Team:Imperial College London/Modelling
From 2010.igem.org
| Line 14: | Line 14: | ||
|style="font-family: helvetica, arial, sans-serif;font-size:1.2em;color:#ea8828;" align="right"|[[Team:Imperial_College_London/Modelling/Protein_Display/Objectives|Objectives]] | [[Team:Imperial_College_London/Modelling/Protein_Display/Detailed_Description|Description]] | [[Team:Imperial_College_London/Modelling/Protein_Display/Parameters_and_Constants|Constants]] | [[Team:Imperial_College_London/Modelling/Protein_Display/Results_and_Conclusion|Results]] | [[Team:Imperial_College_London/Modelling/Protein_Display/Download_MatLab_Files|MATLAB Code]] | |style="font-family: helvetica, arial, sans-serif;font-size:1.2em;color:#ea8828;" align="right"|[[Team:Imperial_College_London/Modelling/Protein_Display/Objectives|Objectives]] | [[Team:Imperial_College_London/Modelling/Protein_Display/Detailed_Description|Description]] | [[Team:Imperial_College_London/Modelling/Protein_Display/Parameters_and_Constants|Constants]] | [[Team:Imperial_College_London/Modelling/Protein_Display/Results_and_Conclusion|Results]] | [[Team:Imperial_College_London/Modelling/Protein_Display/Download_MatLab_Files|MATLAB Code]] | ||
|} | |} | ||
| - | We came up with a novel idea of detecting organisms that we do not have a specific receptor for. In our particular example, the protease of Schistosoma is meant to cleave a protein displayed on the bacteria's cell wall. The cleaved peptide is supposed to be recognized by the receptor which would activate the colour expression. This solution raised questions about the risk of false positive or whether there are any chances for ComD receptors to be activated in the diluted environment. Modelling of this module would enable us to answer these questions.<br/><br/> | + | We came up with a novel idea of detecting organisms that we do not have a specific receptor for. In our particular example, the protease of Schistosoma is meant to cleave a protein displayed on the bacteria's cell wall (<i>the pink rectangle on the diagram below</i>). The cleaved peptide is supposed to be recognized by the receptor which would activate the colour expression. This solution raised questions about the risk of false positive or whether there are any chances for ComD receptors to be activated in the diluted environment. Modelling of this module would enable us to answer these questions.<br/><br/> |
{| style="width:800px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;" | {| style="width:800px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;" | ||
|2. <b>Signalling Model</b> | |2. <b>Signalling Model</b> | ||
|style="font-family: helvetica, arial, sans-serif;font-size:1.2em;color:#ea8828;" align="right"|[[Team:Imperial_College_London/Modelling/Signalling/Objectives|Objectives]] | [[Team:Imperial_College_London/Modelling/Signalling/Detailed_Description|Description]] | [[Team:Imperial_College_London/Modelling/Signalling/Parameters_and_Constants|Constants]] | [[Team:Imperial_College_London/Modelling/Signalling/Results_and_Conclusion|Results]] | [[Team:Imperial_College_London/Modelling/Signalling/Download_MatLab_Files|MATLAB Code]] | |style="font-family: helvetica, arial, sans-serif;font-size:1.2em;color:#ea8828;" align="right"|[[Team:Imperial_College_London/Modelling/Signalling/Objectives|Objectives]] | [[Team:Imperial_College_London/Modelling/Signalling/Detailed_Description|Description]] | [[Team:Imperial_College_London/Modelling/Signalling/Parameters_and_Constants|Constants]] | [[Team:Imperial_College_London/Modelling/Signalling/Results_and_Conclusion|Results]] | [[Team:Imperial_College_London/Modelling/Signalling/Download_MatLab_Files|MATLAB Code]] | ||
|} | |} | ||
| - | We decided to use the ComCDE signalling pathway from ''S.pneumoniae'' and so questions arose on whether it would work appropriately in ''B.subtilis''. We modelled this system to make sure that the signalling pathway would be working as anticipated.<br/><br/> | + | We decided to use the ComCDE signalling pathway from ''S.pneumoniae'' and so questions arose on whether it would work appropriately in ''B.subtilis''. We modelled this system to make sure that the signalling pathway would be working as anticipated (<i>the green rectangle on the diagram below</i>).<br/><br/> |
{| style="width:800px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;" | {| style="width:800px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;" | ||
|3. <b>Fast Response Model</b> | |3. <b>Fast Response Model</b> | ||
|style="font-family: helvetica, arial, sans-serif;font-size:1.2em;color:#ea8828;" align="right"|[[Team:Imperial_College_London/Modelling/Output/Objectives|Objectives]] | [[Team:Imperial_College_London/Modelling/Output/Detailed_Description|Description]] | [[Team:Imperial_College_London/Modelling/Output/Parameters_and_Constants|Constants]] | [[Team:Imperial_College_London/Modelling/Output/Results_and_Conclusion|Results]] | [[Team:Imperial_College_London/Modelling/Output/Download_MatLab_Files|MATLAB Code]] | |style="font-family: helvetica, arial, sans-serif;font-size:1.2em;color:#ea8828;" align="right"|[[Team:Imperial_College_London/Modelling/Output/Objectives|Objectives]] | [[Team:Imperial_College_London/Modelling/Output/Detailed_Description|Description]] | [[Team:Imperial_College_London/Modelling/Output/Parameters_and_Constants|Constants]] | [[Team:Imperial_College_London/Modelling/Output/Results_and_Conclusion|Results]] | [[Team:Imperial_College_London/Modelling/Output/Download_MatLab_Files|MATLAB Code]] | ||
|} | |} | ||
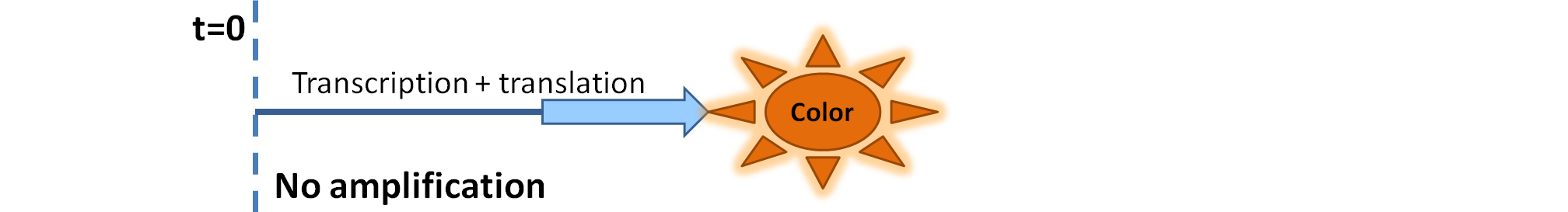
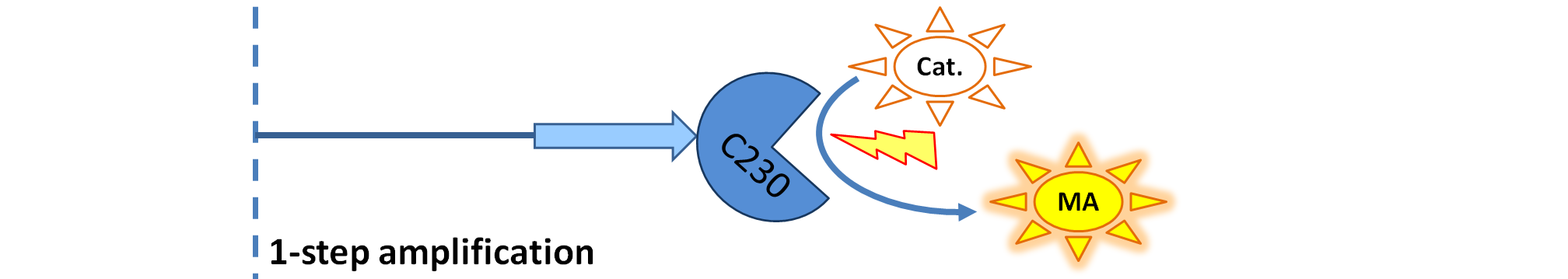
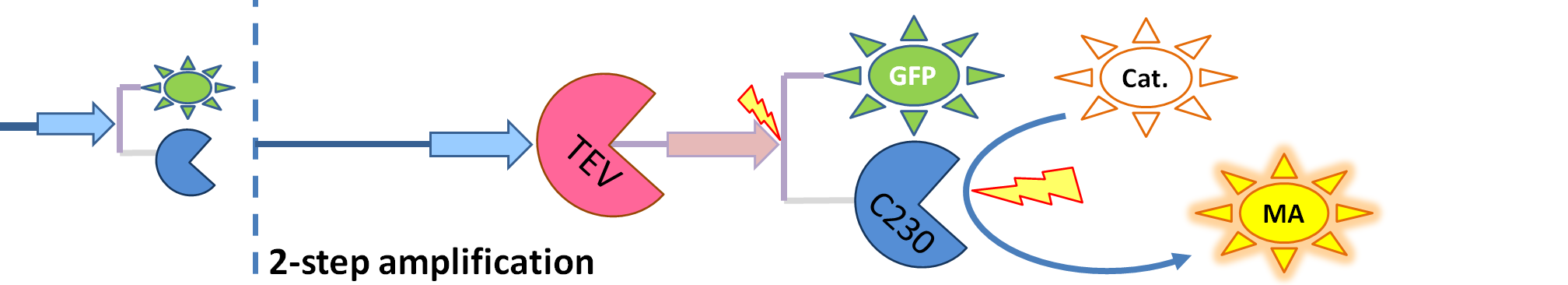
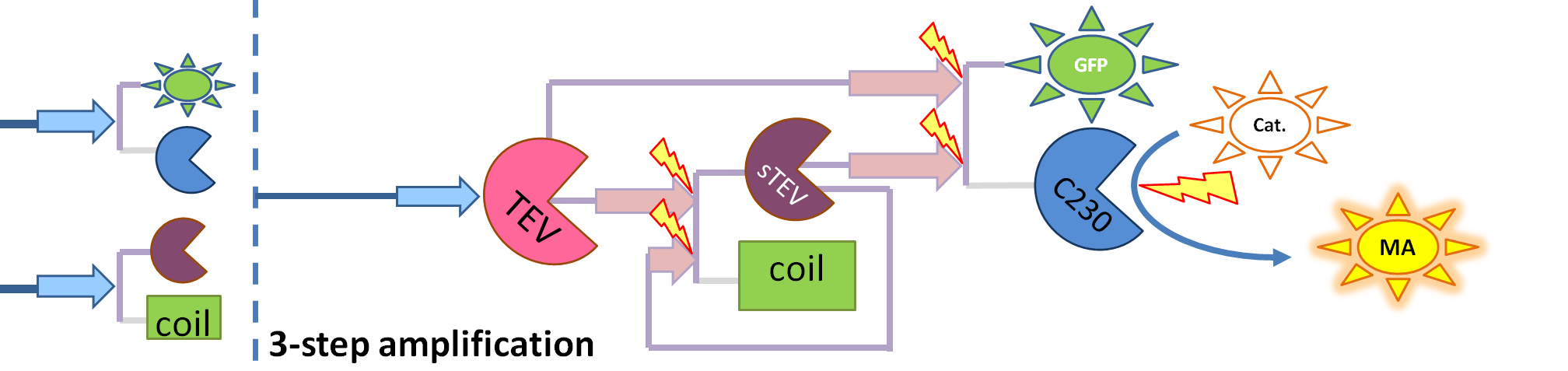
| - | We came up with an idea of using the amplification of a colour output, which would show within minutes of the stimulus being added. The question that arose was whether amplification will actually perform better than simple production (i.e. transcription and translation) in the cellular environment. Furthermore, we had difficulties deciding whether we should design the amplification module to consist of 1,2 or even more amplification steps. These issues were quite difficult to answer, so we decided to employ modelling.<br/> | + | We came up with an idea of using the amplification of a colour output, which would show within minutes of the stimulus being added (<i>the blue rectangle on the diagram below</i>). The question that arose was whether amplification will actually perform better than simple production (i.e. transcription and translation) in the cellular environment. Furthermore, we had difficulties deciding whether we should design the amplification module to consist of 1,2 or even more amplification steps. These issues were quite difficult to answer, so we decided to employ modelling.<br/> |
</ol> | </ol> | ||
| + | <div ALIGN=CENTER> | ||
| + | {| style="width:404px;background:#e7e7e7;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;margin- top:5px;padding: 2px;" cellspacing="5"; | ||
| + | |- | ||
| + | |[[Image:IC_Overview.jpg|400px]] | ||
| + | |- | ||
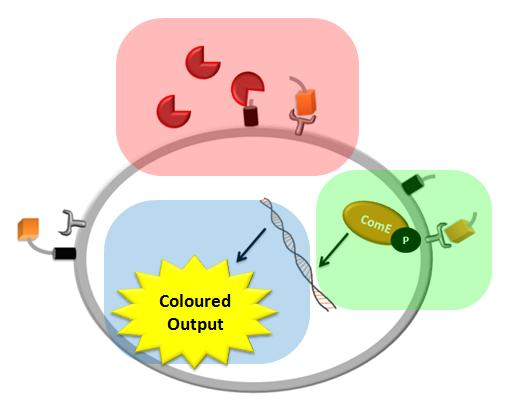
| + | |Each of the shadowed rectangles corresponds to the model of a particular module of the system. Pink corresponds to the model of the detection mechanism, green corresponds to the model of the signal transduction and blue corresponds to the model of the fast response module. | ||
| + | |} | ||
| + | </div> | ||
|} | |} | ||
Revision as of 01:19, 28 October 2010
| Modelling | Overview | Detection Model | Signaling Model | Fast Response Model | Interactions |
| A major part of the project consisted of modelling each module. This enabled us to decide which ideas we should implement. Look at the Fast Response page for a great example of how modelling has made a major impact on our design! | |
| Introduction to modelling | ||||||
| During the design of our construct three major questions arose which could be answered by computer modelling:
We came up with a novel idea of detecting organisms that we do not have a specific receptor for. In our particular example, the protease of Schistosoma is meant to cleave a protein displayed on the bacteria's cell wall (the pink rectangle on the diagram below). The cleaved peptide is supposed to be recognized by the receptor which would activate the colour expression. This solution raised questions about the risk of false positive or whether there are any chances for ComD receptors to be activated in the diluted environment. Modelling of this module would enable us to answer these questions.
We decided to use the ComCDE signalling pathway from S.pneumoniae and so questions arose on whether it would work appropriately in B.subtilis. We modelled this system to make sure that the signalling pathway would be working as anticipated (the green rectangle on the diagram below).
We came up with an idea of using the amplification of a colour output, which would show within minutes of the stimulus being added (the blue rectangle on the diagram below). The question that arose was whether amplification will actually perform better than simple production (i.e. transcription and translation) in the cellular environment. Furthermore, we had difficulties deciding whether we should design the amplification module to consist of 1,2 or even more amplification steps. These issues were quite difficult to answer, so we decided to employ modelling. |
| Results & Conclusions |
Detection Model
|
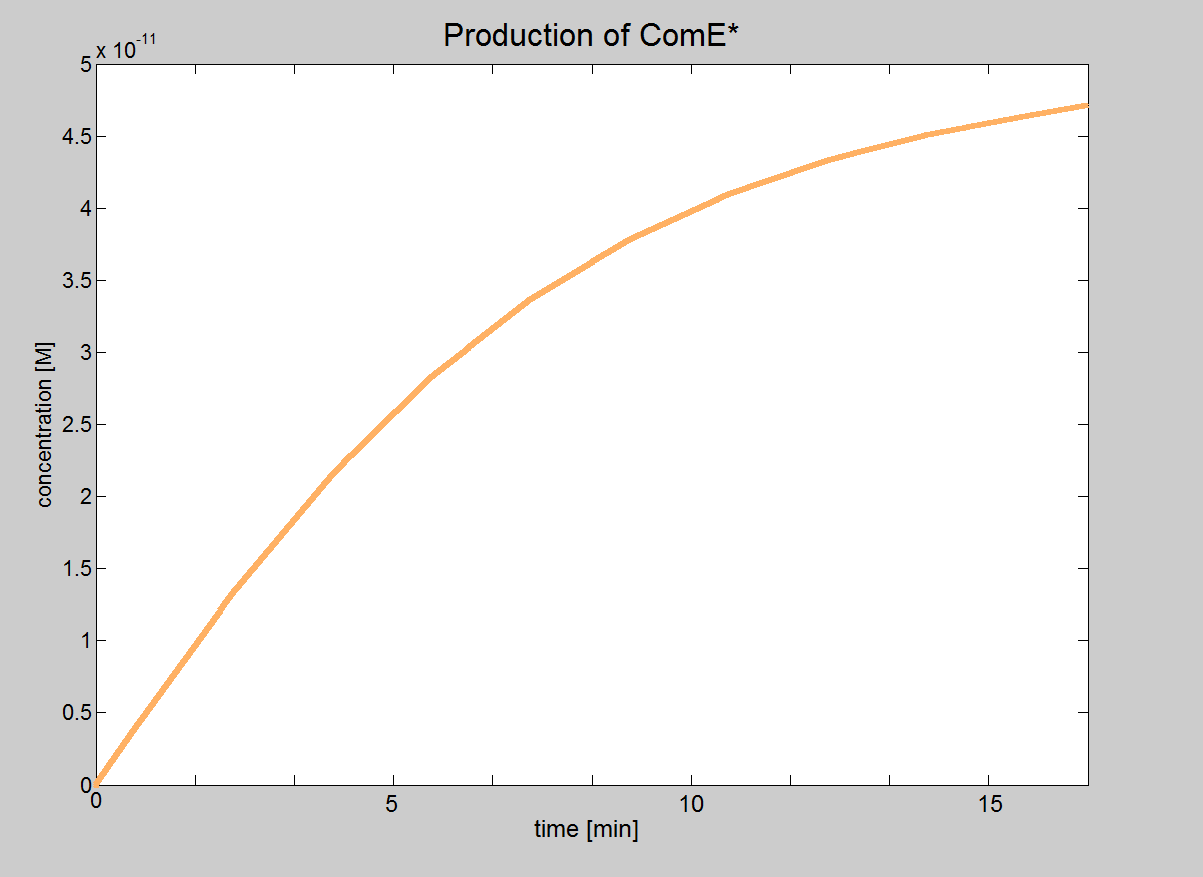
| Signalling Model Even though our model of the signalling module is more simplistic than the real life situation, it provided very important results. We were able to determine under which conditions the signalling pathway would be working and could obtain the major constraints of our system. These constraints are that the necessary concentrations for ComD and AIP are reached before signal transduction is started. In this model, we will treat phosphorylated ComE (referred to as ComE*) as our "output". Phosphorylated ComE is a transcription factor, which will bind to the DNA to enable transcription. |
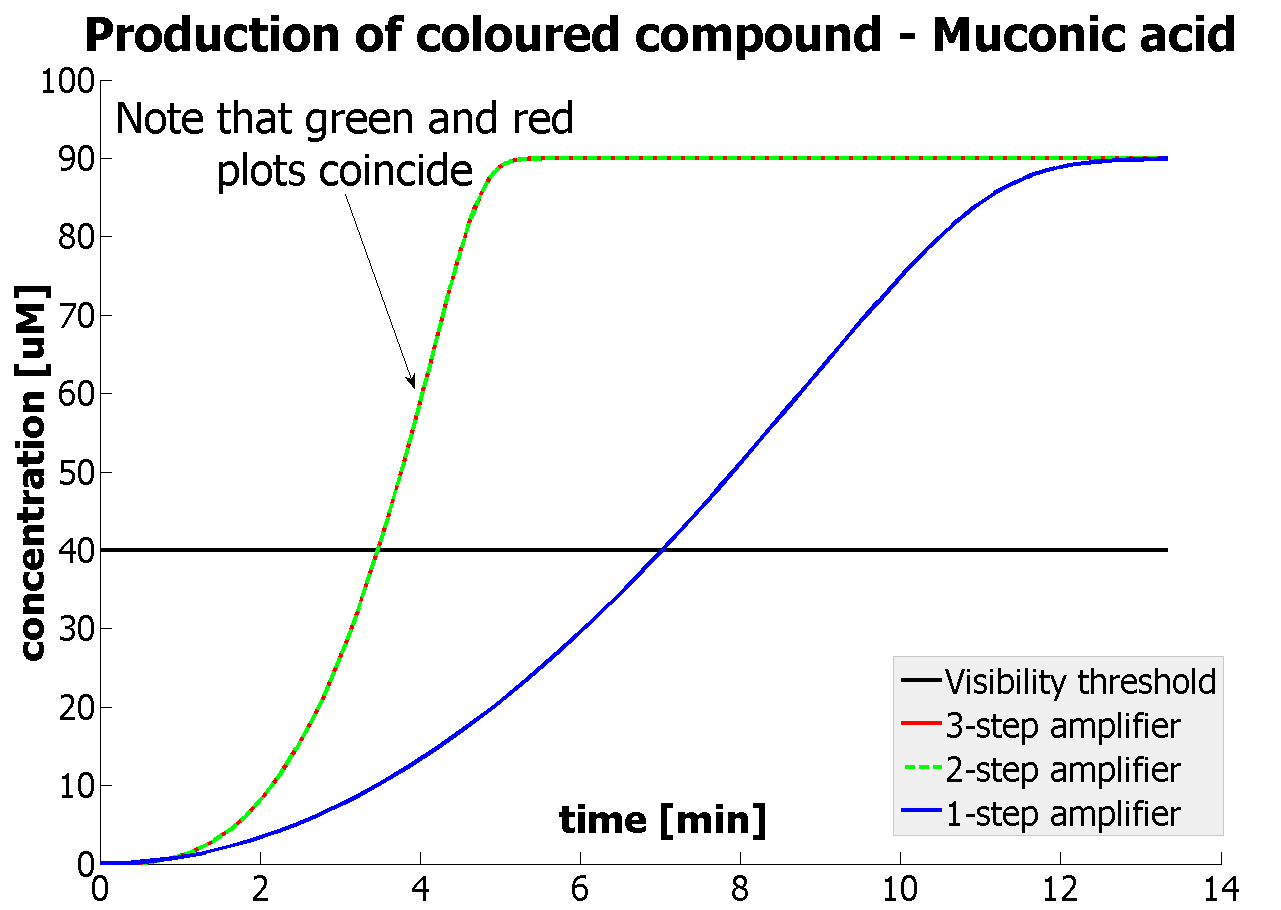
Fast Response Model
|
 "
"