Team:Newcastle/Filamentous Cells
From 2010.igem.org
(Difference between revisions)
RachelBoyd (Talk | contribs) (New page: {{Team:Newcastle/mainbanner}} '''Filamentous Cells''' {| |- |SOS response is believe to be a universal bacteria phenomenon first studied in E.coli -LexA, recA |- |In Bacillus subtillis (...) |
|||
| Line 3: | Line 3: | ||
{| | {| | ||
|- | |- | ||
| - | |SOS response is believe to be a universal bacteria phenomenon first studied in E.coli -LexA, recA | + | |SOS response is believe to be a universal bacteria phenomenon first studied in ''E.coli'' -LexA, recA |
|- | |- | ||
| - | |In Bacillus subtillis (gram positive) dinR protein is homologous to lexA (Repressor of din-damage inducible genes).din genes include uvrA, uvrB, dinB, dinC dinR and recA. DNA damage inhibits cell division. | + | |In ''Bacillus subtillis'' (gram positive) ''dinR'' protein is homologous to ''lexA'' (Repressor of din-damage inducible genes). din genes include ''uvrA'', ''uvrB'', ''dinB'', ''dinC'' ''dinR'' and ''recA''. DNA damage inhibits cell division. |
|- | |- | ||
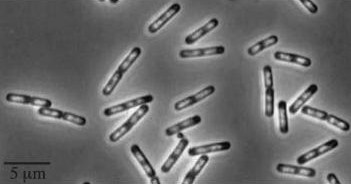
| - | | Wild type Bacillus subtillis | + | | Wild type ''Bacillus subtillis'' |
|- | |- | ||
|[[Image:Wild type Bacillus subtillis.jpg]] | |[[Image:Wild type Bacillus subtillis.jpg]] | ||
|- | |- | ||
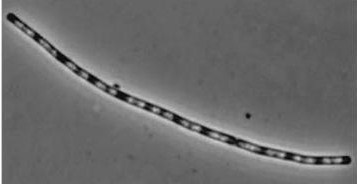
| - | |dinR KO | + | |''dinR'' KO |
|- | |- | ||
|[[Image:dinR KO.jpg]] | |[[Image:dinR KO.jpg]] | ||
|- | |- | ||
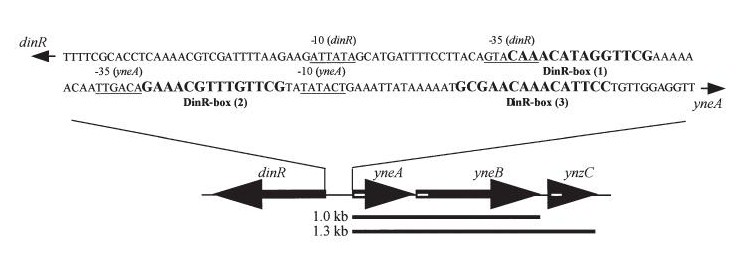
| - | |dinR KO mutant over expressed the divergent (opposite direction) transcript for YneA, YneB and YnzC. These genes form the SOS regulon (recA independent SOS response) | + | |''dinR'' KO mutant over expressed the divergent (opposite direction) transcript for YneA, YneB and YnzC. These genes form the SOS regulon (''recA'' independent SOS response) |
|- | |- | ||
| [[Image:Coding region.jpg]] | | [[Image:Coding region.jpg]] | ||
| Line 25: | Line 25: | ||
|Disruption of YneA in SOS response leads to reduced elongation. Altering YneB and YnzC expression does not affect cell morphology. | |Disruption of YneA in SOS response leads to reduced elongation. Altering YneB and YnzC expression does not affect cell morphology. | ||
|- | |- | ||
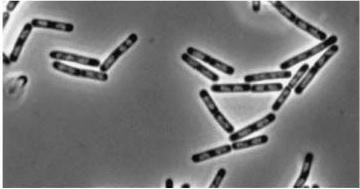
| - | |Double mutant (dinR | + | |Double mutant (''dinR'' YneA) |
|- | |- | ||
|[[Image:Double Mutant.jpg]] | |[[Image:Double Mutant.jpg]] | ||
| Line 34: | Line 34: | ||
assembling other proteins necessary for division at the site. | assembling other proteins necessary for division at the site. | ||
|- | |- | ||
| - | |FtsZ localises to the cell division cycle unless dinR is disrupted or YneA is being induced. | + | |FtsZ localises to the cell division cycle unless ''dinR'' is disrupted or YneA is being induced. |
|- | |- | ||
|YneA suppresses FtsZ ring formation- no proven direct interaction by two-hybrid. | |YneA suppresses FtsZ ring formation- no proven direct interaction by two-hybrid. | ||
| Line 40: | Line 40: | ||
|Filamentous cells less colony formation. | |Filamentous cells less colony formation. | ||
|- | |- | ||
| - | |YneA expression via the inactivation of dinR by Rec A is important. | + | |YneA expression via the inactivation of ''dinR'' by Rec A is important. |
|- | |- | ||
| - | |'''Kawai, Y., Moriya, S., & Ogasawara, N. (2003). Identification of a protein, YneA, responsible for cell division suppression during the SOS response in Bacillus subtilis. Molecular microbiology, 47(4), 1113-22. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/12581363.''' | + | |} |
| + | '''Kawai, Y., Moriya, S., & Ogasawara, N. (2003). Identification of a protein, YneA, responsible for cell division suppression during the SOS response in Bacillus subtilis. Molecular microbiology, 47(4), 1113-22. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/12581363.''' | ||
Revision as of 23:37, 3 June 2010

| |||||||||||||
| |||||||||||||
Filamentous Cells
 "
"