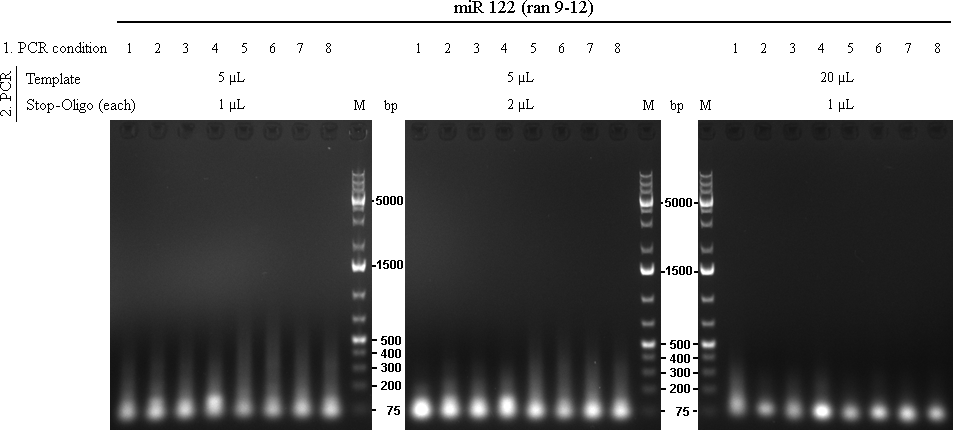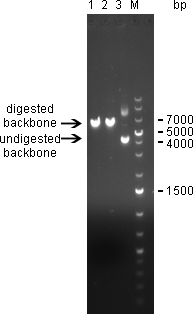Team:Heidelberg/Notebook/BSDesign/September
From 2010.igem.org
(→08/09/2010) |
(→08/09/2010) |
||
| Line 147: | Line 147: | ||
As it is known !!!reference!!! that multiple miRNA binding sites in a row increase the knock-down efficiency of miRNAs, binding patterns may help to find the right threshold for specific tissue targeting. | As it is known !!!reference!!! that multiple miRNA binding sites in a row increase the knock-down efficiency of miRNAs, binding patterns may help to find the right threshold for specific tissue targeting. | ||
| - | Therefore we created a spacer sequence by rational design with a GC content of 50%. This spacer need to be as innert as possible, in terms of miRNA recognition, so the sequence was then tested for compatibility with other miRNAs. We found | + | Therefore we created a spacer sequence to seperate the single binding sites by rational design with a GC content of 50%. This spacer need to be as innert as possible, in terms of miRNA recognition, so the sequence was then tested for compatibility with other miRNAs. We found two 15bp-spacer, which make up a total 30bp-spacer, with low compatiblity to any miRNA given by the tools mentioned. Highest achieved mean free energy was higher than -30 kcal/mol (the higher, the less effective is the binding), whereas a perfect binding miRNA was, in our cases, always under -40 kcal/mol. |
| - | + | ||
| - | + | ||
For our random assembly PCR approach, we created oligos using this spacer-sequence as annealing site. Therefore we split the spacer in two halfs, and placed a miRNA binding site in between: | For our random assembly PCR approach, we created oligos using this spacer-sequence as annealing site. Therefore we split the spacer in two halfs, and placed a miRNA binding site in between: | ||
:<span style="color:#78b41e">CACTGAATCCAACTG</span>-miRNA binding site-<span style="color:#009be1">GCATACATGGACTGC</span> | :<span style="color:#78b41e">CACTGAATCCAACTG</span>-miRNA binding site-<span style="color:#009be1">GCATACATGGACTGC</span> | ||
| - | + | Spacer sequence: | |
| + | : <span style="color:#009be1">SECOND HALF</span><span style="color:#78b41e">FIRST HALF</span> | ||
| + | :<span style="color:#009be1">GCATACATGGACTGC</span><span style="color:#78b41e">CACTGAATCCAACTG</span> | ||
| - | Additionally | + | Additionally we |
---- | ---- | ||
Revision as of 14:50, 27 October 2010

Binding Site Design - September08/09/2010Oligo design for random assembly PCR (raPCR) Using endogenous miRNAs for cell-identification, a detectable difference in miRNA-expression levels need to be present. Therefore we sent isolated RNA from HeLa, HUH7 and HEK-293 cells either infected with AAV or non-infected to [http://www.febit.com febit] for microarray analysation of micro-RNA expression levels. The obtained results were delivered in an evaluated form, showing relative expression levels between the different cell lines and/or conditions. For further experiments, the highest relative differences of two approaches were looked up:
1. comparison in one cell line between control and AAV infected status
Positive or negative log-values indicate upregulation or downregulation in AAV infected cells compared to non-infected cells of the same cell type, respectively.
2. comparison in one condition between different cell linesHere are the most up- and downregulated miRNAs shown. One of those were picked for further experiments.
Beside this, it is known that hsa-mir-122 is expressed in liver uniquely, at least for humans. Additionally, mm-mir-375/376a are uniquely expressed in mouse livers. As we are heading for divergent prospective experiments, first we want to show that the principle of using endogenous miRNAs for tissue identification tasks is compatible with our constructs for luciferase measurements (compare to Synthetic miRNA-Kit) and our new measurement standard miMeasure. As it is known !!!reference!!! that multiple miRNA binding sites in a row increase the knock-down efficiency of miRNAs, binding patterns may help to find the right threshold for specific tissue targeting. Therefore we created a spacer sequence to seperate the single binding sites by rational design with a GC content of 50%. This spacer need to be as innert as possible, in terms of miRNA recognition, so the sequence was then tested for compatibility with other miRNAs. We found two 15bp-spacer, which make up a total 30bp-spacer, with low compatiblity to any miRNA given by the tools mentioned. Highest achieved mean free energy was higher than -30 kcal/mol (the higher, the less effective is the binding), whereas a perfect binding miRNA was, in our cases, always under -40 kcal/mol. For our random assembly PCR approach, we created oligos using this spacer-sequence as annealing site. Therefore we split the spacer in two halfs, and placed a miRNA binding site in between:
Spacer sequence:
Additionally we 13/09/2010
Assay:
Restriction digest was performed for approx. 5h
stop-oligos: raPCR_AS13-stop_rev_NotI and raPCR_AS13-stop_fw_XhoI (ra017/018) Oligos were used in standard conc. (100µM)
for the next PCR, three assay will tried:
stop-oligos: raPCR_AS13-stop_rev_NotI and raPCR_AS13-stop_fw_XhoI
In total there were 72 reactions. Each was run with 2x Phusion Mastermix, missing volume was filled with water.
DNA was stored in fridge afterwards
14/09/2010The 72 PCRs from 13/10/2010 were analysed on 1% agarose gel.
File:2010-09-14 - raPCR miR122-klein fertig.png raPCR using miR-122 oligos The initial 8 PCRs from the first raPCR were purified using Qiagen PCR Purification Kit, eluted in 32 µL and used as template for the second raPCR.
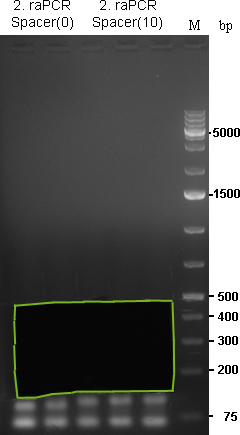
As we are looking for multiple binding sites, lanes with longest smear, meaning more long binding sites, were choosen for preparative gel:
As the gel volume was too much for dissolving in a single 2 mL tube, each part was splitted for dna extraction and brought together on the column. Gel extractions were done according to Qiagen Gel-extraction kit protocal and eluted in 30 µL water.
15/09/2010Sample code:
First row of cloning will be done with miR-122 samples. Others will follow. Samples prepared: 122-0S,-0L,-10S,-10L Gel extracted samples were digested with NotI/XhoI for cloning into psiCheck-2 vector: 5µL DNA (2µL for backbone) in a total volume of 30µL using 1µL XhoI and 0.6µL NotI enzyme, for 1.5 h at 37°C. The digested DNA was then purified using Qiagen nucleotide removal kit and eluted in 30µL. Subsequently, digested fragments were ligated over night at room temperature. Ligation assay for Fermentas T4 ligase: 2µL Buffer 1µL Ligase 7µL water 1µL Backbone (6000bp, psiCHECK-2) 9µL purified restriction digest
16/09/2010Transformation of ligations: 5µL ligation assay in 50µL TOP10 E.coli 25 min on ice 45sec heat shock on 42°C 1.5-2h shaking at 37°C plated 200µL on Ampicillin-LB/Agar-Plates after incubating ~8h, at 37°C, the plates were incubated overnight at room temperature
17/09/2010Colonies were visible in reasonable numbers on every plate Colony PCRs were performed to check for positive clones. Primer for colony PCR were stop-oligos, used in the raPCR. The PCR was performed in a total volume of 20 µL. One colony was dissolved in 20µL water. 5µL of this bacteria solution was used as PCR template. PCR conditions as recommended from Fermentas (see link above). From each plate, 10 colonies were picked (40 in total). Colony PCRs were then analysed on 1.5% agarose gel. Result: all negative Troubleshooting.... Minipreps were prepared (5mL - LB-ampicillin) for each sample for text digestion (over night, shaking @37°C)
18/09/2010Plasmid DNA was purified from over night cultures using Qiagen Plasmid Miniprep Kit according to the protocol. Elution was performed in 30 µL water. Concentrations ranged from approx. 400 to 788 ng/µL. Test digestion with NotI/XhoI was performed for 1h at @37°C and analysed on an 1.5% agarose gel. No insert was visible.
19/09/2010Troubleshooting: Possible Problems
Testing steps:
Over night digestion of backbone was performed at 37°C. 0.5 µg DNA was digested with 1 µL Enzyme in NEB Buffer 3 + BSA in a total volume of 30µL
20/09/2010Digestion of psiCHECK-2 plasmid was analysed on 1% agarose gel: Here we can see that both enzymes cut the plasmid. The linearized vector (visible at 6 kpb) in general shows up as a higher band than its undigested version, which is here visible at 4 kbp. Where the undigested plasmid shows concatemers, those are not visible after digestion, which proofs again for succesful digestion. The new digested plasmid-backbone was used for repeat of the ligation.
21/09/2010Ligation was performed for 4h @25°C (1µL NotI/XhoI-linearised psiCHECK2-plasmid + 4µL digested raPCR product) using Fermentas T4 Ligase
28/09/2010
28/09/2010
29/09/2010
(all those were made with Spacer(0) )
30/09/2010
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"