Team:Heidelberg/Project/miRNA Kit
From 2010.igem.org
(→raw data) |
(→raw data) |
||
| Line 38: | Line 38: | ||
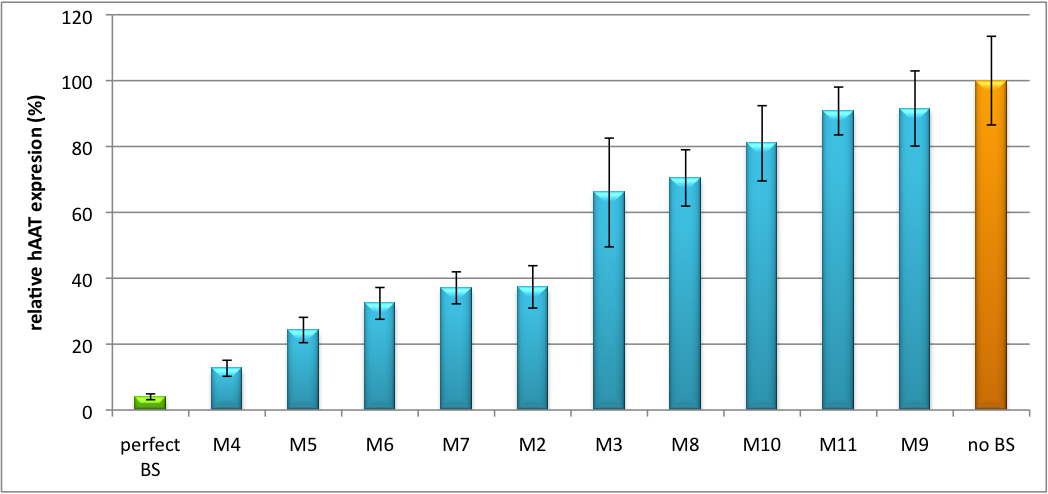
[[Image:Tuning U6 1.png|thumb|center|600px|'''Figure 2: Tuning of gene expression through different imperfect shRNA miR binding sites in pBS_U6.''' Gene expression quantified via dual luciferase assay for M constructs containing different imperfect binding sites.]] | [[Image:Tuning U6 1.png|thumb|center|600px|'''Figure 2: Tuning of gene expression through different imperfect shRNA miR binding sites in pBS_U6.''' Gene expression quantified via dual luciferase assay for M constructs containing different imperfect binding sites.]] | ||
Strikingly, the order of constructs in terms of knockdown for the imperfect binding sites is similar. M4, M5 and M6 always show strong knockdown, whereas M9, M10 and M11 show only loose down-regulation. Consulting the binding site sequences, the weak knockdown can be addressed to bulges in the supplementary region or complete lack of the 3' region of the binding site. Still high strength could be maintained due to only single nucleotide exchanges in the central region of the binding site. | Strikingly, the order of constructs in terms of knockdown for the imperfect binding sites is similar. M4, M5 and M6 always show strong knockdown, whereas M9, M10 and M11 show only loose down-regulation. Consulting the binding site sequences, the weak knockdown can be addressed to bulges in the supplementary region or complete lack of the 3' region of the binding site. Still high strength could be maintained due to only single nucleotide exchanges in the central region of the binding site. | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<!--After creating a binding site library and testing the miRNA-binding site interaction <i>in vitro</i>, we were able to compute an [https://2010.igem.org/Team:Heidelberg/Modeling/miGUI <i>in silico</i> model] based on a machine learning approach to predict knockdown efficiencies. A more detailed description of the different binding sites, we characterized can be found in our [https://2010.igem.org/Team:Heidelberg/Project/miMeasure measurements] page. | <!--After creating a binding site library and testing the miRNA-binding site interaction <i>in vitro</i>, we were able to compute an [https://2010.igem.org/Team:Heidelberg/Modeling/miGUI <i>in silico</i> model] based on a machine learning approach to predict knockdown efficiencies. A more detailed description of the different binding sites, we characterized can be found in our [https://2010.igem.org/Team:Heidelberg/Project/miMeasure measurements] page. | ||
Revision as of 06:05, 27 October 2010

|
|
||
 "
"