Team:Heidelberg/Project/miMeasure
From 2010.igem.org
(→Analysis of endogeneous miRNA) |
Laura Nadine (Talk | contribs) |
||
| Line 25: | Line 25: | ||
=====Analysis of mutated/ randomized binding sites against synthetic miRNA===== | =====Analysis of mutated/ randomized binding sites against synthetic miRNA===== | ||
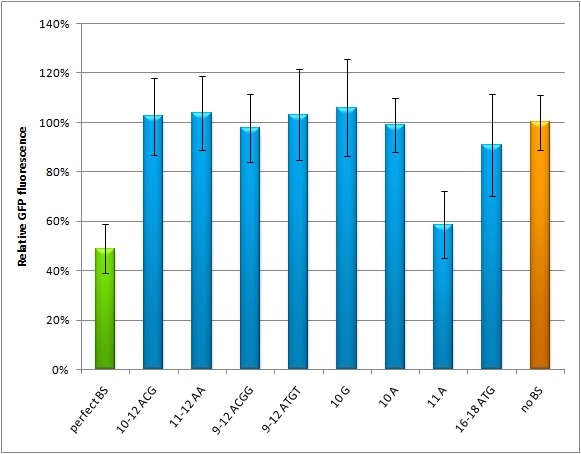
| - | We used microscopy analysis to determine the EGFP expression in relation to EBFP2. EBFP2 serves as a normalization for transfection efficiency. Nine miMeasure constructs with different binding sites were designed. The binding sites are either mutated at one site, or they contain randomly changed sites within a certain range. The construct representing the 100% knock-down is the perfect binding site, which is complementary to the synthetic miRNA miRsAg. The negative control represents 0% knock-down, since there is no binding site cloned into this miMeasure construct. | + | We used microscopy analysis to determine the EGFP expression in relation to EBFP2. EBFP2 serves as a normalization for transfection efficiency. Nine miMeasure constructs with different binding sites were designed. The binding sites are either mutated at one site, or they contain randomly changed sites within a certain range. The construct representing the 100% knock-down is the perfect binding site, which is complementary to the synthetic miRNA miRsAg. The negative control represents 0% knock-down, since there is no binding site cloned into this miMeasure construct. M12 contains the perfect binding site. The GFP/BFP-ratio stand for the level of GFP-expression normalized to one copy per cell. When we compare the GFP/BFP-ratio between the constructs, there is a significant difference of GFP expression in the control (miMeasure without binding site) and the construct containing the perfect binding site for the cotransfected synthetic miRNA. The modified binding sites don't supress GFP-expression as much as the perfect one. So GFP expression is just in between, except for constructs M20 and M22. |
Only the nucleotide 9 is mutated in M20. The change doesn't disturb the knock-down efficiency. Thus down-regulation of GFP is of similar extent as M12, which contains the perfect binding site. The seed region is altered in M22. Since the seed region is considered the most important site for knock-down efficiency, its change diminishes the knock-down capability of the binding site completely. So the GFP expression in this case is as high as the negative control, where no binding site was inserted into the miMeasure plasmid. | Only the nucleotide 9 is mutated in M20. The change doesn't disturb the knock-down efficiency. Thus down-regulation of GFP is of similar extent as M12, which contains the perfect binding site. The seed region is altered in M22. Since the seed region is considered the most important site for knock-down efficiency, its change diminishes the knock-down capability of the binding site completely. So the GFP expression in this case is as high as the negative control, where no binding site was inserted into the miMeasure plasmid. | ||
| - | [[Image:M12-M22_HeLa_daten.jpg|thumb| | + | [[Image:M12-M22_HeLa_daten.jpg|thumb|500px|center|'''GFP/BFP ratio normalized by the negative control''' The data are generated by confocal microscopy of Hela cells, which were transfected with different miMeasure constructs M12-M22 including the negative control (miMeasure construct without binding site). The negative control equals to 1.]] |
| - | =====Analysis of miraPCR generated binding sites against natural miRNA | + | ===Analysis of binding sites against synthetic miRNA miRsAg=== |
| + | |||
| + | 5000 HeLa cells were seeded on day one in each well of the 96 well plate. Transfection of the constructs (M12-M22) with four different conditions were carried out on day two. The ratio of transfection is 1 (M construct) : 5 (stuffer/ miRsAg/ pcDNA5/ shRNA3) with a total amount of 50ng DNA. | ||
| + | |||
| + | Condition '''a''': cotransfection with stuffer (salmon sperm DNA) | ||
| + | |||
| + | Condition '''b''': cotransfection with synthetic RNA miRsAg | ||
| + | |||
| + | Condition '''c''': cotransfection with empty pcDNA5 | ||
| + | |||
| + | Condition '''d''': cotransfection with synthetic shRNA3 | ||
| + | |||
| + | A control consisting of the empty miMeasure plasmid (without binding site) was also cotransfected with the same conditions a, b, c and d. The cells were used for measurements on day three. | ||
| + | |||
| + | {| class="wikitable sortable" border="0" style="text-align: left" | ||
| + | |-bgcolor=#cccccc | ||
| + | |+ align="top, left"|'''used binding sites and their features''' | ||
| + | |sequence||binding site feature||Name/number | ||
| + | |- | ||
| + | |ctcagtttactagtgccatttgttc||perfect binding site against miRsAg||M12 | ||
| + | |- | ||
| + | |ctcagtttactagacgcatttgttc||miMeasure with randomised nucleotides 10-12||M13 | ||
| + | |- | ||
| + | |ctcagtttactagtaacatttgttc||miMeasure with randomised nucleotides 10-12||M14 | ||
| + | |- | ||
| + | |ctcagtttactagacggatttgttc||miMeasure with randomised nucleotides 9-12||M15 | ||
| + | |- | ||
| + | |ctcagtttactagatgtatttgttc||miMeasure with randomised nucleotides 9-12||M16 | ||
| + | |- | ||
| + | |ctcagtttactagtggcatttgttc||miMeasure with mutated nucleotide 10||M17 | ||
| + | |- | ||
| + | |ctcagtttactagtgacatttgttc||miMeasure with mutated nucleotide 10||M18 | ||
| + | |- | ||
| + | |ctcagtttactagtaccatttgttc||miMeasure with mutated nucleotide 9||M20 | ||
| + | |- | ||
| + | |ctcagttatgtagtgccatttgttc||miMeasure with mutated nucleotide 9||M22 | ||
| + | |- | ||
| + | |-||miMeasure without any binding site||NC (negative control) | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | |||
| + | ===Analysis of miraPCR generated binding sites against natural miRNA=== | ||
The miraPCR generates binding sites, which are aligned by chance with each other, whereby different random spacer regions are inserted in between. If perfect binding sites are aligned, this is predicted to give a stronger knock-down than a single one. If imperfect binding sites are aligned, it is also supposed to be stronger than a single one. In this way binding sites with different knock-down efficiencies can be generated. | The miraPCR generates binding sites, which are aligned by chance with each other, whereby different random spacer regions are inserted in between. If perfect binding sites are aligned, this is predicted to give a stronger knock-down than a single one. If imperfect binding sites are aligned, it is also supposed to be stronger than a single one. In this way binding sites with different knock-down efficiencies can be generated. | ||
| - | This experiment shows, that the binding site pattern with 3 aligned perfect binding sites | + | This experiment shows, that the binding site pattern with 3 aligned perfect binding sites miM-1.3-7 gives the strongest knock-down. Whereby the binding site pattern with two imperfect binding sites 1-8 is weaker, but still stronger than the negative control. If one binding site is randomized from nucleotide 9-12 or 9-22 miM-r12, miM-r22, it loses its capability of protein down-regulation. |
| + | |||
| + | HeLa and HuH7 cells were transfected with the constructs miM-r12, miM-r22, miM-1.3-7 and miM-3.1-8 (see table2). These constructs contain binding sites against miR-122. Since HuH7 cells express miR-122, the constructs will be affected in the HuH7 cells without cotransfecting any miRNAs, whereas miR-122 were cotransfected for the HeLa cells. | ||
| + | The transfection is done with lipofectamin. 70ng of total DNA is used and for HeLa cells the miRNA to miMeasure construct ratio was 2:1. The cells were imaged one day after trasnfection with the Leica confocal microspe. The result shows the ratio between GFP and BFP normalized to the negative control, where miMeasure constructs without binding sites are transfected. The miMeasure construct with one perfect binding site is also transfected. | ||
| - | + | <center> | |
| - | + | {| class="wikitable sortable" border="0" style="text-align: left" | |
| + | |-bgcolor=#cccccc | ||
| + | |+ align="top, left"|'''raPCR designed binding sites and their features | ||
| + | |binding site feature'''||Name/number | ||
| + | |- | ||
| + | |miMeasure with 3 aligned perfect binding sites||miM-1.3-7 | ||
| + | |- | ||
| + | |miMeasure with two imperfect binding sites||miM-3.1-8 | ||
| + | |- | ||
| + | |miMeasure with randomised nucleotides 9-12||miM-r12 | ||
| + | |- | ||
| + | |miMeasure with randomised nucleotides 9-22||miM-r22 | ||
| + | |- | ||
| + | |} | ||
| + | </center> | ||
==Discussion== | ==Discussion== | ||
Revision as of 07:36, 27 October 2010

|
|
||||||||||||||||||||||||||||||||||||||||||
 "
"