Team:Imperial College London/Lab Diaries/XylE team
From 2010.igem.org
| Line 794: | Line 794: | ||
|- | |- | ||
| | | | ||
| + | |||
| + | <html> | ||
| + | <table width="850px" border="0"> | ||
| + | <tr> | ||
| + | <td style="background-color:#FFFF66"> | ||
| + | <b>Week 11</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;"><b>Monday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;"><b>Tuesday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;"><b>Wednesday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;"><b>Thursday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;"><b>Friday</b> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="background-color:#FFCC66;width:100px;text-align:top;"><b>Morning</b> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | <li>Gel-analysis of amplification-PCR for XylE-2</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | <li>midi-prep of Pveg-RBS vector</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | <li>In preparation of ''catechol cell-death assay'' tranformation of Top10 cells with CMR plasmid (psb1C3)</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | </ul> | ||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td style="background-color:#FFCC66;width:50px;text-align:top;"><b>Lunch Break</b> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:50px;width:200px;text-align:top;"> | ||
| + | </td> | ||
| + | <td style="background-color:#eeeeee;height:50px;width:200px;text-align:top;"> | ||
| + | </td> | ||
| + | <td style="background-color:#eeeeee;height:50px;width:200px;text-align:top;"> | ||
| + | </td> | ||
| + | <td style="background-color:#eeeeee;height:50px;width:200px;text-align:top;"> | ||
| + | </td> | ||
| + | <td style="background-color:#eeeeee;height:50px;width:200px;text-align:top;"> | ||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td style="background-color:#FFCC66;width:100px;text-align:top;"><b>Afternoon</b> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | <li>Gel-purification of amplification-PCR for XylE-2</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | <li>plating and overnight incubation of transformed top10 CMR</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | <li>overnight cultures for catechol cell death assay: Top10 cells XylE/CMR </li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:top;"> | ||
| + | <ul> | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </html> | ||
'''Monday 13th september''' | '''Monday 13th september''' | ||
Overnights weren't set up on sunday so they were made up alongside some assay cultures. J23101-XylE-B0014 colonies 8 and 10 of the replica plate #2 were picked. Chris also provided a replica plate containing 3k3 vector colonies. This was over a year old and he was unsure whether it was the correct plasmid or if the cells would grow up. I picked all available colonies 102 150 151 and 260 of kanamycin resistance. Lastly for assays 2x LB 2xM9 cultures were made 5ml +5ul antibiotic. | Overnights weren't set up on sunday so they were made up alongside some assay cultures. J23101-XylE-B0014 colonies 8 and 10 of the replica plate #2 were picked. Chris also provided a replica plate containing 3k3 vector colonies. This was over a year old and he was unsure whether it was the correct plasmid or if the cells would grow up. I picked all available colonies 102 150 151 and 260 of kanamycin resistance. Lastly for assays 2x LB 2xM9 cultures were made 5ml +5ul antibiotic. | ||
| - | + | '''Tues 14th September''' | |
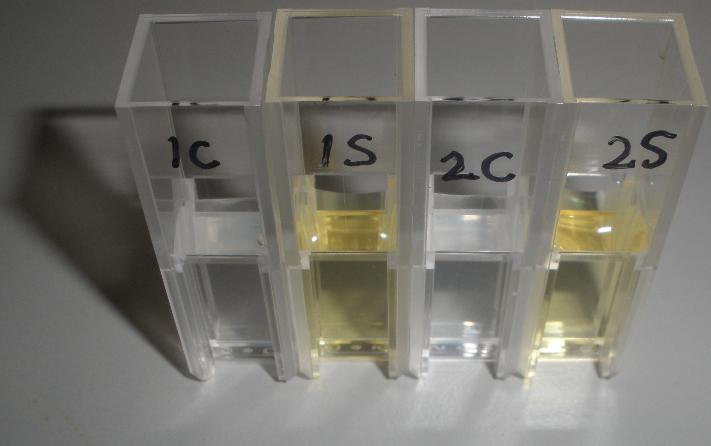
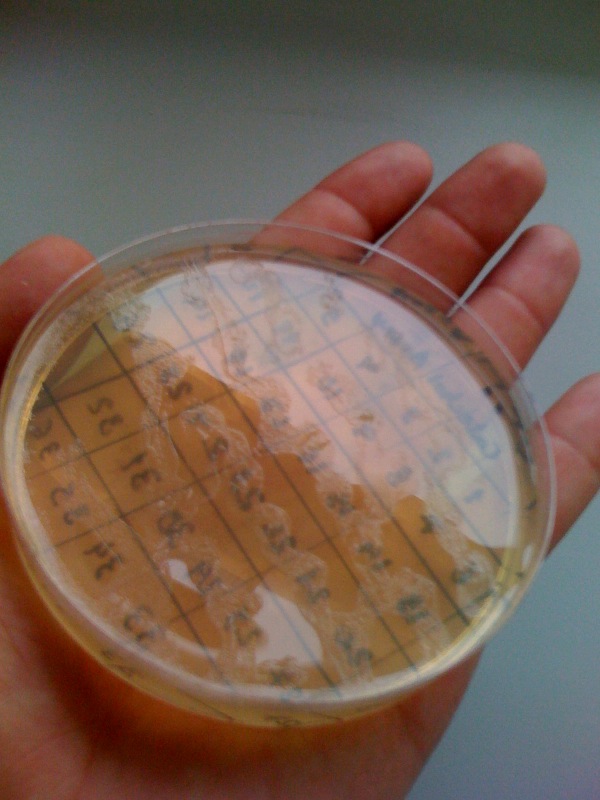
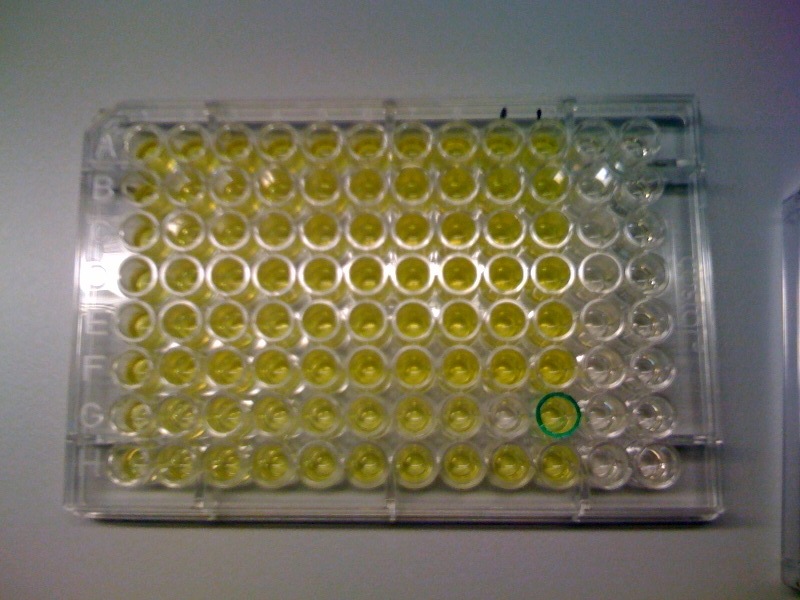
[[Image:CS1.JPG|thumb|right|300px|test confirming that yellow product of XylE enzmymatic reaction is leaks back from the cytosol into the solution.1C and 2C is the XylE producing cells after centrifuging and redissolving of the pellet and 1S and 2S is the supernatant after centrifuging]] | [[Image:CS1.JPG|thumb|right|300px|test confirming that yellow product of XylE enzmymatic reaction is leaks back from the cytosol into the solution.1C and 2C is the XylE producing cells after centrifuging and redissolving of the pellet and 1S and 2S is the supernatant after centrifuging]] | ||
*assay on plate reader of: | *assay on plate reader of: | ||
| Line 808: | Line 906: | ||
Piotr: | Piotr: | ||
| - | *Mini prep of 1 sample (6 samples lost due to mistake) of GFP-Xyle fusion protein and its digestion with Spe and Xba (gel to be run on the next day) | + | * Mini prep of 1 sample (6 samples lost due to mistake) of GFP-Xyle fusion protein and its digestion with Spe and Xba (gel to be run on the next day) |
| - | *Preparing E.Coli colonies for the next day for mini prep to be redone | + | * Preparing E.Coli colonies for the next day for mini prep to be redone |
| - | *PCR of 6 samples of GFP-Xyle fusion | + | * PCR of 6 samples of GFP-Xyle fusion |
| - | + | '''Weds 15th September''' | |
| - | *Midi preps of overnights. | + | * Midi preps of overnights. |
| - | *In preparation for assays determining the effect of catechol and/or breakdown product on cell viability we prepared Top10 cells, one strain containing pVeg-XylE-terminator and the other containing a CMR plasmid of similar size. As the XylE cells have already been prepared, only Top10 transformation with CMR-plasmid had to be carried out. | + | * In preparation for assays determining the effect of catechol and/or breakdown product on cell viability we prepared Top10 cells, one strain containing pVeg-XylE-terminator and the other containing a CMR plasmid of similar size. As the XylE cells have already been prepared, only Top10 transformation with CMR-plasmid had to be carried out. |
| - | + | '''Thurs 16th September''' | |
*Top10 CMR transformation was successful. Four overnight cultures of each Top10 XylE and Top10 CMR were set off with either M9 or LB medium respectively. | *Top10 CMR transformation was successful. Four overnight cultures of each Top10 XylE and Top10 CMR were set off with either M9 or LB medium respectively. | ||
*Piotr: Did mini preps from bacteria with blunt ended Xyle-GFP fusion and did diagnostic using both: | *Piotr: Did mini preps from bacteria with blunt ended Xyle-GFP fusion and did diagnostic using both: | ||
| Line 827: | Line 925: | ||
The PCR reactions were not very conclusive on the gels but digests allowed to determine that colonies 1->5 seem to have the right sizes of DNA in them and on Friday they will be prepared to be sent off for sequencing | The PCR reactions were not very conclusive on the gels but digests allowed to determine that colonies 1->5 seem to have the right sizes of DNA in them and on Friday they will be prepared to be sent off for sequencing | ||
| - | * | + | * Continuation of the midi from overnight step. A gel analysis was run which showed the correct bands were present. A gel purification of the protein was then performed and the bands excised. |
| - | + | '''Friday 17th September''' | |
*Catechol assay of transformed E.coli Top ten(J23101-XylE-terminator) in M9 medium + data analysis | *Catechol assay of transformed E.coli Top ten(J23101-XylE-terminator) in M9 medium + data analysis | ||
**[[Image:M9 catechol concentrations assay alternative.xls|Data analysis]] | **[[Image:M9 catechol concentrations assay alternative.xls|Data analysis]] | ||
| Line 842: | Line 940: | ||
|- | |- | ||
| | | | ||
| - | + | '''Monday 20th Sept''' | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Performed ligation again. Errors in the sequencing we received back regarding J23101-XylE-B0014. | Performed ligation again. Errors in the sequencing we received back regarding J23101-XylE-B0014. | ||
One was a log error (in XylE) phew. The other is in PSB1C3 out of the scar site and should be OK. | One was a log error (in XylE) phew. The other is in PSB1C3 out of the scar site and should be OK. | ||
Chris is currently assembling the pVEG promoter+ RBS in a vector. Once he has finished this, I will combine XylE-GFP and XylE with this promoter for comparison characterization with J23101 (in E.coli and Bacillus) | Chris is currently assembling the pVEG promoter+ RBS in a vector. Once he has finished this, I will combine XylE-GFP and XylE with this promoter for comparison characterization with J23101 (in E.coli and Bacillus) | ||
| - | + | '''Tues 21st Sept''' | |
| - | *Ligation to be transformed in a bit. (Issue with the vector not being cut efficiently=failed ligations?) | + | * Ligation to be transformed in a bit. (Issue with the vector not being cut efficiently=failed ligations?) |
| - | *I am cutting the original XylE midi as well as PSB1C3 containing B0014 and ligating them today (hopefully) the vector is purified, waiting on the gel run and then extraction of XylE insert. A ligation ratio must be determined from a gel analysis then dephosphorylation of the vector and then ligation. This will produce a promoter-less XylE+terminator vector which will be readily available to switch in the desired promoter (E+S cut) | + | * I am cutting the original XylE midi as well as PSB1C3 containing B0014 and ligating them today (hopefully) the vector is purified, waiting on the gel run and then extraction of XylE insert. A ligation ratio must be determined from a gel analysis then dephosphorylation of the vector and then ligation. This will produce a promoter-less XylE+terminator vector which will be readily available to switch in the desired promoter (E+S cut) |
| - | *Florian is performing a mini prep on pVeg promoter which if it is correctly identified in the gel i can use to ligate to my XylE + PSB1C3-B0014. | + | * Florian is performing a mini prep on pVeg promoter which if it is correctly identified in the gel i can use to ligate to my XylE + PSB1C3-B0014. |
| - | *Piotr is PCRing the reverse version of XylE that will be under the control of inducible promoter LacI (not delivered from synthesis yet) that will become a testing construct. | + | * Piotr is PCRing the reverse version of XylE that will be under the control of inducible promoter LacI (not delivered from synthesis yet) that will become a testing construct. |
| - | + | ||
| - | *Piotr's PCR has been successful for 3 samples (1. recommended annealing temp; 2. recommended + 2 degrees and 3. recommended + 4 degrees). Sample no1. has been run on the gel and gel purified. It has been cut overnight with SpeI and tommorrow in the morning XbaI will be added. | + | '''Wed 22nd Sept''' |
| + | * Piotr's PCR has been successful for 3 samples (1. recommended annealing temp; 2. recommended + 2 degrees and 3. recommended + 4 degrees). Sample no1. has been run on the gel and gel purified. It has been cut overnight with SpeI and tommorrow in the morning XbaI will be added. | ||
* I (Maddie!) performed steps towards the ligation (XylE-3K3) again from scratch (inefficient cutting steps??) including digest, gel analysis and extraction. | * I (Maddie!) performed steps towards the ligation (XylE-3K3) again from scratch (inefficient cutting steps??) including digest, gel analysis and extraction. | ||
| - | *I ligated PSBC3-B0014 and XylE | + | * I ligated PSBC3-B0014 and XylE |
| - | *I picked successfully transformed colonies for a replica plate and set up overnight mini cultures of 1,2,3 ComC DE promoter FWD and 4,5,6 ComC promoter Rev | + | * I picked successfully transformed colonies for a replica plate and set up overnight mini cultures of 1,2,3 ComC DE promoter FWD and 4,5,6 ComC promoter Rev |
| - | + | '''Thurs 23rd Sept First day of AUTUMN!!''' | |
*I determined the ligation ratio for XylE and 3k3 (1:3) performed dephosphorylation of vector then ligation, to be transformed along with the other ligation tomorrow hopefully. | *I determined the ligation ratio for XylE and 3k3 (1:3) performed dephosphorylation of vector then ligation, to be transformed along with the other ligation tomorrow hopefully. | ||
*I did mini preps of the Comc DE promoters, performed a ES test digest and am currently waiting for gel results. I shall then set up overnight midis. | *I did mini preps of the Comc DE promoters, performed a ES test digest and am currently waiting for gel results. I shall then set up overnight midis. | ||
| Line 882: | Line 975: | ||
|- | |- | ||
| | | | ||
| - | + | '''Monday 27th Sept''' | |
| - | + | * Successfull transformations: now have XylE-PSB1C3-B0014 (no promoter) and 3K3-J23101-XylE-B0014. No background plate colonies..many colonies for 3K3-XylE and a few for PSB1C3-XylE. Colonies have been replica plated (plus catechol assay plate for 3K3-XylE) and set up for mini prepping tomorrow. | |
| - | + | * Sumo tagged XylE will be given to us by Kirsten at some point this week and we will have to purify it. This can then be used for in vitro testing and comparisons made with the linker-XylE protein. | |
| - | + | * pVeg and ComC DE promoter FWD can be added into the promoter-less construct after it has been midi-prepped and digested accordingly. ComC DE promoter REV can be added onto the reverse XylE construct (PCR'd by Piotr) AFTER it has been put into PSB1C3 and is more manageable. Kirsten is taking care of the GFP-XylE fusion for now, including putting the pVeg promoter plus RBS infront of it (proving difficult.) We still need to receive LacI promoter to create the inducible test version of the fusion XylE-GFP protein. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | *Successfull transformations: now have XylE-PSB1C3-B0014 (no promoter) and 3K3-J23101-XylE-B0014. No background plate colonies..many colonies for 3K3-XylE and a few for PSB1C3-XylE. Colonies have been replica plated (plus catechol assay plate for 3K3-XylE) and set up for mini prepping tomorrow. | + | |
| - | *Sumo tagged XylE will be given to us by Kirsten at some point this week and we will have to purify it. This can then be used for in vitro testing and comparisons made with the linker-XylE protein. | + | |
| - | *pVeg and ComC DE promoter FWD can be added into the promoter-less construct after it has been midi-prepped and digested accordingly. ComC DE promoter REV can be added onto the reverse XylE construct (PCR'd by Piotr) AFTER it has been put into PSB1C3 and is more manageable. Kirsten is taking care of the GFP-XylE fusion for now, including putting the pVeg promoter plus RBS infront of it (proving difficult.) We still need to receive LacI promoter to create the inducible test version of the fusion XylE-GFP protein. | + | |
| - | + | '''Tues 28th Sept''' | |
| - | *Performed mini preps and then E+S / E+S AND AseI digests of colonies 1-7 3k3-XylE and 1-3 C3-B0014-XylE respectively. Gel runs showed them to be the correct thing :)))))) so, 3k3-XylE final vector is now left as 2-7 minis (catechol assay test showed colony 1 was background so DO NOT USE) in my DNA box (orange tubes). Colony 1 of the C3-B0014-XylE was selected for midi prepping and will be performed tomorrow. | + | * Performed mini preps and then E+S / E+S AND AseI digests of colonies 1-7 3k3-XylE and 1-3 C3-B0014-XylE respectively. Gel runs showed them to be the correct thing :)))))) so, 3k3-XylE final vector is now left as 2-7 minis (catechol assay test showed colony 1 was background so DO NOT USE) in my DNA box (orange tubes). Colony 1 of the C3-B0014-XylE was selected for midi prepping and will be performed tomorrow. |
| - | + | '''Weds 29th Sept''' | |
| - | *Midi prep of XylE-GFP fusion protein along with C3-B0014-XylE. The C3-B0014-XylE midi will then be used in 2x cloning digests (E+S) to insert pVeg and ComC DE FWD promoters. These will be obtained and also digested E+S to get out of their current vectors a gel analysis and purification will be run of all 3 in parallel, followed by a ligation ratio gel analysis, dephosphorylation of vector then ligation. | + | * Midi prep of XylE-GFP fusion protein along with C3-B0014-XylE. The C3-B0014-XylE midi will then be used in 2x cloning digests (E+S) to insert pVeg and ComC DE FWD promoters. These will be obtained and also digested E+S to get out of their current vectors a gel analysis and purification will be run of all 3 in parallel, followed by a ligation ratio gel analysis, dephosphorylation of vector then ligation. |
| - | *Primers for J23101-XylE arrive today so they can be used to PCR out J23101-XylE (From my J23101-Xyl-E midi) with blunt ends and this will be ligated into the final Spec vector so that we can get some data with Bacillus. | + | * Primers for J23101-XylE arrive today so they can be used to PCR out J23101-XylE (From my J23101-Xyl-E midi) with blunt ends and this will be ligated into the final Spec vector so that we can get some data with Bacillus. |
| - | *J23101-XylE-B0014 XylE-3K3 and PSB1C3-B0014-XylE were all sent off for sequencing | + | * J23101-XylE-B0014 XylE-3K3 and PSB1C3-B0014-XylE were all sent off for sequencing |
| - | + | '''Thurs 30th Sept''' | |
| - | *I gel purified ComCDE FWD promoter, C3-XylE-B0014, pVeg vector and XylE-B0014. Did a gel analysis to get ligation ratio. | + | * I gel purified ComCDE FWD promoter, C3-XylE-B0014, pVeg vector and XylE-B0014. Did a gel analysis to get ligation ratio. |
| - | *Set up midi culture of J23101-XylE-B0014 colony 8 | + | * Set up midi culture of J23101-XylE-B0014 colony 8 |
| - | *Set up midi culture to repeat GFP-XylE midi prep | + | * Set up midi culture to repeat GFP-XylE midi prep |
| - | *Set up 6x minis (plus replica plate) of final Spec colonies 1-6 | + | * Set up 6x minis (plus replica plate) of final Spec colonies 1-6 |
| - | *PCR reaction using newly arrived primers to get blunt ended J23101-XylE. | + | * PCR reaction using newly arrived primers to get blunt ended J23101-XylE. |
| - | + | '''Friday 1st Oct''' | |
*morning maddie, from earl's court! :) | *morning maddie, from earl's court! :) | ||
ahahahaha awesome. hey you :) | ahahahaha awesome. hey you :) | ||
| - | * | + | * Today I peformed ligations for ComCDE FWD promoter and PSB1C3-XylE-B0014 and also PSB1C3-pVeg and XylE-B0014 they will be left overnight and transformed by chris tmro thaaanks. |
| - | *I ran the PCR Kirill performed yday to get out blunt ended J23101-XylE on a gel; there was no template DNA! so I've just set up the reaction again... hopefully i'll purify it later for ligation into the final Spec vector that kyasha has maaaaade :))) then we can test XylE in Bacillus. | + | * I ran the PCR Kirill performed yday to get out blunt ended J23101-XylE on a gel; there was no template DNA! so I've just set up the reaction again... hopefully i'll purify it later for ligation into the final Spec vector that kyasha has maaaaade :))) then we can test XylE in Bacillus. |
| - | *I'm currently performing Midi's of my J23101-XylE-B0014 (running low) and of GFP-XylE (3rd time!) | + | * I'm currently performing Midi's of my J23101-XylE-B0014 (running low) and of GFP-XylE (3rd time!) |
|} | |} | ||
| Line 952: | Line 1,039: | ||
|- | |- | ||
| | | | ||
| - | |||
| - | + | '''Monday 4th Oct''' | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Did a replica plate and catechol assay plate/ minis 2 6 7 9 12 16. (Successful Pveg-XylE transformation) | Did a replica plate and catechol assay plate/ minis 2 6 7 9 12 16. (Successful Pveg-XylE transformation) | ||
| - | + | '''Tues 5th Oct''' | |
4 16 colonies were background the rest turned yellow. Mini prepped 2 6 7 9 12 16. Made overnight assay cultures of 3k3-XylE and C3-xylE | 4 16 colonies were background the rest turned yellow. Mini prepped 2 6 7 9 12 16. Made overnight assay cultures of 3k3-XylE and C3-xylE | ||
| - | + | '''Weds 6th Oct''' | |
| - | *Digested mini preps 2 6 7 9 12 16 with E+S will run on gel to determine if it's the correct plasmid, then will select a colony to midi prep tomorrow. | + | * Digested mini preps 2 6 7 9 12 16 with E+S will run on gel to determine if it's the correct plasmid, then will select a colony to midi prep tomorrow. |
| - | *Will perform the same assays already performed by Nick with the 3k3-XylE constructs to compare results. | + | * Will perform the same assays already performed by Nick with the 3k3-XylE constructs to compare results. |
| - | *Just realised Im working with background colony 16!! disregard this after the gel run! | + | * Just realised Im working with background colony 16!! disregard this after the gel run! |
| - | *I'm currently keeping my undigested PVeg-XylE minis in florians DNA box (orange lids and labels) | + | * I'm currently keeping my undigested PVeg-XylE minis in florians DNA box (orange lids and labels) |
| - | + | '''Thursday 7th Oct''' | |
| - | *Labpartner, goodmorning and from greece!! :p | + | * Labpartner, goodmorning and from greece!! :p |
| - | *Niiiiiiiiiiiiiiick!!! you left the country! you were too ugly. good luck babe! | + | * Niiiiiiiiiiiiiiick!!! you left the country! you were too ugly. good luck babe! |
| - | *Midi prepped PVEG-XylE-B0014 in PSB1C3; will digest out the insert (E+S) and put it into 3K3(E+X). Then both the midi (in PSB1C3) and the ligation 3k3-PVEG-XylE-B0014 can be transformed into TOP10. J23101-XylE-B0014 in 3K3 Vector also needs to be put into TOP10 currently only the PSB1C3 version is in TOP10. We need the rest for comparison testing as TOP10 is the more widely used testing strain. | + | * Midi prepped PVEG-XylE-B0014 in PSB1C3; will digest out the insert (E+S) and put it into 3K3(E+X). Then both the midi (in PSB1C3) and the ligation 3k3-PVEG-XylE-B0014 can be transformed into TOP10. J23101-XylE-B0014 in 3K3 Vector also needs to be put into TOP10 currently only the PSB1C3 version is in TOP10. We need the rest for comparison testing as TOP10 is the more widely used testing strain. |
| - | *Florian kindly digested PVeg-XylE-B0014-C3 with E+S | + | * Florian kindly digested PVeg-XylE-B0014-C3 with E+S |
| - | *Ran PCR to get out blunt PVeg-XylE-B0014 using primers. Did 3 reactions at variations around 60 degrees. | + | * Ran PCR to get out blunt PVeg-XylE-B0014 using primers. Did 3 reactions at variations around 60 degrees. |
| - | + | '''Friday 8th Oct''' | |
| - | *Ran gel analysis of my PCRs 1, 2 and 3* appear to have worked. Samples 1 and 3* show stronger bands and so should be used in the next ligation step into the SPEC vector (which kyasha will be doing) then transformation into Bacillus (meeee.) | + | * Ran gel analysis of my PCRs 1, 2 and 3* appear to have worked. Samples 1 and 3* show stronger bands and so should be used in the next ligation step into the SPEC vector (which kyasha will be doing) then transformation into Bacillus (meeee.) |
| - | *Chris is purifying digested PVeg-XylE-B0014-C3 to get the insert out and this will then be ran on a gel alongside cut (E+X) 3k3 to determine a ratio for ligation. | + | * Chris is purifying digested PVeg-XylE-B0014-C3 to get the insert out and this will then be ran on a gel alongside cut (E+X) 3k3 to determine a ratio for ligation. |
| - | *Meeting today at 4pm, no one is here!! it'll be me florian ben and piotr attending..yikes | + | * Meeting today at 4pm, no one is here!! it'll be me florian ben and piotr attending..yikes |
| - | + | '''Sunday 10th of the 10th of the 2010!!!!! and im in LAB ''' | |
| - | *I did ligations of PVeg-XylE into 3k3 and of Reverse-XylE into the digested C-Tev LacI vector. | + | * I did ligations of PVeg-XylE into 3k3 and of Reverse-XylE into the digested C-Tev LacI vector. |
| - | *I also set up assay cultures for monday 2x PVegXE 2x XE-C3 2x XE-3K3 and 2x GFP-XylE!!! | + | * I also set up assay cultures for monday 2x PVegXE 2x XE-C3 2x XE-3K3 and 2x GFP-XylE!!! |
|} | |} | ||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| Line 994: | Line 1,075: | ||
|- | |- | ||
| | | | ||
| - | + | ||
| - | + | '''Monday 11th Oct''' | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
*No Nicolas Kylilis, what a douche! | *No Nicolas Kylilis, what a douche! | ||
*Diluted my overnights 250ul into 5ml to grow up to around 0.5 OD | *Diluted my overnights 250ul into 5ml to grow up to around 0.5 OD | ||
| Line 1,006: | Line 1,083: | ||
*Finalising the polo-shirt logo design, it's looking fiiiine, we're gonna look goooood ;) | *Finalising the polo-shirt logo design, it's looking fiiiine, we're gonna look goooood ;) | ||
| - | + | '''Tues 12th Oct''' | |
*Transformations worked! we now have PVEG-XylE in 3k3 and reverse-xylE with lacI promoter (C-TEV vector)i did replica plates, catechol assay plates and overnights for minis. Will mini prep tmro and transform them into TOP10 and we should be good to go with all constructs. | *Transformations worked! we now have PVEG-XylE in 3k3 and reverse-xylE with lacI promoter (C-TEV vector)i did replica plates, catechol assay plates and overnights for minis. Will mini prep tmro and transform them into TOP10 and we should be good to go with all constructs. | ||
* Assays messed up, my fault :( 3k3 didnt respond to catechol, so i think that I picked a background colony, i wont pick it again. The results are a bit ridiculous and will probably be chucked. | * Assays messed up, my fault :( 3k3 didnt respond to catechol, so i think that I picked a background colony, i wont pick it again. The results are a bit ridiculous and will probably be chucked. | ||
| - | + | '''Weds 13th Oct''' | |
mini prepped 3k3-Pveg-XylE-B0014 and went to the school workshop! set off overnight midi of culture 9 | mini prepped 3k3-Pveg-XylE-B0014 and went to the school workshop! set off overnight midi of culture 9 | ||
| - | + | '''Thurs 14th Oct''' | |
Midi prepped 3k3-Pveg | Midi prepped 3k3-Pveg | ||
| - | + | '''Fri 15th''' | |
Transformed all my constructs into T0P10 3k3-J23101/pveg and PSB1C3-J23101/PVEG | Transformed all my constructs into T0P10 3k3-J23101/pveg and PSB1C3-J23101/PVEG | ||
|} | |} | ||
==Output Photo Gallery== | ==Output Photo Gallery== | ||
Revision as of 22:42, 24 October 2010
| Lab Diaries | Overview | Surface Protein Team | XylE Team | Vectors Team | Modelling Team |
| Here are the technical diaries for our project. We've split them up into three lab teams and the modelling team. We think it's really important that absolutely anyone can find out what we've been doing. For a really detailed look at what we did, and when, you've come to the right place! | |
| Team XylE |
Follow our progress: Click me!
| XylE team Lab Objectives |
|
| Week 6 | ||||||||||||||||||||||||
|
we constructed the standard E.coli promoter J23101 with sticky ends. These ends are complementary to restriction sites made by EcoRI and SpeI enzyme. This promoter will be later used in 3A assemply to construct a promoter-RBS-XylE design in a psB1C3 vector. E.coli will be transformed with this final construct plasmid to assess XylE activity and characterization. It will also be one of the submitted biobricks.
these cultures are going to be used tomorrow for mini-prepping. Miniprep will allow us to isolate E.coli's plasmid DNA(which contains the XylE gene). Friday, 13-Aug-2010
Mini-prep is usually used to confirm that our gene of interest has not been changed in any way, as the isolated plasnid id sent for sequencing. However, since XylE was taken from the registry, we assume that it is fine and no sequencing is required. The mini-prep will later be used for the midi-prep (that gives out higher yeilds of DNA needed for cloning).
|
| Week 7 | ||||||||||||||||||||||||
|
Tuesday, 17-Aug-2010
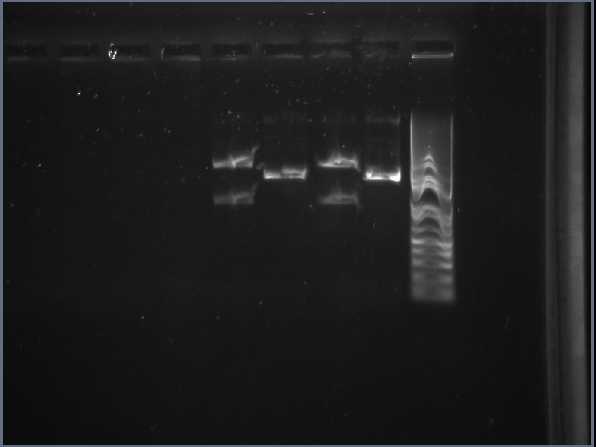
gel analysis of XylE, J23101 promoter and pSB1C3 vector samples to determine the volume ratios of samples to be used for 3A assemply ligation
Thursday, 19-Aug-2010
Friday, 20-Aug-2010 The J23101 gene in a biobrick vector containg RFP gene
|
| Week 8 | ||||||||||||||||||||||||
|
Monday, 23-Aug
Tuesday, 24th-Aug
Wednesday, 25th-Aug Performed gel analysis on the purified XylE and J23101 to obtain ratios for ligation. First gel was scrapped as it produced appauling(explanation for Nick:really bad) results, 2nd gel run was successful.
Thursday, 26th-Aug
Friday, 27th-Aug
Saturday, 28th-Aug
Sunday, 29th-Aug
|
| Week 10 | ||||||||||||||||||||||||
|
Tuesday 7th Sept
Wednesday, 8th Sep
Thursday, 9th Sep
Friday, 10 sep The transformation was a SUCCESS. 2x replica plates were made plate 1# 1-6 plate #2 6-11; colony 6 and colony 9 of plates 1 and 2 respectively were transfered into a 5ml liquid culture + 5ul CmR. These will later be turned into glycerol stocks. After the replica plates have grown up mini preps on a number of colonies shall be performed - this hopefully will eliminate the contaminating plasmid DNA. This will be followed by a midi prep. Saturday, 11 Sep The transformation of E.Coli with PSB1-C3 with insert did not work :( |
| Week 11 | ||||||||||||||||||||||||
|
Monday 13th september Overnights weren't set up on sunday so they were made up alongside some assay cultures. J23101-XylE-B0014 colonies 8 and 10 of the replica plate #2 were picked. Chris also provided a replica plate containing 3k3 vector colonies. This was over a year old and he was unsure whether it was the correct plasmid or if the cells would grow up. I picked all available colonies 102 150 151 and 260 of kanamycin resistance. Lastly for assays 2x LB 2xM9 cultures were made 5ml +5ul antibiotic. Tues 14th September
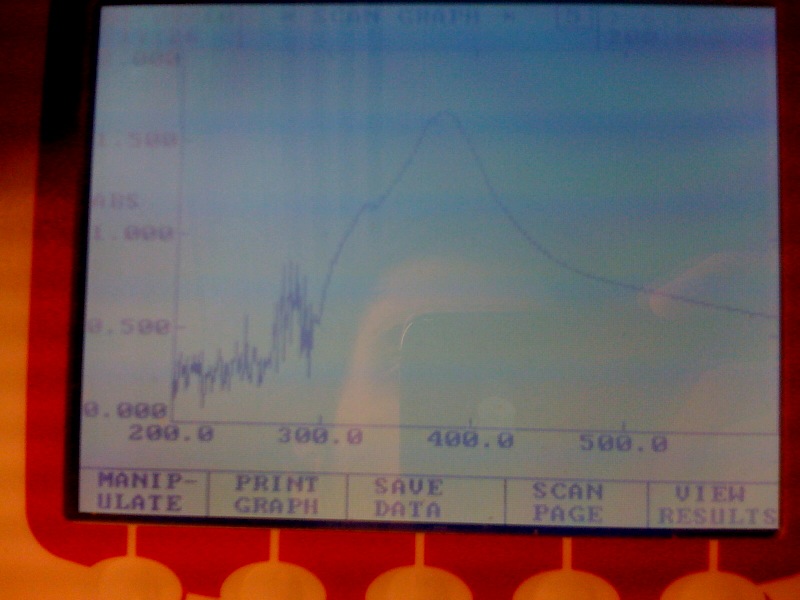
The assay was carried out with E.coli, top ten spcies, transformed with J23101-XylE-B0014 in pSB1C3 vector.The overnight culture wastransfered in new medium this morning for 4 hrs before assaying. LB medium was used for dilutions and blank. Catechol was diluted in ddH2O. Data analysis of the assay
Piotr:
Weds 15th September
Thurs 16th September
- digest with XbaI and SpeI - PCR reactions with the following primers added: HIS-GFP, XylE-Xba and the other sample with HIS-GFP and GFP-flag The PCR reactions were not very conclusive on the gels but digests allowed to determine that colonies 1->5 seem to have the right sizes of DNA in them and on Friday they will be prepared to be sent off for sequencing
Friday 17th September
The gel purifications of 3k3 vector and XylE were used for ligation. 3K3 was dephosphorylated and then ligated in a ratio 5:1 with the insert. This was left overnight and Chris tried to transform with it on sunday.. ligation and transformation failed :-( |
| Week 12 |
|
Monday 20th Sept Performed ligation again. Errors in the sequencing we received back regarding J23101-XylE-B0014. One was a log error (in XylE) phew. The other is in PSB1C3 out of the scar site and should be OK. Chris is currently assembling the pVEG promoter+ RBS in a vector. Once he has finished this, I will combine XylE-GFP and XylE with this promoter for comparison characterization with J23101 (in E.coli and Bacillus) Tues 21st Sept
Wed 22nd Sept
Thurs 23rd Sept First day of AUTUMN!!
Fri 24th Sept
|
| Week 13 |
|
Monday 27th Sept
Tues 28th Sept
Weds 29th Sept
Thurs 30th Sept
Friday 1st Oct
ahahahaha awesome. hey you :)
|
have a look :)it's from John Hoppkins wiki 2008. There are one or two good ones! Love: Before I heard the doctors tell The dangers of a kiss; I had considered kissing you. The nearest thing to bliss. But now I know biology and sit and sigh and moan; six million mad bacteria and I thought we were alone!
|
| Week 14 |
|
Monday 4th Oct Did a replica plate and catechol assay plate/ minis 2 6 7 9 12 16. (Successful Pveg-XylE transformation) Tues 5th Oct 4 16 colonies were background the rest turned yellow. Mini prepped 2 6 7 9 12 16. Made overnight assay cultures of 3k3-XylE and C3-xylE Weds 6th Oct
Friday 8th Oct
Sunday 10th of the 10th of the 2010!!!!! and im in LAB
|
| Week 15 |
|
Monday 11th Oct
Tues 12th Oct
Weds 13th Oct mini prepped 3k3-Pveg-XylE-B0014 and went to the school workshop! set off overnight midi of culture 9 Thurs 14th Oct Midi prepped 3k3-Pveg Fri 15th Transformed all my constructs into T0P10 3k3-J23101/pveg and PSB1C3-J23101/PVEG |
 "
"