother. Therefore the circullar permutation of capD did not have a negative effect on catalytic actiivty, suggesting that the autoprocessing feature is not required for catalysis but simply a regulatory function for the protein.
Catalytic Residue knock-outs show that capD_cp is catalyzing the reaction as expected
Before we can predict which mutations increase hydrolysis capability, we need to validate that the circularly permuted version of CapD has measurable activity for further assessments. We also hypothesize a threonine residue in the catalytic site of CapD_CP plays an important role in the catalysis reaction and mutating it will eliminate all enzymatic activity. Thus we created two mutants, T2V and T2A, to act as negative controls. The result (figure 1 below) of this assay confirms our hypothesis that CapD_CP has enzymatic activity to the two catalytic knockouts. The relatively flat activity curves of the knockout mutants confirm the hypothesis of the threonine's role in the catalytic site.
'Designing CapD_CP to be a better Hydrolase'
Screening Mutant Libraries
By standardizing the activity slope of each design relative to CapD_CP, a scatterplot easily portrays the qualities of each mutant. Several designs show negative catalytic curves similar to the catalytic knockouts. Some immediately show a negative activity curve meaning decrease in transpeptidation, hydrolysis, or both. T20S is a promising mutant hydrolase design.
Characterization of best hits
blah blah blah
 "
"