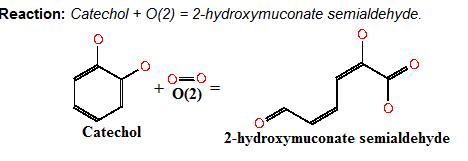Team:Imperial College London/Modules/Fast Response
From 2010.igem.org
| Line 36: | Line 36: | ||
</div> | </div> | ||
| | | | ||
| - | Catechol (2,3) dioxygenase ( | + | Catechol (2,3) dioxygenase (C23O) is the protein product of the XylE gene. It originates from ''Pseudomonas putida'' and the active protein is made up of a homotetramer of C2,3O monomers. The enzyme catalyses the conversion of a colourless substrate (catechol or substituted catechols) into a bright yellow product (2-hydroxymuconic semialdehyde) within seconds of substrate addition. |
|} | |} | ||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
Revision as of 00:19, 28 October 2010
| Modules | Overview | Detection | Signaling | Fast Response |
| Our design consists of three modules; Detection, Signaling and a Fast Response, each of which can be exchanged with other systems. We used a combination of modelling and human practices to define our specifications. Take a look at the overview page to get a feel for the outline, then head to the full module pages to find out how we did it. | |
| Fast Response Module | Overview of the output |
|
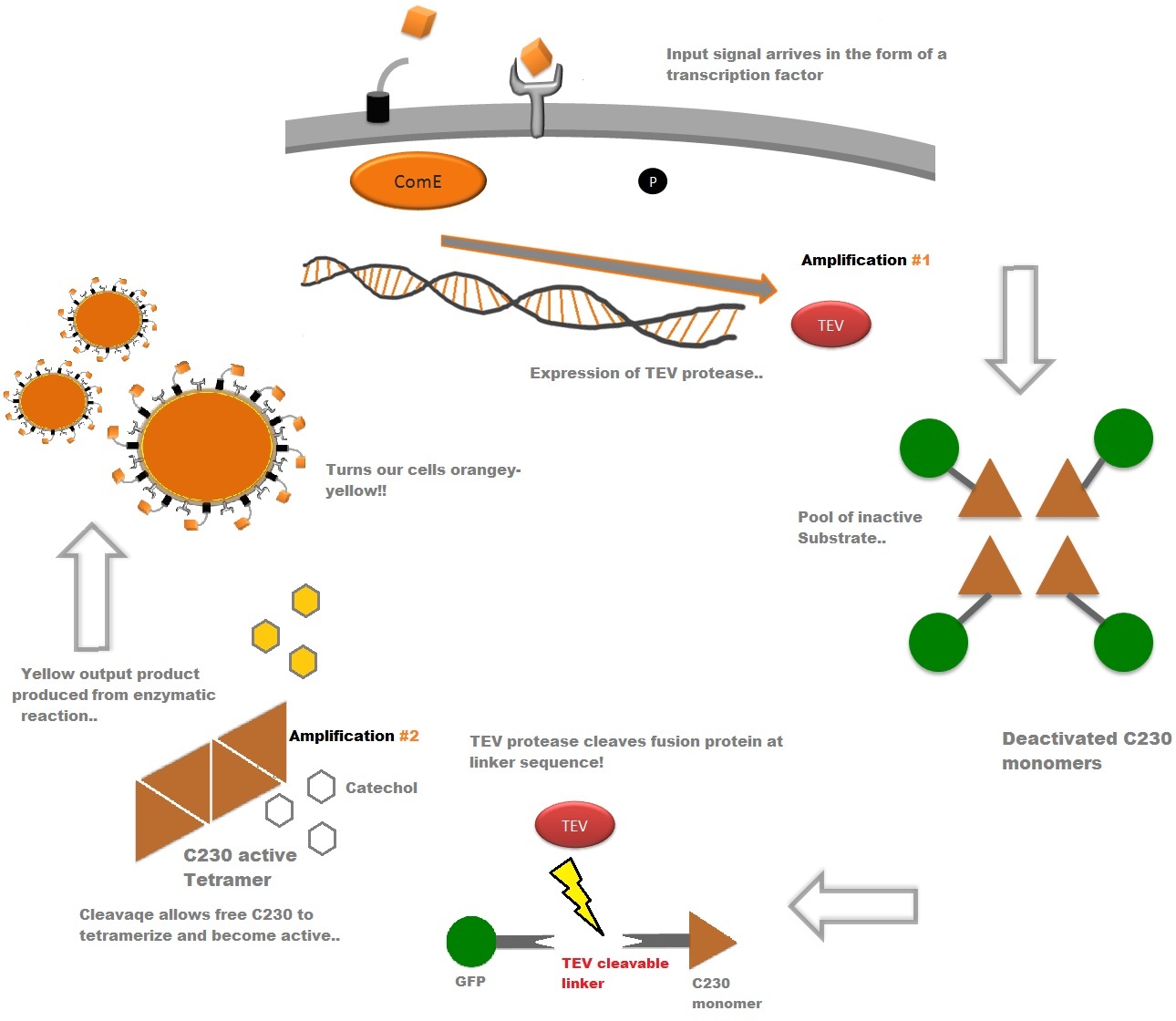
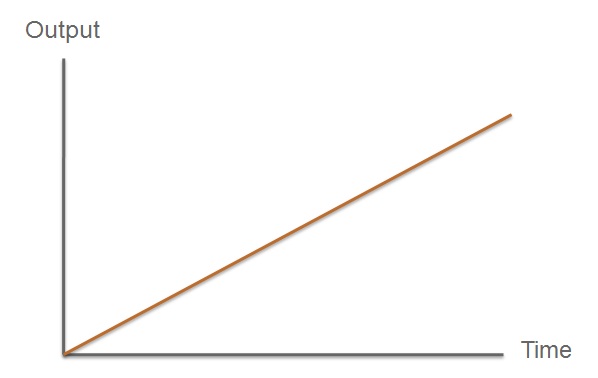
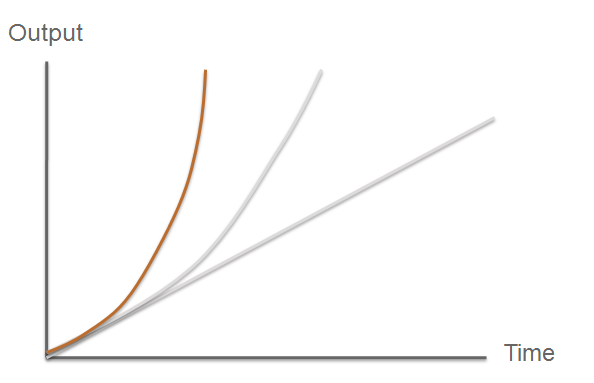
The third module of our bacterial detector consists of a fast response reporter module that upon an input signal arrival gives out a yellow signal output visible by the naked eye. Traditional reporters in biochemical research usually rely on transcription-translation of a specific reporter molecule (such as Green Fluorescent protein) which is time consuming and/or might involve sophisticated equipment for its detection. In this module we introduce a novel reporter system where expression of a low molecular weight protease allows this protease to act on an existing pool of substrate. Due to this double amplification step, the output signal should be several orders of magnitude faster than traditional reporters. Below is a quick overview of our system, followed by a more in-depth analysis of the system. |
| Video of a C230 assay! |
| C230 and C230-GFP fusion | |
|
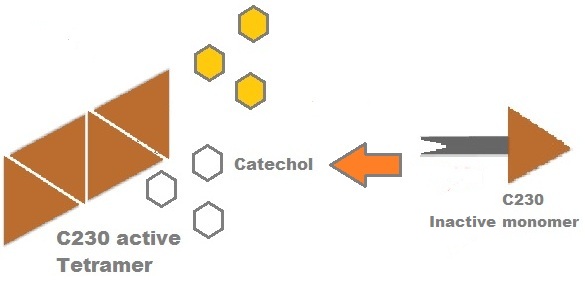
Catechol (2,3) dioxygenase (C23O) is the protein product of the XylE gene. It originates from Pseudomonas putida and the active protein is made up of a homotetramer of C2,3O monomers. The enzyme catalyses the conversion of a colourless substrate (catechol or substituted catechols) into a bright yellow product (2-hydroxymuconic semialdehyde) within seconds of substrate addition. | |
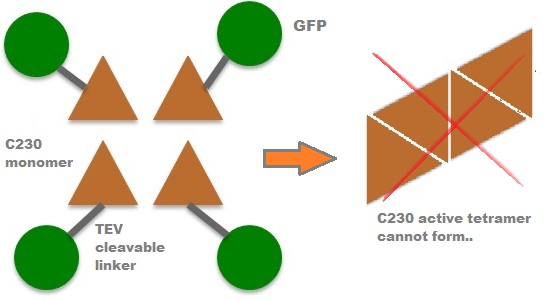
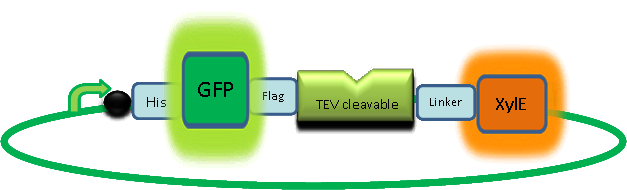
| In our project, we take advantage of the fact that Catechol-2,3-dioxygenase is only active as a tetramer as the active site forms as a result of intersubunit and interface interactions after oligomerization. We have constructed a gene product that encodes a fusion protein. This fusion protein is made of GFP fused to the N-terminus of Catechol-2,3-dioxygenase. The GFP does not allow Catechol-2,3-dioxygenase monomers to associate due to a steric hindrance effect, so that the active tetramer cannot form. Instead the fusion protein remains as a monomer in the cytosol of the cell. |
|
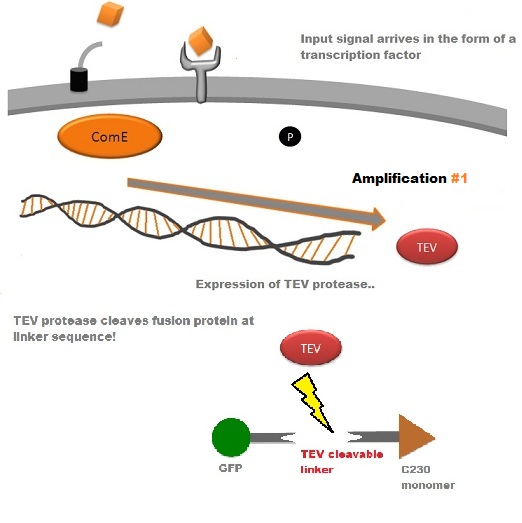
A very important feature of the fusion protein is its potential for induction of the system. A closer look at the fusion gene construct reveals the secret of the inducibility of our system. The GFP gene is fused to the XylE gene through a protein sequence that is recognised and cleaved by TEV protease. This amino acid sequence is recognized by the active site of the TEV protease so that when TEV is present in the cell, GFP is cleaved off Catechol-2,3-dioxygenase monomers. The free monomers are no longer sterically hindered by the GFP and are now able to associate with each other. Tetramerization of the monomers produces the active Catechol-2,3-dioxygenase enzyme, which in the presence of catechol substrate produces a yellow color. |
| Our GFP-XylE gene construct has also some other addition features, which make it easier to work with. To start with it contains an N-terminal His tag which would assist in purification and for in vitro characterization. The same applies to the flag tag sequence which can be recognized by monoclonal antibodies for easier detection and purification. The linker sequence is there to ensure that TEV recognition sequence is readily accessible to the protease for cleavage. |
| TEV protease | |
|
TEV protease is a natural polyprotein cutting viral protease. In our system ComE transcription factors bind to their native ComCDE promoter which was engineered as the promoter for TEV protease. This means that TEV protease is only transcribed and translated upon detection of the Schistosoma parasite. Why TEV protease? This enzyme has been used extensively in protein engineering as it has a number of key qualities including a high specificity (ENYLFQG), and importantly for our system it has a relatively low molecular weight (242 aa) compared to other protease candidates available. This means that the synthesis of the protease will be relatively quick. For more information about TEV, please see the PDB file with the accession number 1Q31. TEV protease expression in E.Coli (and consequently B .subtilis) is made efficient by codon optimisation and inducing expression of additional tRNAs. |
| Fast < faster < Imperial College Turbo FAST | |
|
So what are the key features that make our system a novel example of a fast response mechanism? | Modelling |
| |
| |
| |
| C23O | Additional Material | |||
| After observing C2,3O in Pymol, and in addition information obtained from Kita et al, which shows that the projecting loop is needed for dimerization and that this is very near to the N-terminus, we chose to engineer (by straightforward DNA synthesis) GFP plus linker (GGGSGGGS ENYLFQG) onto the N-termini of our C230 monomers.
Below is the reaction performed by our systems enzyme:
|
| References |
|
[1] An archetypical extradiol-cleaving catecholic dioxygenase: the crystal structure of catechol 2,3-dioxygenase (metapyrocatechase) from Pseudomonas putida mt-2 Akiko Kita1, Shin-ichi Kita2, Ikuhide Fujisawa1, Koji Inaka3, Tetsuo Ishida4, Kihachiro Horiike4, Mitsuhiro Nozaki4 and Kunio Miki5, |
 "
"