Team:Valencia/prion
From 2010.igem.org
(→Prions) |
|||
| Line 16: | Line 16: | ||
==Prions== | ==Prions== | ||
| - | In 1982 Stanley B. Prusiner created the term “prion” (or proteinacius infectious particle) to name the exclusively proteic infectious agent responsible of the transmissible spongiform encephalopathies (TSEs), a group of mammalian neurodegenerative disorders. According | + | In 1982 Stanley B. Prusiner created the term “prion” (or proteinacius infectious particle) to name the exclusively proteic infectious agent responsible of the transmissible spongiform encephalopathies (TSEs), a group of mammalian neurodegenerative disorders. According to the widely supported “protein-only” model, the prion mechanism of transmissibility arise from the ability of the prion form of the protein to promote the conformational change of the normal cellular form to the infectious prion forms (Prusiner, 1998). The infectious forms are mis-folded proteins that induce by polymerization the formation of an amyloid fold constituted by tightly packed beta sheets. These aggregates are insoluble fibrils that display resistance to proteolytic digestion and have affinity for aromatic dyes. |
===Fungal prions=== | ===Fungal prions=== | ||
Revision as of 18:02, 23 October 2010
Time goes by...
(El tiempo pasa...)
Follow us:

Our main sponsors:

Our institutions:

Visitor location:
Using a Prion Switch
Contents |
Prions
In 1982 Stanley B. Prusiner created the term “prion” (or proteinacius infectious particle) to name the exclusively proteic infectious agent responsible of the transmissible spongiform encephalopathies (TSEs), a group of mammalian neurodegenerative disorders. According to the widely supported “protein-only” model, the prion mechanism of transmissibility arise from the ability of the prion form of the protein to promote the conformational change of the normal cellular form to the infectious prion forms (Prusiner, 1998). The infectious forms are mis-folded proteins that induce by polymerization the formation of an amyloid fold constituted by tightly packed beta sheets. These aggregates are insoluble fibrils that display resistance to proteolytic digestion and have affinity for aromatic dyes.
Fungal prions
In 1994 Reed Wickner proposed the prion nature of Ure2, a protein involved in the nitrogen metabolism of the yeast Saccharomyces cerevisae, to explain the unusual dominant and cytoplasmatic inheritance of the phenotype [URE3] first described by Cox (1965). In later years a wide array of genetic and biochemical evidence have supported that the prionic behaviour is present in other proteins of the yeast such as Sup35, Rnq1 and Swi1 and in HET-s, a protein involved in the mechanism of genetic incompatibility between strains of Podospora anserina.
These prions can produce amyloid-like fibrils similar to those associated with the mammalian prions. The molecular architecture of these amyloids have been studied using solid-state NMR spectroscopy and it has been found that the fibrils formed by Ure2p, Rnq1, and Sup35 share a common parallel and in-register β-structure (Fig.1) (Wickner et al. 2008). But these fungal prions have no sequence similarity with their mammalian counterparts. Besides they are not generally pathogenic and might have a beneficial role providing an evolutionary advantage to their hosts (). The prion domains of these fungal proteins have an exceptionally high content (~40%) of two polar and amyloidogenic amino acids, glutamine and asparagine.
Sup35p
[PSI+] is a non-Mendelian trait of Saccharomyces cerevisae that supress nonsense codons. This phenotype is due to a self-replication conformation (prion state) of a protein encoded by the gene Sup35. This protein, Sup35p, is the yeast eukariotyc release factor 3 (eRF3) and forms the translation termination complex with Sup45p (eRF1). The function of Sup45p is releasing the nascent polypeptide chain from the ribosome through GTP hydrolysis when Sup45p recognize a stop codon.
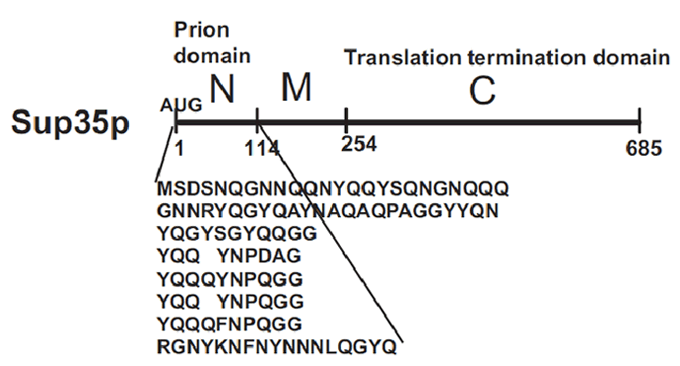
Sup35p is 685 amino acids long and has three distinct parts (Fig.2). The NH2-terminus (N) is termed the prion-forming domain (PrD) because plays a critical role in Sup35p’s changes in proteic conformation and it is responsible for its prion behaviour. This domain is 114 amino acids long and has a high content in glutamine and asparagine. The middle region (M) provides a solubilizing and/or spacing function. Finally, the COOH-terminus (C) is responsible for the translation-termination activity.
In [PSI+] cells, most Sup35p is insoluble and nonfunctional, causing an increase in the translational read-through of stop codons. This trait is heritable because Sup35p in the amyloid state as every prion influences new Sup35p to adopt the same conformation and passes from mother cell to daughter. In [psi-] cells, the translation-termination factor Sup35p is soluble and functional.
Prion switch
Parts
The switch is formed by two different parts: the activator and the reporter. The activator part is a construction of two fragments: the NM domains of the protein Sup35, which confers to the protein the prionic activity, and the GR526 portion, which contains the DNA-binding and transcription-activation domains. The ligand-binding domain of the protein GR was eliminated, decoupling the response of the protein to the presence of glucocorticoids, and thus generating a constitutive transcription activation factor. The normal activity of this protein results in the activation of the genes preceded by the GRE (Glucocorticoid Response Element). When exposed to heat shock or other stress conditions, the NM domains start the prionic activity, eventually inhibiting the activation of transcription.
 "
"