Team:TU Munich/Lab
From 2010.igem.org
(→PCR) |
|||
| Line 5: | Line 5: | ||
<!-- ############## WIKI-PAGE STARTS HERE ############## --> | <!-- ############## WIKI-PAGE STARTS HERE ############## --> | ||
| + | |||
| + | =Lab proceedings= | ||
| + | The idea behind our project is to change the way BioBricks have been used up to now. Over the years, many receptors and signals have been constructed as BioBricks during the annual iGEM competition, but still it is not possible to interconnect these Bricks in a complex biological network resuting in a cell, that is able to respond to its environment giving differenciated responses depending on the input signals. (Beispiel: cambridge hat das gemacht, xx dies, aber eine zelle kann nicht beides...<br> | ||
| + | We plan to create biological switches, that can function as locial gates inside a cell. Our switches rely on RNA/RNA-interactions, regulating transcriptional termination. This is a major advance of our concept, as regular switches rely on complex regulation including proteins and/or metabolites. Thus, our switches shall offer a greater robustness and their behaviour should be easier to predict. [[switch|Read more]] (hier sollte noch das hochskalieren erwähnt werden...<br> | ||
| + | These switches can further be used to build up a logical network inside a bacterial cell, enabling every scientist to connect as many functionalities (in form of BioBricks) as designated. We plan to offer simulation on each specifically designed network. | ||
| + | |||
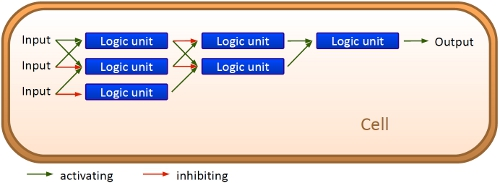
| + | <br><br>Over the years, many teams participating in the iGEM competition spent their time on constructing receptors and systems to detect a certain input that a variety of gorgeous oppurtunities is available so far.[[Image:TUM2010 network.png|thumb|300 px|right|Our visioon: A logic network inside the cell]] Nevertheless, until now it is not possible to link all those functionalities and build up a network giving differenciated responses to several of those input signals, where the molecular response depends on the complex composition of the environment a cell faces. We would like to offer this possibility to everyone. | ||
| + | <br> | ||
| + | The logic network we want to apply will be based on devices, that can be easily upscaled and therefor offer the chance to build networks of any wanted complexicity. Our devices rely on pure RNA/RNA interactions and thus their behaviour is well predictable. | ||
| + | <br> | ||
| + | |||
| + | The concept we rely on for our design of RNA-switches is based on the principle of [https://2010.igem.org/Team:TU_Munich/Glossary#Attenuation/ '''attenuation''']. | ||
| + | |||
| + | = Experiments = | ||
| + | We designed several experiments to test our switches, all of them based on fluorescence measurements. We designed experiment setting for measurements ''in vivo'' as well as ''in vitro''. Our ''in vitro'' measurements relied on two different experiment set-ups. While the first was based on a commercial ''E. coli''-lysate, the latter was reporting on a transcriptional level only, eliminating most of the possible side-effects one could expect in the complex behaviour of a living cell or cell-lysate. [[Experiments_main|Read more]] | ||
| + | |||
| + | = Results = | ||
| + | We ...blablabla | ||
| + | [[Resultss_main|Read more]] | ||
| + | |||
On this page you can find our protocols for standard molecular biology procedure as well as the full notebook containing lab progress. | On this page you can find our protocols for standard molecular biology procedure as well as the full notebook containing lab progress. | ||
Revision as of 13:38, 2 October 2010
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
Lab proceedingsThe idea behind our project is to change the way BioBricks have been used up to now. Over the years, many receptors and signals have been constructed as BioBricks during the annual iGEM competition, but still it is not possible to interconnect these Bricks in a complex biological network resuting in a cell, that is able to respond to its environment giving differenciated responses depending on the input signals. (Beispiel: cambridge hat das gemacht, xx dies, aber eine zelle kann nicht beides... Over the years, many teams participating in the iGEM competition spent their time on constructing receptors and systems to detect a certain input that a variety of gorgeous oppurtunities is available so far. Nevertheless, until now it is not possible to link all those functionalities and build up a network giving differenciated responses to several of those input signals, where the molecular response depends on the complex composition of the environment a cell faces. We would like to offer this possibility to everyone.
The concept we rely on for our design of RNA-switches is based on the principle of attenuation. ExperimentsWe designed several experiments to test our switches, all of them based on fluorescence measurements. We designed experiment setting for measurements in vivo as well as in vitro. Our in vitro measurements relied on two different experiment set-ups. While the first was based on a commercial E. coli-lysate, the latter was reporting on a transcriptional level only, eliminating most of the possible side-effects one could expect in the complex behaviour of a living cell or cell-lysate. Read more ResultsWe ...blablabla Read more
ProtocolsGelsAgarose Gels
Running of Gels mix samples 1:1 with formamide loading dye (stored @ -20°C) carefully remove comb blow air into pockets with a 50 µl syringe fill samples into pockets run the gel (usually about 200 V) PCR
used protocols
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Application | DNA Binding Buffer : Sample | Example |
| Plasmid, genomic DNA (>2 kb) | 2 : 1 | 200 μl : 100 μl |
| PCR, cDNA, DNA fragment | 5 : 1 | 500 μl : 100 μl |
| ssDNA (e.g., M13 phage) | 7 : 1 | 700 μl : 100 μl |
- Transfer mixture to a provided Zymo-Spin™ Column1 in a Collection Tube.
- Centrifuge at ≥10,000 x g for 30 seconds. Discard the flow-through.
- Add 200 μl Wash Buffer to the column. Centrifuge at ≥10,000 x g for 30 seconds. Repeat wash step.
- Add ≥6 μl water2,3 directly to the column matrix. Transfer the column to a 1.5 ml microcentrifuge tube and centrifuge at ≥10,000 x g for 30 seconds to elute the DNA.
Ultra-pure DNA in water is now ready for use.
QIAquick purification Kit
[http://www1.qiagen.com/literature/render.aspx?id=103715 Handbook]
Procedure
1. Add 5 volumes of Buffer PB to 1 volume of the PCR sample and mix. It is not necessary to remove mineral oil or kerosene. For example, add 500 μl of Buffer PB to 100 μl PCR sample (not including oil).
2. If pH indicator I has beein added to Buffer PB, check that the color of the mixture is yellow. If the color of the mixture is orange or violet, add 10 μl of 3 M sodium acetate, pH 5.0, and mix. The color of the mixture will turn to yellow.
3. Place a QIAquick spin column in a provided 2 ml collection tube.
4. To bind DNA, apply the sample to the QIAquick column and centrifuge for 30–60 s. We changed it to 3 min @ 6000rpm !
5. Discard flow-through. Place the QIAquick column back into the same tube. Collection tubes are re-used to reduce plastic waste.
6. To wash, add 0.75 ml Buffer PE to the QIAquick column and centrifuge for 30–60 s.
7. Discard flow-through and place the QIAquick column back in the same tube. Centrifuge the column for an additional 1 min.repeat!
IMPORTANT: Residual ethanol from Buffer PE will not be completely removed unless the flow-through is discarded before this additional centrifugation.
8. Place QIAquick column in a clean 1.5 ml microcentrifuge tube.
9. To elute DNA, add 50 μl Buffer EB (10 mM Tris·Cl, pH 8.5) or water (pH 7.0–8.5) to the center of the QIAquick membrane and centrifuge the column for 1 min. Alternatively, for increased DNA concentration, add 30 μl elution buffer to the center of the QIAquick membrane, let the column stand for 1 min, and then centrifuge.
IMPORTANT: Ensure that the elution buffer is dispensed directly onto the QIAquick membrane for complete elution of bound DNA. The average eluate volume is 48 μl from 50 μl elution buffer volume, and 28 μl from 30 μl elution buffer. Elution efficiency is dependent on pH. The maximum elution efficiency is achieved between pH 7.0 and 8.5. When using water, make sure that the pH value is within this range, and store DNA at –20°C as DNA may degrade in the absence of a buffering agent. The purified DNA can also be eluted in TE buffer (10 mM Tris·Cl, 1 mM EDTA, pH 8.0), but the EDTA may inhibit subsequent enzymatic reactions.
10. If the purified DNA is to be analyzed on a gel, add 1 volume of Loading Dye to 5 volumes of purified DNA. Mix the solution by pipetting up and down before loading the gel.
Gel samples
ZYMO RESEARCH Gel DNA Recovery Kit
[http://www.acgtinc.com/PDF_files/Sample%20Prepation_ACGT/Zymoclean%20Gel%20DNA%20Recovery%20Kit_Zymo%20Research.pdf Product informartion]
Protocol
- Excise the DNA fragment1 from the agarose gel using a razor blade or scalpel and transfer it to a 1.5 ml microcentrifuge tube.
- Add 3 volumes of ADB to each volume of agarose excised from the gel (e.g. for 100 μl (mg) of agarose gel slice add 300 μl of ADB).
- Incubate at 37-55 °C for 5-10 minutes until the gel slice is completely dissolved2. For DNA fragments >8 kb, following the incubation step, add one additional volume (equal to that of the gel slice) of water to the mixture for better DNA recovery (e.g. 100 μl agarose, 300 μl ADB and 100 μl water).
- Transfer the melted agarose solution to a Zymo-SpinTM I Column in a Collection Tube.
- Centrifuge at ≥10,000 x g for 30-60 seconds. Discard the flow-through.
- Add 200 μl of Wash Buffer to the column and centrifuge at ≥10,000 x g for 30 seconds. Discard the flow-through. Repeat the wash step.
- Add ≥6 μl of water3,4 directly to the column matrix. Place column into a 1.5 ml tube and centrifuge ≥10,000 x g for 30-60 seconds to elute DNA.
Ultra-pure DNA in water is now ready for use.
- add other kits here...
Restriction
| Enzyme | 10 units is sufficient, generally 1µl is used |
| DNA | 1 µg |
| 10X NEBuffer | 5 µl (1X) |
| BSA | Add to a final concentration of 100 µg/ml (1X) if necessary |
| Total Reaction Volume | 50 µl |
| Incubation Time | 1 - 1.5 hour |
| Incubation Temperature Enzyme dependent |
XbaI, SpeI, PstI, SpeI : 37 °C |
[http://www.neb.com/nebecomm/tech_reference/restriction_enzymes/buffer_activity_restriction_enzymes.asp activity of restriction enzymes in NEB buffers]
Biobrick standard
[http://openwetware.org/wiki/Restriction_digest Protocols for IGEM standard digestion]
Dephosphorylation
using Antarctic Phosphatase
- Add 1/10 volume of 10X Antarctic Phosphatase Reaction Buffer to 1-5 µg of DNA cut with any restriction endonuclease in any buffer.
- Add 1 µl of Antarctic Phosphatase (5 units) and mix.
- Incubate for 15 minutes at 37°C for 5´ extensions or blunt-ends, 60 minutes for 3´ extensions.
- Heat inactivate (or as required to inactivate the restriction enzyme) for 5 minutes at 65°C.
- Proceed with ligation.
[http://www.neb.com/nebecomm/products/protocol76.asp from NEB]
Ligation
Using T4 Ligase, New England Labs
- 1 µl T4 Ligase (10.000 U)
- 50 ng plasmid
- 3x mol(plasmid) insert
- 2 µl T4 Ligase 10x buffer
- add H2O to reach final volume of 20 µl
- incubation at 22°C for 1 h
- storing at 16 °C for 40 min
Biobrick Standard
[http://parts2.mit.edu/wiki/index.php/Standard_Assembly Standard BioBrick assembly]
Transformation
At Woehlke's S1-Lab !!!
- Thaw competent cells on Ice
- Add DNA, pipette gently to mix
- Let sit for 30 minutes on ice
- Incubate cells for 45 seconds at 42°C
- Incubate cells on ice for 2 min
- Add 1 ml LB0
- Incubate for 1 hour at 37oC on shaker
- Spread 100-300 μl onto a plate made with appropriate antibiotic.
- Grow overnight at 37 °C.
- Save the rest of the transformants in liquid culture at 4 °C
modified from [http://openwetware.org/wiki/Transforming_chemically_competent_cells open wetware]
Miniprep
Preparation of BioBricks from distribution 2008
Sequencing
- Monsterplasmids contain GATC-Standard-Primer pBR1 (CGAAAAGTGCCACCTGAC ) directly in front of AATII cleavage site.
- Monsterplasmids contain GATC-Standard-Primer pGFP-FP () approx. 100 bp upstream of Biobrick insert site.
!!! Please always fill in iGEM-Sequencing-YYMMDD (e.g. iGEM-Sequencing-100625 for today´s date) as internal billing number!
 "
"