Team:Harvard/flavor/notebook
From 2010.igem.org
(→06-22-2010 [ top ]) |
(→06-23-2010 [ top ]) |
||
| Line 143: | Line 143: | ||
| - | |||
Revision as of 04:57, 25 October 2010

notebook calendar [ top ]
| Week 1 | 06-14-2010 | 06-15-2010 | 06-16-2010 | 06-17-2010 | 06-18-2010 |
| Week 2 | 06-21-2010 | 06-22-2010 | 06-23-2010 | 06-24-2010 | 06-25-2010 |
| Week 3 | 06-28-2010 | 06-29-2010 | 06-30-2010 | 07-01-2010 | 07-02-2010 |
| Week 4 | 07-05-2010 | 07-06-2010 | 07-07-2010 | 07-08-2010 | 07-09-2010 |
| Week 5 | 07-12-2010 | 07-13-2010 | 07-14-2010 | 07-15-2010 | 07-16-2010 |
| Week 6 | 07-19-2010 | 07-20-2010 | 07-21-2010 | 07-22-2010 | 07-23-2010 |
| Week 7 | 07-26-2010 | 07-27-2010 | 07-28-2010 | 07-29-2010 | 07-30-2010 |
| Week 8 | 08-02-2010 | 08-03-2010 | 08-04-2010 | 08-05-2010 | 08-06-2010 |
| Week 9 | 08-09-2010 | 08-10-2010 | 08-11-2010 | 08-12-2010 | 08-13-2010 |
06-14-2010 [ top ]
- Miraculin and brazzein constructs due to arrive on Wednesday from Mr. Gene
BioBrick Transformation
BioBrick parts from the 2010 iGEM kit were transformed and grown in highly competent TURBO bacteria.
- Wintergreen Scent Pathway:
BBa_J45700 - entire pathway, Ampicillin
BBa_J45004 - BSMT1 only, Ampicillin
(Not in 2010 BB Kit: BBa_J45017 - PchB, PchA)
- Banana Scent Pathway:
BBa_J45250 - ATF3 + Promoter, Ampicillin?
BBa_J45014 - ATF3 only, Ampicillin
(Not in BB 2010 Kit: BBa_J45400 - BAT2 and THI3)
06-15-2010 [ top ]
Primer Designs for Agrobacterium Vector
- Primers were designed to amplify from the Expression Series pORE Agrobacterium plasmid (e3) 1)the pENTcup2 promoter, 2) the tNOS stop sequence, 3) the tNOS stop sequence + additional stop codon and 4) the pHLP promoter.
Note: the NOSterm_BB_R primer works for both tNOS and tNOS+STOP codon sequences.
pENTcup2_BB_F CCTTTCTAGAGGGATCTTCTGCAAGCATCT
pENTcup2_BB_R AAGGCTGCAGCGGCCGCTACTAGTTCCGGTGGGTTTTGAGGT
STOP_NOSterm_BB_F CCTTTCTAGATGAGATCGTTCAAACATTTGG
NOSterm_BB_F CCTTTCTAGAGATCGTTCAAACATTTGGCA
NOSterm_BB_R AAGGCTGCAGCGGCCGCTACTAGTGATCTAGTAACATAGATGACA
pHPL_BB_F CCTTTCTAGAAACGTGGATACTTGGCAGTG
pHPL_BB_R AAGGCTGCAGCGGCCGCTACTAGTCTTTTGAGCTTAGAGGTTTTT
BioBrick Transformation
- Colonies were observed after overnight culture growth, but in low quantities.
- J45700 and J45004 (both Wintergreen Pathway) showed minimal number of colonies on both 10μL and 100μL cultures.
- J45250 and J45014 (both Banana Pathway) showed no colonies on either the 10μL or 100μL cultures.
Codon Usage in Arabidopsis
Using http://gcua.schoedl.de/
Valencene Extraction
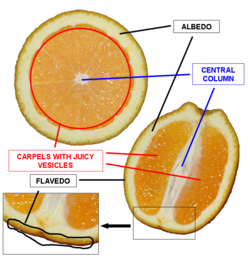
Used RNeasy Plant Mini Kit to extract RNA from the flavedo of an Organic Valencia Orange.
06-16-2010 [ top ]
MiniPrep
- MiniPrep of Wintergreen parts from Registry (J45004 and J45700) following Qiagen MiniPrep protocol. MiniPrep three samples of each part.
- DNA concentrations:
J45004-1: 59.1 ng/μL
J45004-2: 72.9 ng/μL
J45004-3: 88.1 ng/μL
J45700-1: 146.7 ng/μL
J45700-2: 187.4 ng/μL
J45700-3: 175.5 ng/μL
Digestion with Enzymes
- For J45004, we added: 9μL DNA, 1μL buffer, 1μL xbaI restriction enzyme (slow), and .5μL pstI restriction enzyme (fast).
- For J45700, we added 5μL DNA, 4μL H2O, 1μL buffer, 1μL xbaI restriction enzyme (slow), and .5μL pstI restriction enzyme (fast).
- We let mixtures sit for 30 minutes due to the use of xbaI restriction enzyme (slow). After the 30 minutes, we added 2.5μL dye to each mix.
Gel
- We loaded 12.5μL of 1kb ladder to well 1 of the gel (numbered left to right). We then loaded 12.5μL of J45004-1, J45004-2, and J45004-3 to wells 2,3, and 4, respectively. We loaded J45700-1, J45700-2, J4500-3 in wells 5, 6, and 7, respectively (see images below for well locations).
- Ran on 1% agarose gel.
06-17-2010 [ top ]
PCR Purification of pORE Vector Parts
Following QIAgen PCR Purification Kit Protocol the following PCR products were purified:
- pENTCUP2 Promoter - 16.1 ng/μL
- NOSterminator Sequence - 76.3 ng/μL
- STOP codon + NOSterminator Sequence - 76.5 ng/μL
To do: follow up with restriction digest ✓ and Agarose gel ✓ to confirm PCR and PCR purification.
Restriction Digest and Agarose Gel
Restriction Digest reactions were set up as follows:
pENTCUP2 Promoter:
- 27μL PCR product
- 3μL Loading Buffer w/ dye
- 1μL Xba1 Fast Enzyme
- 1μL Pst1 Fast Enzyme
NOSterm and NOSterm + STOP:
- 9μL PCR product
- 1μL Loading Buffer w/ dye
- 0.5μL Xba1 Fast Enzyme
- 0.5μL Pst1 Fast Enzyme
BioBrick Plasmid V0120 (Ampicillin Resistance):
- 3μL DNA
- 1μL Loading Buffer w/ dye
- 6μL diH2O
- 0.5μL Xba1 Fast Enzyme
- 0.5μL Pst1 Fast Enzyme
Reactions were allowed to proceed for 45 minutes at 37°C.
06-24-2010 [ top ]
06-25-2010 [ top ]
We began this week trying to ligate the pENTCUP2 promoter, NosT terminator, and NosT terminator plus stop codon into the V0120 vector. We ran into several problems during this process.
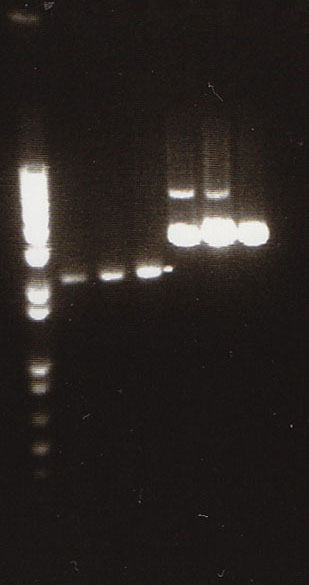
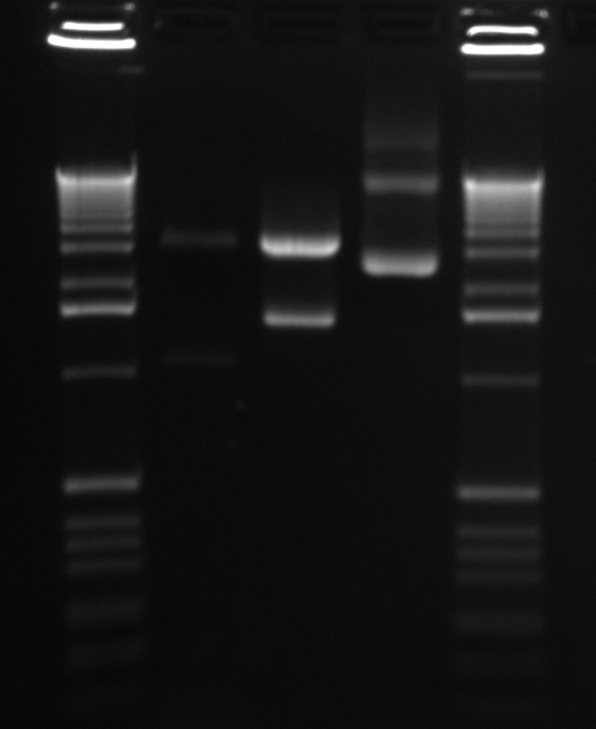
We had previously digested the three inserts as well as the promoter with XbaI and PstI fast digest enzymes, ligated, and transformed the ligated vectors into e. coli. Unfortunately, we did not find any colonies of cells after over 14 hours of incubation. Furthermore, on Friday June 18th, after digesting all three inserts and vector and running them on a gel, we did not see any NosT + stop DNA. Therefore, on Monday June 21, we began by digesting more NosT + stop and V0120 vector with xba1 and pstI fast digestive enzymes. The gel showed DNA length to be consistent with the given parts. (See gel below)
- Gel Lanes (both inserts and vector were digested with xbaI/pstI): 1. 1kb plus ladder. 2. pENTCUP2 promoter. 3. NosT terminator. 4. NosT + stop terminator 5. V0120 digested.
- Gel Lanes (both inserts and vector were digested with xbaI/pstI): 1. 1kb plus ladder. 2. NosT + stop terminator. 3. V0120 digested.
Upon gel extraction and gel purification, we ligated the inserts into the vectors with a 2:1 insert to vector ratio (previously we had been using a 4:1 insert to vector ratio). We then transformed the plasmids into e.coli and plated; no colony growth was seen the next morning.
We used Team Vector's R3 ligation (they have successfully ligated and transformed) as a positive control. However, the bacteria containing the ligated plasmids were plated on Ampicillin plates, which turned out to be the wrong resistance.
Troubleshooting
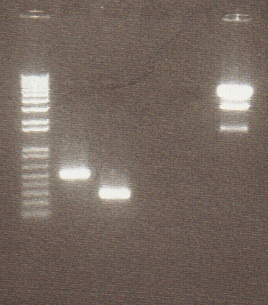
In order to get a better understanding of the problem with our ligation, we ran a diagnostic gel with the V0120 backbone and vectors B15 and B21.
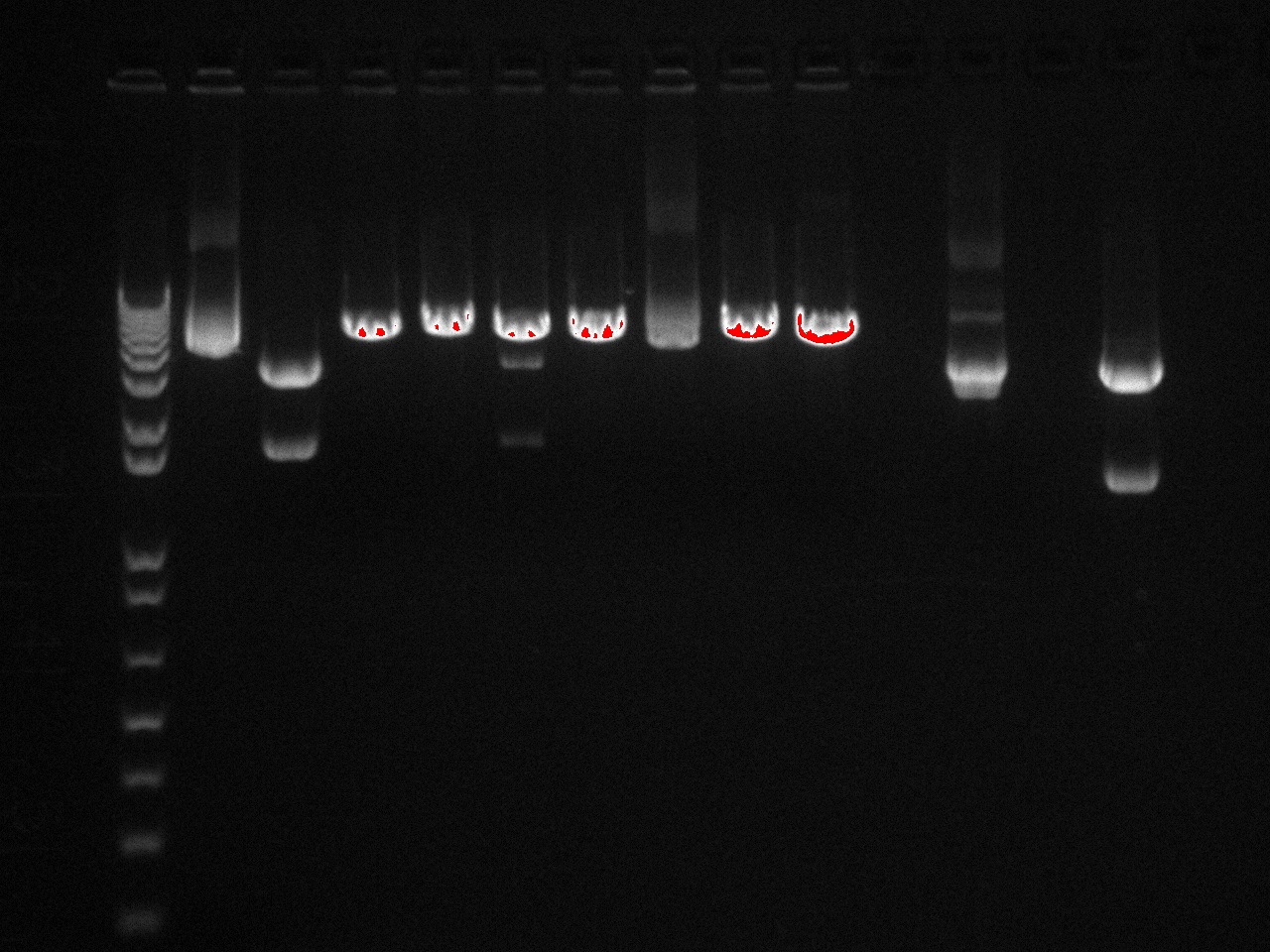
Gel Lanes: 1. 1 kb plus ladder 2. v0120 undigested 3. v0120 eco/spe 4. v0120 eco/xba 5. v0120 spe/pstI 6. v0120 xba/pst 7. v0120 ecoRI 8. v0120 xba1 9. v0120 pstI 10. v0120 spe 11. blank 12. B15 (StrepII tag) xba/pstI 13. B21 (YFP) xba/pstI
We concluded from the gel that xba1 is the problem in our ligation step. The pattern of bands in the v0120 xba1/pstI digest along with the v0120 xba1 digest illustrate that xba1 is not cutting correctly.
We began a slow enzyme xba1/pstI digest on Friday, but the gel was run in mismatching buffers (TAE and TBE).
- Gel Lanes (inserts and vector digested with xba1/pstI): 1. 1 kb plus ladder. 2. pENTCUP2 3. NosT 4. NosT & stop 5. v0120
Valencene
Digestion of PCR products aimed to isolate valencene gene showed very short length on gel. These are not the correct size and are most likely primer dimers. PCR products were digested with EcoRI/SpeI. 5 μL of each sample was digested with pstI to be sure that the pstI site was in the sequence of DNA.
- Gel Lanes: 1. 1 kb plus ladder. 2. Flavedo EcoRI/SpeI. 3. Flavedo PstI 4. Open 5. Fruit EcoRI/SpeI. 6. Fruit pstI
06-28-2010 [ top ]
Miniprep of Miraculin and Brazzein Constructs
- Protocol used was from the Qiagen Miniprep Kit
| Name | Concentration (ng/μL) |
|---|---|
| Miraculin 1 | 159.6 |
| Miraculin 2 | 182.2 |
| Brazzein 1 | 91.2 |
| Brazzein 2 | 33.7 |
DTL (Digestion, Ligation, Transformation) of The Big Three (pENTCUP2, NOSt, NOSt + STOP)
| pENTCUP2 | NOSt | NOSt+STOP | B15 | |
|---|---|---|---|---|
| DNA | 4 | 7 | 12 | 7 |
| NEB Buffer 3 (10x) | 2 | 2 | 2 | 2 |
| diH2O | 10 | 7 | 2 | 7 |
| Xba1 | 1 | 1 | 1 | 1 |
| Pst1 | 1 | 1 | 1 | 1 |
| BSA (10x) | 2 | 2 | 2 | 2 |
- Digestions were left for 1:30 at 37°C
- 2.2 μL of DNA Loading Buffer were added to each reaction and loaded onto a 1% Agarose gel (TAE buffer). Gel was ran at 125 V for 30 min.
| Lane | Contents |
|---|---|
| Lane 1 | 1KB Plus Ladder |
| Lane 2 | pENTCUP2 |
| Lane 3 | NOSt |
| Lane 4 | NOSt+STOP |
| Lane 5 | B15 digested |
| Lane 6 | B15 undigested |
- The undigested B15 in lane 6 appears as two bands on account of supercoiling of the B15 plasmid.
- Bands in lanes 2 - 5 were cut out and purified using a Qiagen Gel Purification kit
- The gel purification was done with 300 μL Buffer PB and 400 μL Buffer QG
| Name | Concentration (ng/μL) |
|---|---|
| pENTCUP2 | 16.5 |
| NOSt | 10.2 |
| NOSt+STOP | 0.4 |
| B15 (backbone vector) | 10.9 |
- Note the very low concentration of NOSt+STOP. It is uncertain as to what caused such a concentration, but transformation proceeded anyways with max volume
- The ligation reactions were preformed at at 2:1 (insert to backbone) Molar Ratio
- The sole exception being the NOSt+STOP ligation, in which the lack of product forced us to use 1.8 ng DNA
| pENTCUP2 | NOSt | NOSt+STOP | Control | |
|---|---|---|---|---|
| Insert | 1 | 1 | 1 | 1 |
| Vector | 2 | 2 | 2 | 2 |
| Buffer (10x) | 5 | 5 | 5 | 5 |
| Ligase | 1 | 1 | 1 | 1 |
| H2O | 11 | 11 | 0 | 12 |
- Ligation reactions were preformed as per the Silver Lab ligation protocol (with the differences in concentration and ratio made as per the table above).
- Transformations to E. Coli were preformed as per the Silver lab transformation protocol with ampicillin plates.
- Plates were left overnight.
06-29-2010 [ top ]
Results of ligation and transformation of The Big Three (pENTCUP2, NosT, NosT + stop) with B15 backbone into E.coli
A few colonies were found on the pENTCUP2 + Backbone plate, NosT + stop and backbone plate, and b15 backbone control plate. Colonies were picked from the plates and placed in 3ml of LB and agarose in a culture tube. The tubes were placed in the 37°C shaker for 8 hours. 2 mL of LB+AMP was added to the cultures, bring the total volume to 5 mL of LB+AMP. Cultures were left to shake overnight.
DLT of the Sweethearts - Miraculin and Brazzein (and B21)
| Miraculin | Brazzein | B21 | |
|---|---|---|---|
| DNA | 6 | 11 | 7 |
| FD Buffer (10x) | 2 | 2 | 2 |
| diH2O | 10 | 5 | 9 |
| EcoRI | 1 | 1 | 1 |
| SpeI | 1 | 1 | 1 |
Placed in 37°C waterbath for 20 minutes.
Digestion Gel
Gel Lanes:
1. 1 kb plus ladder
2. Miraculin digested with EcoRI/SpeI
3. Brazzein digested with EcoRI/SpeI
4. B21 digested with EcoRI/SpeI
5. 1 kb plus ladder
| Name | Concentration (ng/μL) |
|---|---|
| Miraculin | 9.5 |
| Brazzein | 10.2 |
| B21 Vector (V0120) | 8.4 |
| Miraculin | Brazzein | Control | |
|---|---|---|---|
| Insert | 3.5 | 0.75 | 0 |
| Vector | 6 | 6 | 6 |
| Buffer (10x) | 2 | 2 | 2 |
| Ligase | 1 | 1 | 1 |
| H2O | 6.5 | 10.25 | 0 |
- The ligation of Miraculin was done at a 3:1 ratio (insert to vector) due to its larger size (~700 bp)
- The ligation of Brazzein was done at a 2:1 ratio due to its smaller size (~200 bp)
06-30-2010 [ top ]
Miraculin, Brazzein
- Colonies did grow!
- Started 5 mL cultures of Miraculin and Brazzein biobrick constructs (5 cultures each)
The Big Three
- Began miniprep of cultures of pENT, NOSt+STOP and our Control (for experimental purposes)
| OD | |
|---|---|
| pENTCUP2 #1 | 248.5 ng/μL |
| pENTCUP2 #2 | 447.7 ng/μL |
| NosT + stop #1 | 427.7 ng/μL |
| NosT + stop #2 | 303.9 ng/μL |
| Control #1 | 366.8 ng/μL |
| Control #2 | 359.2 ng/μL |
Digestion
- We digested miniprep samples with xbaI/pstI fast digest enzymes in order to determine if ligation was successful.
| Control #1 | Control #2 | pENTCUP2 #1 | pENTCUP2 #2 | NosT & stop #1 | NosT & stop #2 | |
|---|---|---|---|---|---|---|
| DNA | 2 | 2 | 3 | 2 | 2 | 3 |
| Green FD buffer | 2 | 2 | 2 | 2 | 2 | 2 |
| diH2O | 14 | 14 | 13 | 14 | 14 | 13 |
| xbaI | 1 | 1 | 1 | 1 | 1 | 1 |
| pstI | 1 | 1 | 1 | 1 | 1 | 1 |
Digestion mixes were placed in 37°C water bath for approximately 1 hr.
Big Three PCR digestion
- We simultaneously digested PCR purified pENTCUP2, NosT, and NosT & stop inserts with xbaI/pstI. This way if the above digestion showed the ligation of the inserts into v0120 backbone was not successful, we can continue with these inserts.
| pENTCUP2 | NosT | NosT & stop | |
|---|---|---|---|
| DNA | 4 | 7 | 2 |
| Green FD buffer | 2 | 2 | 2 |
| diH2O | 12 | 9 | 14 |
| xbaI | 1 | 1 | 1 |
| pstI | 1 | 1 | 1 |
Placed in 37°C water bath for 20 minutes.
07-01-2010 [ top ]
- We made glycerol stocks from our Miraculin & v0120 and Brazzein & v0120. 333 μL cultures with 666μL glycerol and placed in -80°C freezer.
- We miniprepped our Miraculin and v0120 cultures and our Brazzein and v0120 cultures. Miniprep done per Qiagen protocol.
ODs
| Sample | OD |
|---|---|
| Miraculin and v0120 #1 | 171.3ng/μL |
| Miraculin and v0120 #2 | 202.0ng/μL |
| Miraculin and v0120 #3 | 106.4ng/μL |
| Miraculin and v0120 #4 | 115.6ng/μL |
| Miraculin and v0120 #5 | 40.5ng/μL |
| Brazzein and v0120 #1 | 41.7ng/μL |
| Brazzein and v0120 #2 | 42.8ng/μL |
| Brazzein and v0120 #3 | 38.8ng/μL |
| Brazzein and v0120 #4 | 0ng/μL |
| Brazzein and v0120 #5 | N/A |
Valencia orange surgery (part two)
- RNA extraction per Qiagen RNAeasy protocol.
- We made sure to apply RNAzap (RNAase-free) spray to all surfaces and instruments.
ODs
| Sample | OD |
|---|---|
| RNA Flavedo extraction #1 | 93.1ng/μL |
| RNA Flavedo extraction #2 | 28.7ng/μL |
07-02-2010 [ top ]
NosT + stop and v0120
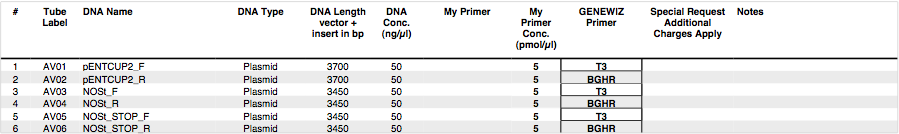
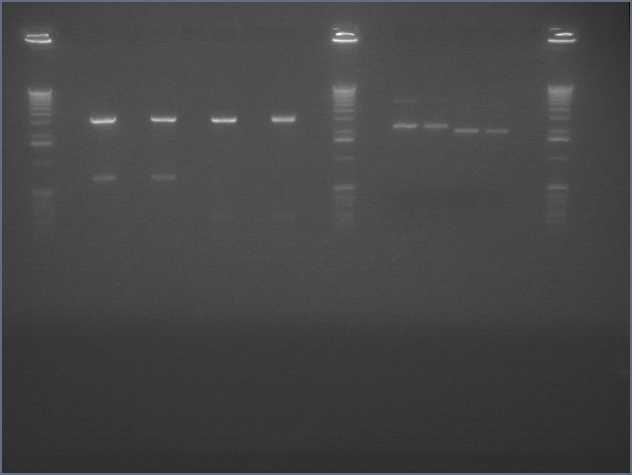
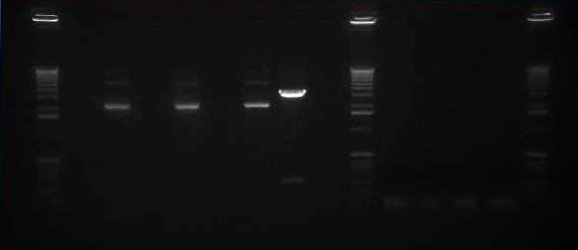
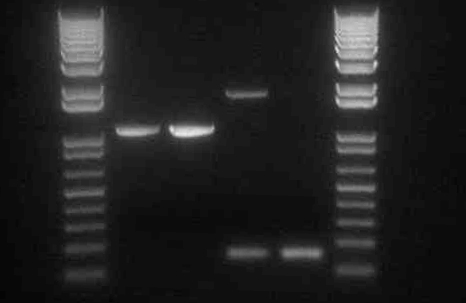
We ran an xba/pstI fast enzyme digest on our miniprepped NosT + stop and v0120 plasmid. The gel was consistent with a successful ligation. (See lanes 7-10 in gel image below).
Valencene
After running a reverse transcriptase reaction on our RNA extraction, we ran PCR on the cDNA with Valencene specific primers. The gel did not show the proper fragment length of the Valencene gene. Instead, it appears primer dimers formed again. (See lanes 2-5)
07-05-2010 [ top ]
07-06-2010 [ top ]
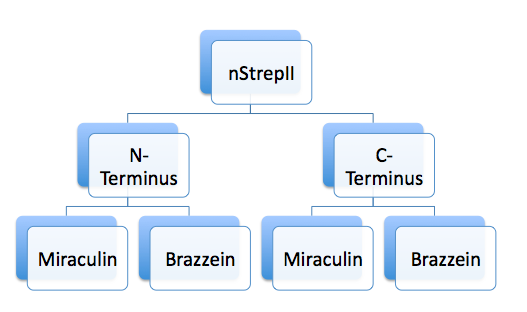
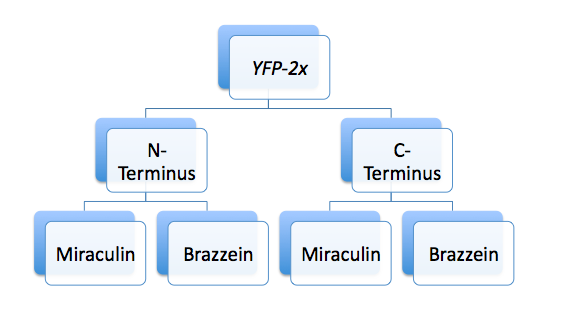
Tagging Miraculin and Brazzein with YFP and StrepII tags
Tag: -E--N--X--STREP/YFP--S--N--P-
Insert: E--N--X--Miraculin/Brazzein--S--N--P-
- The 'tag' biobrick (either YFP or STREPII) was used as the vector; Miraculin and Brazzein were used as the insert.
- Tags were ligated to both the N- and C-terminal of Miraculin and Brazzein
| Vector | Insert | |
|---|---|---|
| N-Terminus | Spe1/Pst1 | Xba1/Pst1 |
| C-Terminus | EcoR1/Xba1 | EcoR1/Spe1 |
| B21 S/P | B21 E/X | Miraculin X/P | Miraculin E/S | Brazzein X/P | Brazzein E/S | |
|---|---|---|---|---|---|---|
| diH2O | 13 | 13 | 11 | 11 | 0 | 0 |
| Green FD buffer (10x) | 2 | 2 | 2 | 2 | 2 | 2 |
| DNA | 3 | 3 | 5 | 5 | 16 | 16 |
| Spe1 | 1 | - | - | 1 | - | 1 |
| PstI | 1 | - | 1 | - | 1 | - |
| EcoR1 | - | 1 | - | 1 | - | 1 |
| Xba1 | - | 1 | 1 | - | 1 | - |
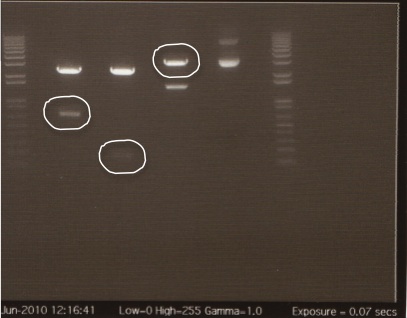
Gel
The gel showed DNA fragments consistent with Miraculin and Brazzein. The digestion of B21 appeared to be succesful, but the DNA sequence cut out was too small to see on the gel.
Gel Lanes:
1. 1 kb plus ladder
2. B21 speI/pstI
4. B21 ecoRI/xbaI
6. Miraculin xbaI/pstI
8. Miraculin ecoRI/speI
10. Brazzein xbaI/pstI
12. Brazzein ecoRI/pstI
14. 1kb plus ladder
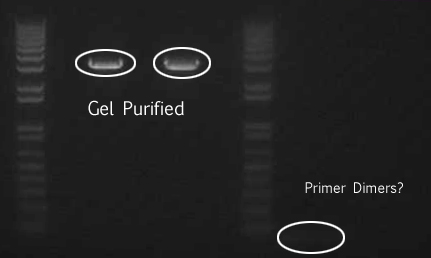
We extracted the circled bands and gel purified the gel per the Qiagen gel purification protocol.
ODs of gel purified DNA
B21 speI/pstI: 15.9 ng/μL
B21 ecoRI/xbaI: 11.0 ng/μL
Miraculin xbaI/pstI: 9.7 ng/μL
Miraculin ecoRI/speI: 7.1 ng/μL
Brazzein xbaI/pstI: 2.6 ng/μL
Brazzein ecoRI/pstI: 5.0 ng/μL
Ligation
We did 6 different ligation reactions. The chart below shows the different reactions. Ligation reactions were left at room temperature for 15 minutes.
| Miraculin xbaI/pstI w/ B21 speI/pstI | Miraculin ecoRI/speI w/ B21 ecoRI/xbaI | Brazzein xbaI/pstI w/ B21 speI/pstI | Brazzein ecoRI/speI w/ B21 ecoRI/xbaI | Control B21 speI/pstI | Control B21 ecoRI/xbaI | |
|---|---|---|---|---|---|---|
| DNA Insert | 2 | 3 | 3 | 1 | 0 | 0 |
| T4 DNA ligase buffer (10x) | 2 | 2 | 2 | 2 | 2 | 2 |
| diH2O | 12 | 9 | 11 | 11 | 14 | 12 |
| T4 DNA ligase | 1 | 1 | 1 | 1 | 1 | 1 |
| DNA Backbone | 1 | 1 | 1 | 1 | 1 | 1 |
Transformation
We mixed 5μL ligation mix with 15μL Turbo e. coli cells. These were placed in ice for 30 minutes and then heat shocked in 42°C water bath for 30 seconds. The cells were placed back on ice for 2 minutes. 170 μL of SOC broth was added to each Eppendorf tube. The transformed e. coli were then plated on LB Ampicillin plates and left in the 37°C incubator overnight.
07-07-2010 [ top ]
Sequencing
V0120 Plasmids containing the pENTCUP2 plant promoter, NOS terminator and NOS terminator + STOP were sent to GENEWIZ for sequencing. Sequencing results are expected tomorrow.
Cultures
5 mL cultures were started from the YFP-2x construct from yesterday, as well as the B15 (StrepII) tag.
- 2 x B21 E/X + Brazz E/S - C-Terminus
- 2 x B21 S/P + Brazz X/P - N-Terminus
- 2 x B21 E/X + Mira E/S - C-Terminus
- 2 x B21 S/P + Mira X/P - N-Terminus
- 2 x B15 Plate #1
- 2 x B15 Plate #2
Cultures were placed in a 37°C incubator and left to shake overnight.
Primers for Wintergreen pathway parts
J45004
- Left Primer: 5' cctttctagaatggaagttgttgaagttcttca 3'
- Right Primer: 5' aaggctgcagcggccgctactagtttaatttattttggtcaagga 3' (last 5 bp omitted to meet 45 bp maximum)
J45017
- Left Primer: 5' cctttctagaatgaaaactcccgaagactgc 3'
- Right Primer: 5' aaggctgcagcggccgctactagtttattaggcgacgccgc 3'
07-08-2010 [ top ]
- Re-submitted sequencing order, with correct primers, to GeneWiz
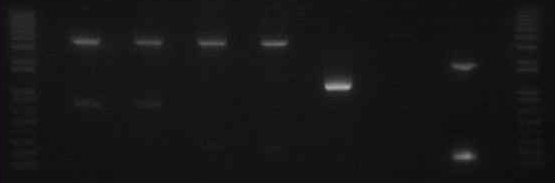
- Did a confirmation digest of the Miraculin/Brazzein + YFP-2x constructs
- Ladder
- B15 E/X
- B15 S/P
- Mira+YFP N1
- Mira+YFP N2
- Mira+YFP C1
- Mira+YFP C2
- Brazz+YFP N1
- Brazz+YFP N2
- Brazz+YFP C1
- Brazz+YFP C2
- Ladder
- It appears that the digestion of Mira+YFP C2 (lane 7) did not work properly. This could have been the result of choosing a bacterial colony that was the result of contamination, or from simply an error in the digestion step.
- Transformed two pDUET expression vectors for Miniprep
Ligated Miraculin/Brazzein to a StrepII tag
| Miraculin xbaI/pstI w/ B15 speI/pstI | Miraculin ecoRI/speI w/ B15 ecoRI/xbaI | Brazzein xbaI/pstI w/ B15 speI/pstI | Brazzein ecoRI/speI w/ B15 ecoRI/xbaI | Control B15 speI/pstI | Control B15 ecoRI/xbaI | |
|---|---|---|---|---|---|---|
| DNA Insert | 3 | 4 | 4 | 2 | 0 | 0 |
| T4 DNA ligase buffer (10x) | 2 | 2 | 2 | 2 | 2 | 2 |
| diH2O | 11 | 10 | 10 | 12 | 14 | 14 |
| T4 DNA ligase | 1 | 1 | 1 | 1 | 1 | 1 |
| DNA Backbone | 3 | 3 | 3 | 3 | 3 | 3 |
- We then transformed the ligated plasmids into Turbo e. coli and plated the bacteria on Amp plates. Plates were left in the 37°C incubator overnight.
07-09-2010 [ top ]
Ligation of Brazzein & Miraculin (w/ YFP tags) into DUET vector
Digestion
| Brazzein & YFP C EcoRI/SpeI | Brazzein & YFP N EcoRI/SpeI | Miraculin & YFP C EcoRI/SpeI | Miraculin & YFP N EcoRI/SpeI | V24 NotI/SpeI | NosT & Stop EcoRI/XbaI | |
|---|---|---|---|---|---|---|
| DNA | 2 | 2 | 2 | 3 | 5 | 2 |
| FD Buffer (10x) | 2 | 2 | 2 | 2 | 2 | 2 |
| diH2O | 14 | 14 | 14 | 13 | 11 | 14 |
| Enzyme | 1 | 1 | 1 | 1 | 1 | 1 |
| Enzyme | 1 | 1 | 1 | 1 | 1 | 1 |
- We are adding the NosT & stop to the C-terminus of the Miraculin and Brazzein constructs. This whole construct will then be ligated into the V24 DUET vector.
Gel
Gel Lanes:
1. 1kb plus ladder
3. Miraculin & YFP N-terminus ecoRI/SpeI
5. Miraculin & YFP C-terminus ecoRI/SpeI
7. Brazzein & YFP N-terminus ecoRI/SpeI
9. Brazzein & YFP C-terminus ecoRI/SpeI
11. V24 NotI/SpeI
13. NosT & Stop EcoRI/XbaI
07-12-2010 [ top ]
- We removed our cultures of e. coli containing Miraculin/Brazzein C/N YFP and NosT & Stop and e. coli containing Miraculin/Brazzein and StrepII tag from the 37°C shaker.
- Glycerol stocks were made (666 μL glycerol and 333 μL cells)
Miniprep
ODs
Miraculin C YFP and NosT & Stop 1: 414.4 ng/μL
Miraculin C YFP and NosT & Stop 2: 304.0 ng/μL
Miraculin N YFP and NosT & Stop 1: 428.6 ng/μL
Miraculin N YFP and NosT & Stop 2: 421.0 ng/μL
Brazzein C YFP and NosT & Stop 1: 572.8 ng/μL
Brazzein C YFP and NosT & Stop 2: 580.3 ng/μL
Brazzein N YFP and NosT & Stop 1: 348.9 ng/μL
Brazzein N YFP and NosT & Stop 2: 639.6 ng/μL
Miraculin N StrepII 1: 888.6 ng/μL
Miraculin N StrepII 2: 614.4 ng/μL
Miraculin C StrepII 1: 677.1 ng/μL
Miraculin C StrepII 2: 222.1 ng/μL
Brazzein N StrepII 1: 352.6 ng/μL
Brazzein N StrepII 2: 331.6 ng/μL
Brazzein C StrepII 1: 209.3 ng/μL
Brazzein C StrepII 2: 346.4 ng/μL
Digestion
| Miraculin C YFP NST | Miraculin N YFP NST | Brazzein N YFP NST | Brazzein C YFP NST | |
|---|---|---|---|---|
| diH2O | 14 | 14 | 13.5 | 14.3 |
| Green FD buffer (10x) | 2 | 2 | 2 | 2 |
| DNA | 2 | 2 | 2.5 | 1.7 |
| NotI | 1 | 1 | 1 | 1 |
| SpeI | 1 | 1 | 1 | 1 |
Gel
Gel Lanes:
1. 1 kb plus ladder
3. Miraculin N YFP NST (NotI/SpeI)
5. Miraculin C YFP NST (NotI/SpeI)
7. Brazzein N YFP NST (NotI/SpeI)
9. Brazzein C YFP NST (NotI/SpeI)
11. 1 kb plus ladder
- Gel purify per Qiagen protocol
ODs
Brazzein C YFP NST (NotI/SpeI): 6.3 ng/μL
Brazzein N YFP NST (NotI/SpeI): 8.5 ng/μL
Miraculin C YFP NST (NotI/SpeI): 3.6 ng/μL
Miraculin N YFP NST (NotI/SpeI): 12.8 ng/μL
07-13-2010 [ top ]
Ligation in V24
- Ligation of of Mira/Brazz+YFP+NOSt+STOP Construct into the V24 pETDUET vector for expression of proteins in IPTG inducible bacteria.
| Mira N + V24 | Mira C + V24 | Brazz N + V24 | Brazz C + V24 | Control | |
|---|---|---|---|---|---|
| diH2O | 7 | 1 | 6 | 4 | 12 |
| T4 DNA ligase buffer (10x) | 2 | 3 | 2 | 2 | 2 |
| DNA Insert | 5 | 20 | 6 | 8 | 0 |
| DNA Backbone | 5 | 5 | 5 | 5 | 5 |
| T4 DNA ligase | 1 | 1 | 1 | 1 | 1 |
- 5μL of Ligation reactio
07-14-2010 [ top ]
Miniprep of STOP + V0120 part
Nanodrop O/D's: STOP V0120 1-1: 134.4 ng/μL STOP V0120 1-2: 98.7 ng/μL STOP V0120 2-1: 55.2 ng/μL STOP V0120 2-2: 121.8 ng/μL
DTL of StrepII+Mira/Brazz with STOP codon
- To insert a STOP codon to the end of our Mira/Brazz StrepII constructs, we used the STOP+V0120 BioBrick as our vector (EcoR1/Xba1 digest) and our Miraculin/Brazzein construct as our insert (EcoR1/Spe1 digest)
- Construct DNA used was from Miniprep 1 (Box 4)
- ~1μg of DNA was digested in each reaction
| Mira Strep N | Mira Strep C | Brazz Strep N | Brazz Strep C | STOP+V0120 BioBrick | |
|---|---|---|---|---|---|
| diH2O | 15 | 14.5 | 13.5 | 11.5 | 6 |
| Green FD buffer (10x) | 2 | 2 | 2 | 2 | 2 |
| DNA | 1 | 1.5 | 2.5 | 4.5 | 10 |
| EcoR1 | 1 | 1 | 1 | 1 | 1 |
| Xba1 | - | - | - | - | 1 |
| Spe1 | 1 | 1 | 1 | 1 | - |
Digestion of Mira/Brazz+Strep and STOP codon BioBrick

- Circled bands were cut and gel purified
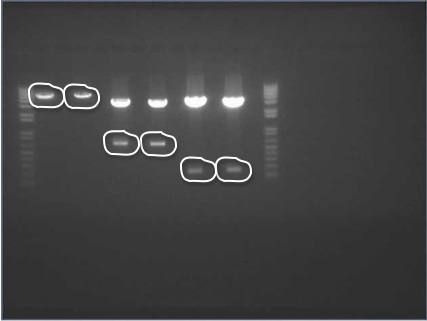
Miraculin + StrepII = 730 bp Brazzein + StrepII = 230 bp STOP + V0120 = 3200 bp
- 1kb Plus Ladder
- Miraculin C Strep E/S
- Miraculin N Strep E/S
- Brazzein C Strep E/S
- Brazzein N Strep E/S
- STOP + V0120 E/X
- 1kb Plus Ladder
Gel Purification Specs Mira N StrepII E/S: 3.0 ng/μL Mira C StrepII E/S: 2.8 ng/μL Brazz N StrepII E/S: 3.4 ng/μL Brazz C StrepII E/S: 1.3 ng/μL STOP+V0120 BB E/X: 7.1 ng/μL
Ligation of Mira/Brazz+Strep and STOP codon BioBrick
| Mira StrepII N + STOP w/ v0120 BB | Mira StrepII C + STOP w/ v0120 BB | Brazz StrepII N + STOP w/ v0120 BB | Brazz StrepII C + STOP w/ v0120 BB | Control STOP w/ v0120 BB | |
|---|---|---|---|---|---|
| diH2O | 4.5 | 4.5 | 9 | 5 | 12 |
| T4 DNA ligase buffer (10x) | 2 | 3 | 2 | 2 | 2 |
| DNA Insert | 7.5 | 7.5 | 2 | 6 | 0 |
| DNA Backbone | 5 | 5 | 6 | 6 | 5 |
| T4 DNA ligase | 1 | 1 | 1 | 1 | 1 |
Transformation
- Transformed 5 μL ligation mix with 15 μL TURBO e. coli cells
- Plated on LB + Amp plates and left in 37°C incubator overnight
07-15-2010 [ top ]
Miraculin/Brazzein YFP N/C and NosT + Stop in V24 DUET vector
- Made glycerol stocks (666 μL glycerol and 333 μL cells)
ODs
Miraculin C1: 168.8 ng/μL
Miraculin C2: 67.2 ng/μL
Miraculin N1: 183.0 ng/μL
Miraculin N2: 173.5 ng/μL
Brazzein N1: 54.8 ng/μL
Brazzein N2: 79.0 ng/μL
Brazzein C1: 161.4 ng/μL
Brazzein C2: 125.0 ng/μL
| Mira N1 | Mira N2 | Mira C1 | Mira C2 | Brazz N1 | Brazz N2 | Brazz C1 | Brazz C2 | |
|---|---|---|---|---|---|---|---|---|
| diH2O | 13.5 | 13.5 | 13.5 | 11 | 11 | 11 | 13.5 | 13 |
| Green FD buffer (10x) | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| DNA | 2.5 | 2.5 | 2.5 | 5 | 5 | 5 | 2.5 | 3 |
| PstI | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| Xba1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
Confirmation Digest of Mira/Brazz + YFP + NOSt + STOP in V24 Expression Vector
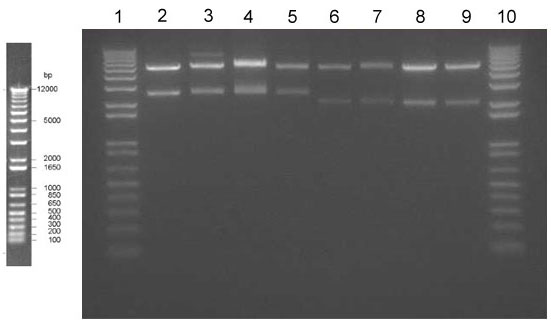
- Ladder
- Mira N1
- Mira N2
- Mira C1
- Mira C2
- Brazz N1
- Brazz N2
- Brazz C1
- Brazz C2
- Ladder
- The N & C designation corresponds to the terminus upon which the YFP was fused.
Expected Lengths
Miraculin + YFP + NOSt + STOP = 700 + 1500 + 250 = 2450 bp
Brazzein + YFP + NOSt + STOP = 300 + 1500 + 250 = 2050 bp
V24 = 5200 bp
07-16-2010 [ top ]
- Did minipreps of the Mira/Brazz + Strep constructs. All constructs had failed to ligate.
- Ran PCR of Valencene gene from extracted genomic DNA. Followed with PCR purification
- Re-digested STOP+V0120 BioBrick, gel purified. Gel Purification failed, no DNA was extracted.
07-19-2010 [ top ]
DTL of Mira/Brazz+StrepII into STOP+V0120
Digestion
| Stop v0120 1 | Stop v0120 2 | |
|---|---|---|
| DNA | 10 | 8 |
| FD Buffer (10x) | 2 | 2 |
| diH2O | 6 | 8 |
| Xba1 | 1 | 1 |
| EcoRI | 1 | 1 |
| Mira N + Stop | Mira C + Stop | Brazz N + Stop | Brazz C + Stop | Stop only Control | |
|---|---|---|---|---|---|
| diH2O | 3 | 3 | 11 | 6 | 14 |
| T4 DNA ligase buffer (10x) | 2 | 3 | 2 | 2 | 2 |
| DNA Insert | 11 | 11 | 3 | 8 | 0 |
| DNA Backbone | 3 | 3 | 3 | 3 | 3 |
| T4 DNA ligase | 1 | 1 | 1 | 1 | 1 |
Confirm of valencene PCR (failed)
- Re-do of Valencene PCR (failed)
07-20-2010 [ top ]
- Picked colonies from Mira/Brazz StrepII STOP constructs
- Only Mira N and Brazz C had colonies, 2 mL cultures were started and will be miniprepped to confirm ligation at the end of today.
- Diluted Mira/Brazz YFP expression colonies 25:1 in LB+AMP for eventual IPTG induction. Will culture @ 37°C for 1 hour then take O/D's.
07-21-2010 [ top ]
Miraculin N/Brazzein C StrepII & Stop
- We ran a confirmation digest on the miniprepped Miraculin N StrepII & Stop and the Brazzein C StrepII & Stop. We digested the samples with XbaI/PstI. We should have used NotI/SpeI in this digest if we planned on inserting into V24 pETDUET vector.
- We digested 558 ng of Mira N1, 516 ng Mira N2, 540 ng Brazz C1, and 551 ng Brazz C2 in 20 μL digestion reactions.
- We also ran undigested samples of all four to help determine why the other ligations (Brazz N and Mira C) did not work.
- We ran 279 ng of Mira N1, 258 ng Mira N2, 270 ng of Brazz C1, and 275.5 ng Brazz C2.
Gel
Gel Lanes:
1. 1kb plus ladder
2. Mira N1 XbaI/PstI
3. Mira N2 XbaI/PstI
4. Brazz C1 XbaI/PstI
5. Brazz C2 XbaI/PstI
6. 1kb plus ladder
7. Mira N1 undigested
8. Mira N2 undigested
9. Brazz C1 undigested
10. Brazz C2 undigested
- Gel shows bands consistent with successful ligation!
- Gel extraction and purification of Miraculin, StrepII & Stop insert and Brazzein, StrepII & Stop insert.
ODs
Mira N1: 5.4 ng/μL
Mira N2: 3.1 ng/μL
Brazz C1: 6.1 ng/μL
Brazz C2: 10.1 ng/μL
Ligation/Transformation of Mira/Brazz+StrepII
- Our Miraculin and Brazzein constructs with StrepII tags (pre-cut with EcoR1/Spe1) were re-ligated to STOP+V0120 backbone
| Insert DNA | BackBone DNA | |
|---|---|---|
| Mira N | 9 ng | 49.5 ng |
| Mira C | 22.4 ng | 49.5 ng |
| Brazz N | 37.4 ng | 49.5 ng |
| Brazz C | 14.3 ng | 53.4 ng |
| Control | 0 ng | 53.4 ng |
NEB T4 Ligase was used for all reactions
Digestion & Gel Purification of V24
- The pETDUET Expression Vector, V24, was digested with Not1/Spe1 for later use.
1039.9 ng of DNA was digested in both reactions
Digestion reactions were run on a 1% Agarose gel at 125V for 30 min
OD's
V24-1 N/S: 13.5 ng/μL
V24-2 N/S: 17.9 ng/μL
07-22-2010 [ top ]
07-23-2010 [ top ]
Genomic DNA extraction from Valencia Oranges
- We used Qiagen DNeasy Plant Mini Kit
- 100 mg (wet) sample from Flavedo tissue of orange
- Samples had little DNA content and dirty 260/280
ODs
Orange genomic DNA 1-1: 1.8ng/μL (260/280: .82)
Orange genomic DNA 1-2: 1.1ng/μL (260/280: .68)
Orange genomic DNA 2-1: 2.6ng/μL (260/280: .85)
Orange genomic DNA 2-2: 2.1ng/μL (260/280: .89)
- PCR ran with between 40ng and 95ng of genomic DNA
- Extension time increased to 1:30
Mira/Brazz StrepII & Stop
- Miniprepped per Qiagen protocol and made glycerol stocks
07-26-2010 [ top ]
- Digested miniprepped Mira/Brazz+StrepII+STOP constructs with Not1/Spe1 for ligation confirmation: failed
- Ran PCR product of Valencia Orange genome to confirm: failed
- Digested Mira/Brazz+StrepII constructs EcoRI/SpeI, STOP+V0120 EcoRI/XbaI
Digestion Reactions
Mira N: 888ng
Mira C: 888ng
Brazz N: 993ng
Brazz C: 1032ng
STOP: 781ng x 2 (max DNA available)
07-27-2010 [ top ]
Ligation of Mira/Brazz + StrepII to STOP + V0120
- Insert to Backbone ration of 3:1
Miraculin: 34ng Brazzein: 11ng Backbone: 50ng
- Ligations were transformed, plated onto LB+AMP and left overnight @ 37°C
PCR of Valencene using Pfx Polymerase
- Pfx polymerase was rumoured to be more accurate at PCR from genomic DNA
Specs:
DNA 1-1: 1.8 ng/μL
DNA 1-2: 1.1 ng/μL
DNA 2-1: 2.6 ng/μL
DNA 2-2: 2.2 ng/μL
- 1μL of each DNA sample was used for PCR. Cycling procedure can be found here: http://www.invitrogen.com/etc/medialib/en/filelibrary/pdf.Par.79411.File.dat/platinumpfx_pps.pdf
Mira/Brazz+StrepII+STOP confirmation digest
Digested DNA:
MN2: 211.2 ng
MC2: 190.2 ng
BN2: 184.4 ng
BC2: 219.2 ng
- Digestions were run on a 1% agarose gel at 125V for 30 minutes.
- Ladder
- MN2
- MC2
- BN2
- BC2
- Ladder
07-28-2010 [ top ]
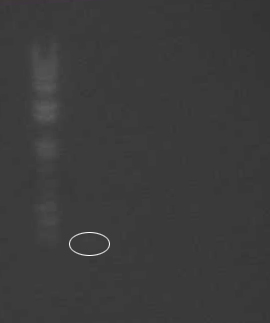
PCR Confirmation
- Ran gel to confirm Valencene PCR: failed
- Ladder
- DNA 1-1
- DNA 1-2
- DNA 2-1
- DNA 2-2
- Ladder
The numerical differentiation refers to the specific genomic DNA sample
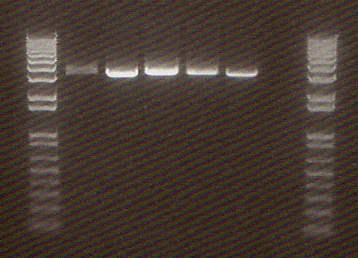
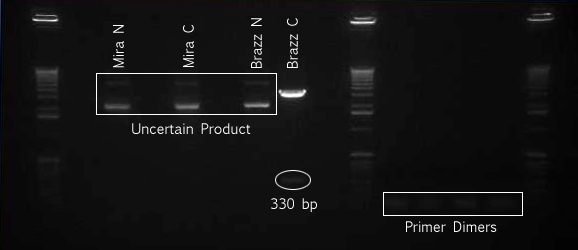
PCR of Wintergreen parts
- Ran PCR to extract J45004 and J45017 parts from the Wintergreen Pathway
Primers:
J45004_F
Left Primer: 5' cctttctagaatggaagttgttgaagttcttca 3'
J45004_R
Right Primer: 5' aaggctgcagcggccgctactagtttaatttattttggtcaagga 3' (last 5 bp omitted to meet 45 bp maximum)
J45017_F
Left Primer: 5' cctttctagaatgaaaactcccgaagactgc 3'
J45017_R
Right Primer: 5' aaggctgcagcggccgctactagtttattaggcgacgccgc 3'
The PCR reaction was set-up as per the specifications from the Phusion Polymerase manual (http://www.neb.com/nebecomm/ManualFiles/manualF-530.pdf). For template DNA, 3.5 ng of the J45700 BioBrick part (the entire wintergreen pathway) was used.
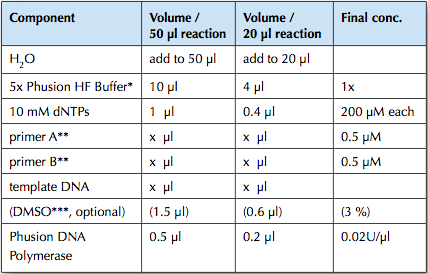
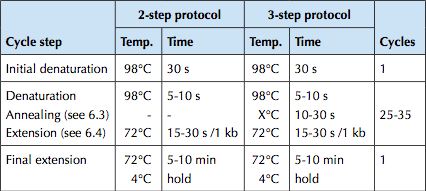
An annealing temperature of 62°C for 15s was used. Polymerase was allowed to extend for 60s; Phusion Polymerase extends at 1kb/15s, our longest construct is 1.7 kb
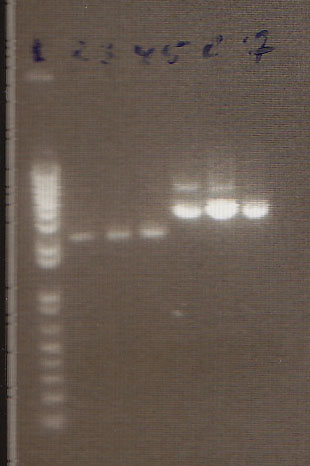
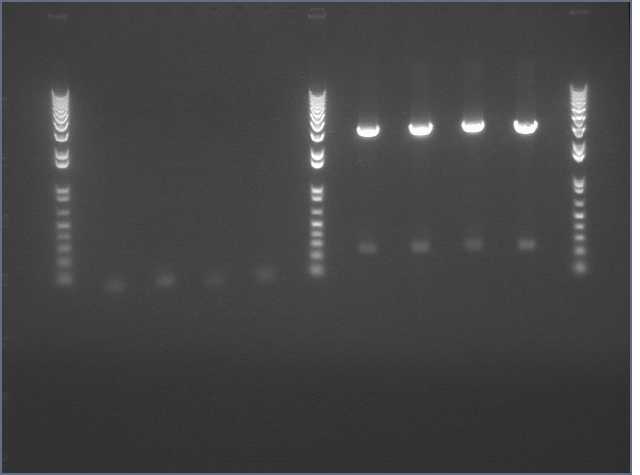
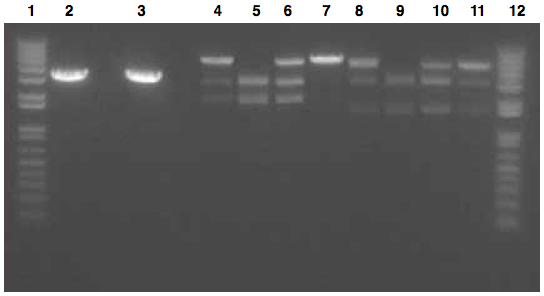
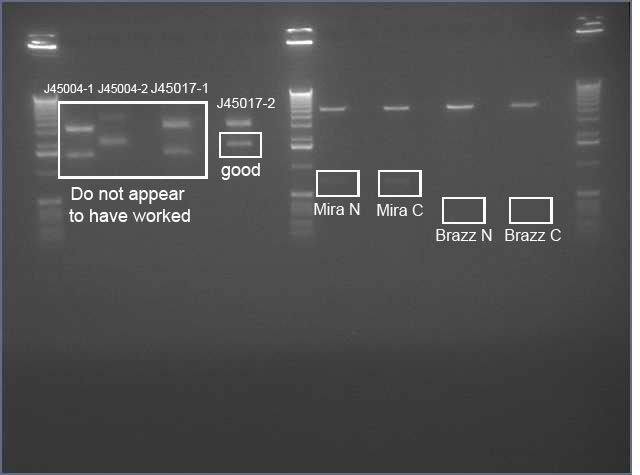
- Ladder
- J45004 PCR #1
- J45004 PCR #2
- J45017 PCR #1
- J45017 PCR #2
- Ladder
- Both J45004 PCR reactions appear to have worked, with product at the expected size of 1.1 kb.
- The J45017 PCR reactions appears to have amplified some of the wrong sequence, as suggested by the short DNA fragment. However, the longer ~2 kb fragment in PCR #1 does appear to be around the correct length.
07-29-2010 [ top ]
Minipreps
- Miniprep of Mira/Brazz+StrepII+STOP in V0120
Gel Extraction
- Gel extraction of J45017:
- Ladder
- Mira N N/S
- Mira C N/S
- Brazz N N/S
- Brazz C N/S
- J45004 X/P
- B21 X/P
- J45017 undigested
- Ladder
07-30-2010 [ top ]
- Ligated StrepII tagged proteins into V24 backbone
- Ligated cut J45004 into V0120 backbone
Ligations were plated and left @ 37°C O/N.
08-02-2010 [ top ]
DTL of Mira/Brazz+StrepII+Stop into V24
- Digested 1 microgram of DNA for each insert and for V24 backbone.
- Digested inserts and backbone with NotI/SpeI
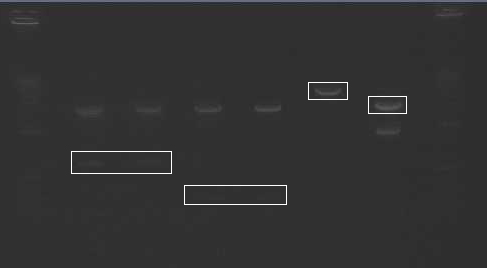
- Gel extracted and purified bands indicated in image below
- Ligated insert into 50ng of V24 backbone with a 3:1 insert to backbone ratio
- Transformed and plated on LB+AMP plates and left in 37°C incubator overnight
DTL of Wintergreen pathway
- Digested 400ng of J45004, 20ng of J45017, and 1 microgram of B21 (for v0120 backbone)
- Digested J45004, J45017, and B21 BB with XbaI/PstI
- PCR purified digested inserts
- Gel extracted and purified B21 BB (see gel image below)
- Ligated inserts into 50ng of B21 v0120 backbone with a 3:1 insert to backbone ratio
- Transformed and plated on LB+AMP plates and left in 37°C incubator overnight
Gel Purification
- Ladder
- Mira N SS
- Mira C SS
- Brazz N SS
- Brazz C SS
- V24 N/S
- B21 X/P
- Ladder
- Gel bands indicated were cut and purified from the gel.
08-03-2010 [ top ]
- Colonies were obtained from all ligations; 5mL cultures were started, left ~5hrs @ 37°C
- Glycerol stocks were made (666μL glycerol: 333μL cells; 50% glycerol final)
- Miniprepped per Qiagen protocol
ODs
004-1: 95.5 ng/μL
004-2: 124.5 ng/μL
017-1: 109.7 ng/μL
017-2: 77.5 ng/μL
Miraculin N+Strep+Stop: 61.5 ng/μL
Miraculin C+Strep+Stop: 78.5 ng/μL
Brazzein N+Strep+Stop: 83.4 ng/μL
Brazzein C+Strep+Stop: 99.4 ng/μL
08-04-2010 [ top ]
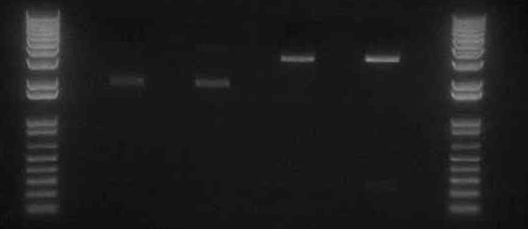
Confirmation Digests
- Ran confirmation digests of miniprepped J45004-1, J45004-2, J45017-1, J45017-2
- Digested approximately 500ng with xbaI/speI
- Ran confirmation digests of miniprepped Miraculin N strep+stop and V24, Miraculin C strep+stop and V24, Brazzein N strep+stop and V24, Brazzein C strep+stop and V24.
- Digested 400-500ng with notI/speI
Retry Ligating j45004 and v0120
- Used PCR purified J45004 that had been previously digested with xbaI/pstI
- Used v0120 from Team Vector that they had digested with xbaI/pstI and used in a successful ligation
- 3:1 insert to backbone ration used in ligation
- 52ng of insert and 50ng v0120 backbone
- J45004 insert is 1100 bp
- Transformed and plated on LB+Amp plates and left overnight
08-05-2010 [ top ]
Transformation of J45004
- 5μL of registry DNA was transformed with 15μL TURBO cells; mixture was plated on LB+AMP and left to grow O/N
08-06-2010 [ top ]
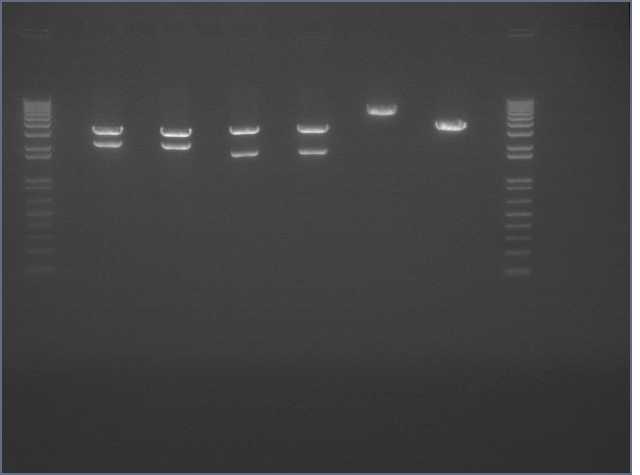
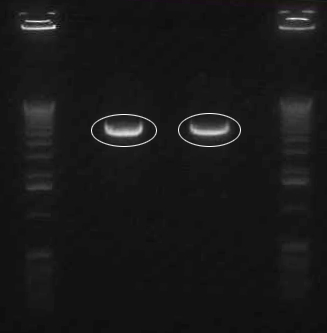
Confirmation Digest of J45004
- J45004 in V0120 backbone; digested ~500ng DNA with Xba1/Pst1
- 1 kb Ladder (Invitrogen)
- 004-1
- 004-2
- 004-3
- Ladder
Backbone: 3.2 kb; Insert: 1.1 kb
Analysis
Both the backbone and the insert in lane 1 look like the correct length (3.2 and 1.1 kb respectively). Lane two consists of a backbone of the correct length, but the insert appears to be ~1.5 kb, too long for the J45004 BioBrick part. Lane 3 appears undigested; we are uncertain as to why.
Transformation of Mira/Brazz StrepII constucts into E45
- 100-200 ng of construct DNA was transformed with 15μL of E45 expression E. Coli; transformation procedure was followed; transformed cells were plated on LB+AMP and left to grow O/N
Transformations: Mira N1 Mira N2 Mira C1 Mira C2 Brazz N1 Brazz N2 Brazz C1 Brazz C2
 "
"