Team:Lethbridge/Modeling
From 2010.igem.org
David.franz (Talk | contribs) (→Homology Modeling) |
Liszabruder (Talk | contribs) |
||
| Line 102: | Line 102: | ||
==<font color="white">Why Metabolic Modeling== | ==<font color="white">Why Metabolic Modeling== | ||
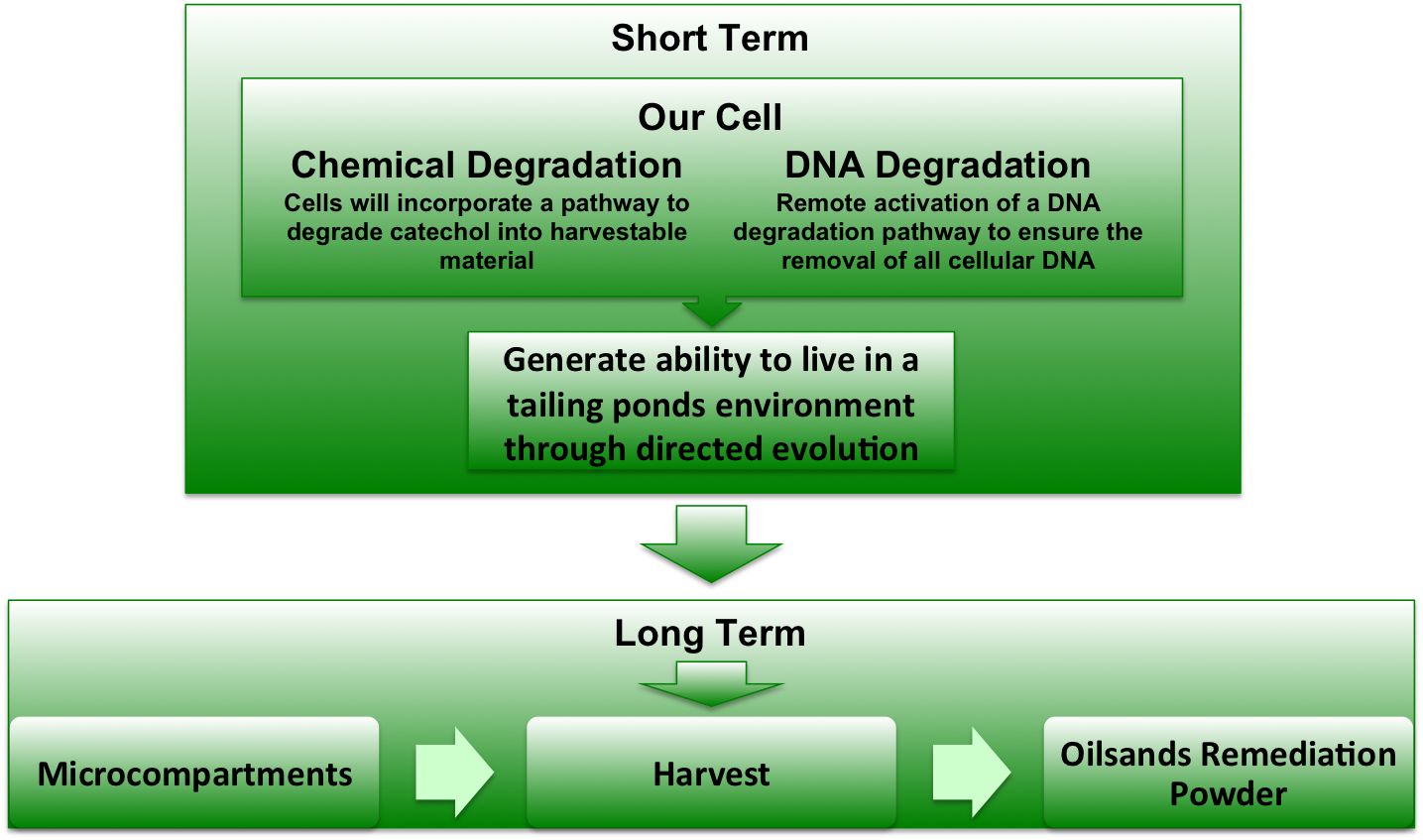
| - | The majority of degredation pathways for oilsands contaminants convert various organic compounds into catechol. Our project this year in the incorporation of the gene <i>xylE</i> into <i>E. coli</i>. The <i>xylE</i> gene encodes catechol-2,3,-dioxygenase, which converts catechol into 2-hydroxymuconate semialdehyde. This semialdehyde can be metabilized by <i>E. coli</i> by means of the glycolysis / Krebbs cycle. | + | The majority of degredation pathways for oilsands contaminants convert various organic compounds into catechol. Our project this year in the incorporation of the gene <i>xylE</i> into <i>E. coli</i>. The <i>xylE</i> gene encodes <html><a href="https://2010.igem.org/Team:Lethbridge/Project/Catechol_Degradation"><font color="green"> catechol-2,3,-dioxygenase</font></a></html>, which converts catechol into 2-hydroxymuconate semialdehyde. This semialdehyde can be metabilized by <i>E. coli</i> by means of the glycolysis / Krebbs cycle. |
[[Image:UofLProjectOverview.jpg|x150px|right|text-top]] | [[Image:UofLProjectOverview.jpg|x150px|right|text-top]] | ||
| Line 110: | Line 110: | ||
==<font color="white">The Model== | ==<font color="white">The Model== | ||
| + | |||
The proposed model will include the entire glycolysis / Krebbs cycle and the entry of catecol and 2-hydroxymuconate semialdehyde into it alongside normal metabolites. Databases (such as BRENDA) have detailed all the pathways and should have most if not all the kinetic parameters required. | The proposed model will include the entire glycolysis / Krebbs cycle and the entry of catecol and 2-hydroxymuconate semialdehyde into it alongside normal metabolites. Databases (such as BRENDA) have detailed all the pathways and should have most if not all the kinetic parameters required. | ||
==<font color="white">The Questions/Answers== | ==<font color="white">The Questions/Answers== | ||
| + | |||
1) What is the maximal rate of catechol degradation by the cell by this pathway? | 1) What is the maximal rate of catechol degradation by the cell by this pathway? | ||
| Line 125: | Line 127: | ||
[[Image:homology1.png|x400px|right|text-top]] | [[Image:homology1.png|x400px|right|text-top]] | ||
| - | Another aspect of our project is working on the localization of catechol-2,3,-dioxygenase (and other proteins) into the interior of microcompartments. To target the protein into the microcompartment (Lumazine modified to have an even more charged interior) requires the fusion of a polyarginine tail to either the C or N terminus of the protein. | + | Another aspect of our project is working on the localization of catechol-2,3,-dioxygenase (and other proteins) into the interior of <html><a href="https://2010.igem.org/Team:Lethbridge/Project/Compartamentalization"><font color="green"> microcompartments</font></a></html>. To target the protein into the microcompartment (Lumazine modified to have an even more charged interior) requires the fusion of a polyarginine tail to either the C or N terminus of the protein. |
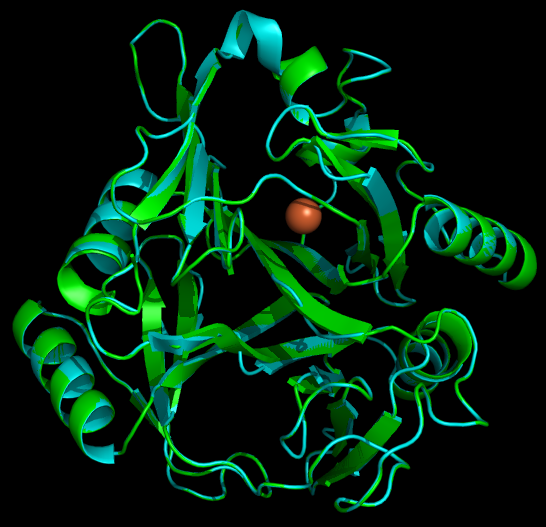
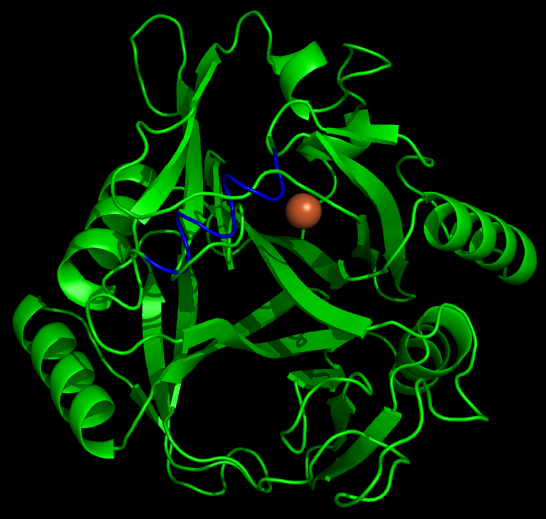
As with any fusion protien, the addition of this polyarginine tail to the protein has the very real potential of blocking (at least partially) the active site of a protein. Thus if one can predict the structure of the protein with the addition of the tail, it is possible to avoid creating inactive/ low efficiency fusion proteins. | As with any fusion protien, the addition of this polyarginine tail to the protein has the very real potential of blocking (at least partially) the active site of a protein. Thus if one can predict the structure of the protein with the addition of the tail, it is possible to avoid creating inactive/ low efficiency fusion proteins. | ||
| Line 134: | Line 136: | ||
| - | To model the | + | To model the xylE structure, the sequence for xylE from <i>Pseudomonas putida</i> (NCBI accession number NP_542866) was aligned with the primary sequence from the crystal structure of xylE from the same organism (pdb: 1MPY; several differences in amino acid sequence were observed) using CLUSTALW (Higgins <i>et al.</i>, 1996). Based on this sequence alignment, a homology model was generated using the alignment mode in SWISSMODEL (Guex <i>et al.</i>, 1997; Kiefer <i>et al.</i>, 2009; Arnold <i>et al.</i>, 2006). To model the placement of an N-terminal arginine tag, the tag was manually added to the N-terminus of the model. Energy minimization was carried out in SWISS-PDB viewer in vacuo utilizing a GROMOS96 energy minimization (Guex <i>et al.</i>, 1997). The resulting pdb file was visualized and manipulated using PYMOL, images were taken using the same software (DeLano, 2006). |
=<font color="white">References= | =<font color="white">References= | ||
| - | + | Arnold, K., Bordoli, L., Kopp, J., and Schwede, T. (2006) The SWISS-MODEL Workspace: A web-based environment for protein structure homology modelling., Bioinformatics 22, 195-201. | |
| - | + | <br> | |
| - | + | DeLano, W. L. (2006) PyMOL, DeLano Scientific. | |
| - | + | <br> | |
| - | + | Guex, N., and Peitsch, M. C. (1997) SWISS-MODEL and the Swiss-PdbViewer: An environment for comparative protein modelling, <i>Electrophoresis</i> 18, 2714-2723. | |
| - | + | <br> | |
| - | + | Higgins, D. G., Thompson, J. D., and Gibson, T. J. (1996) Using CLUSTAL for multiple sequence alignments, <i>Methods Enzymol.</i> 266, 383-402. | |
| - | + | <br> | |
| - | + | Kiefer, F., Arnold, K., Künzli, M., Bordoli, L., and Schwede, T. (2009) The SWISS-MODEL Repository and associated resources, <i>Nucleic Acids Res</i> 37, D387-D392. | |
| - | + | ||
| - | + | ||
| - | + | ||
<br> | <br> | ||
 "
"