Team:Newcastle/Filamentous Cells
From 2010.igem.org
Swoodhouse (Talk | contribs) (→Computational model) |
Swoodhouse (Talk | contribs) (→Computational model) |
||
| Line 42: | Line 42: | ||
|Visualisation of the model's biochemical network in CellDesigner. | |Visualisation of the model's biochemical network in CellDesigner. | ||
|} | |} | ||
| + | |||
| + | Downloads: | ||
| + | *[[Media:Newcastle_filamentous.mod.txt|SBML-shorthand]] | ||
| + | *[[Media:Newcastle_filamentous.xml.txt|SBML]] | ||
==Cloning strategy== | ==Cloning strategy== | ||
Revision as of 16:16, 26 October 2010

| |||||||||||||
| |||||||||||||
Contents |
Filamentous cell formation by overexpression of yneA
Bacillus subtilis cell division is dependent on FtsZ. FtsZ forms a 30 subunit ring at the midpoint of the cell and contracts.
YneA indirectly stops the formation of the FtsZ ring. In nature, yneA is expressed during SOS response, allowing the cell to repair DNA damage before continuing with the division cycle.
It is hypothesized that YneA acts through unknown transmembrane proteins to inhibit FtsZ ring formation; we call these unknown components "Blackbox proteins".
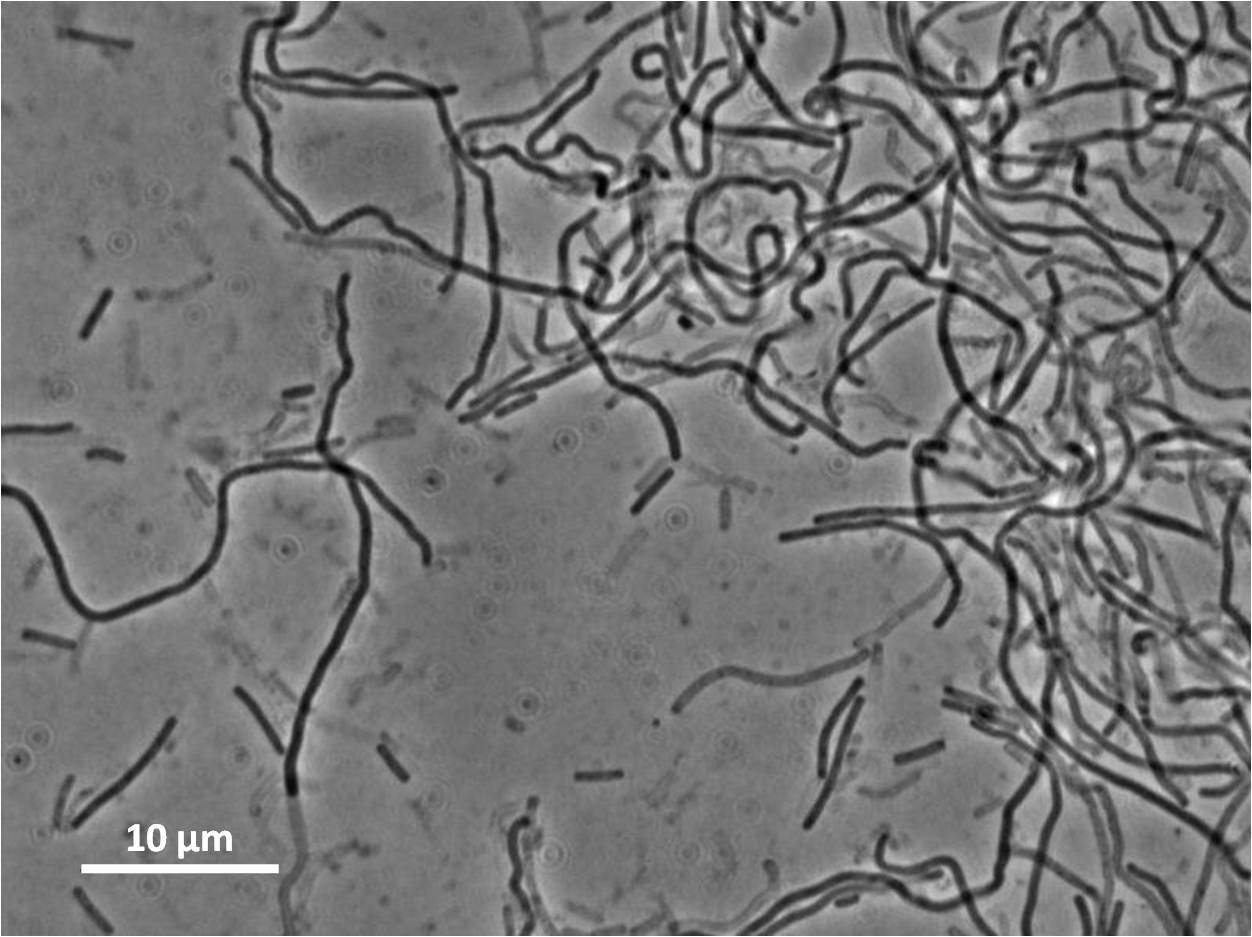
By expressing YneA and therefore inhibiting FtsZ ring formation, the cells will grow filamentous.
Part
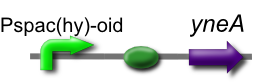
Our IPTG-inducible filamentous cell formation part puts yneA under the control of the [http://partsregistry.org/wiki/index.php?title=Part:BBa_K302003 strongly LacI-repressible promoter that we designed, hyperspankoid]. In the presence of LacI, induction with IPTG will result in a filamentous cell phenotype.
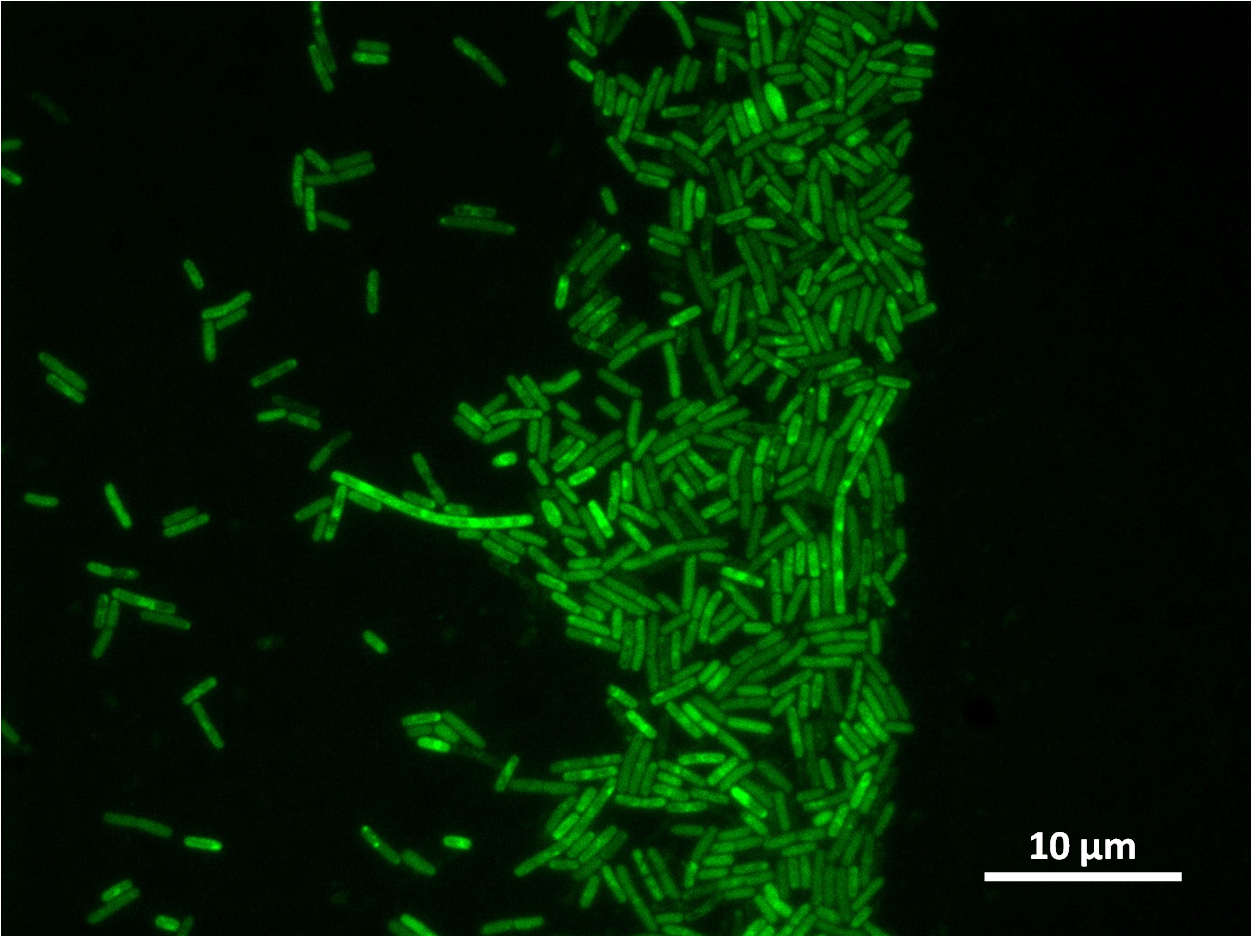
The part has no terminator, allowing for transcriptional fusion with gfp and visualisation under the microscope.
This is part [http://partsregistry.org/Part:BBa_K302012 BBa_K302012] on the [http://partsregistry.org parts registry].
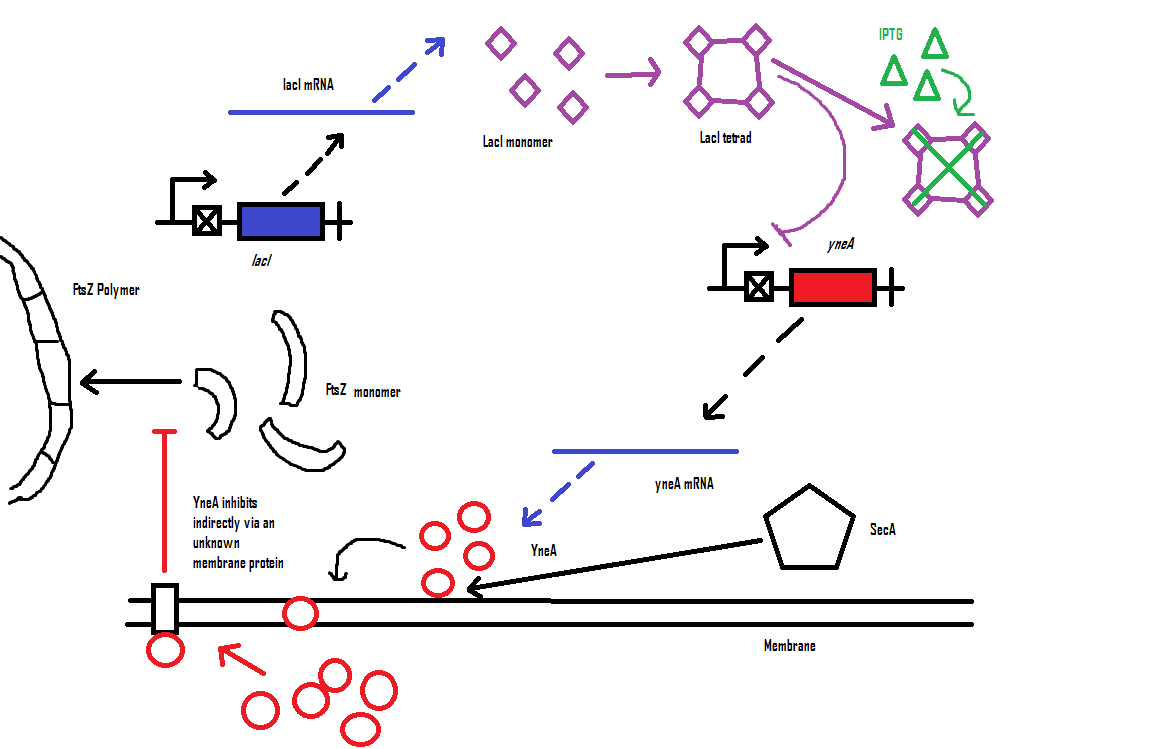
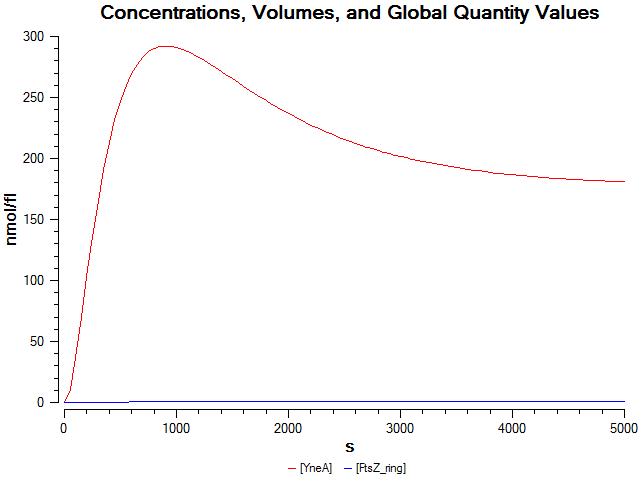
Computational model
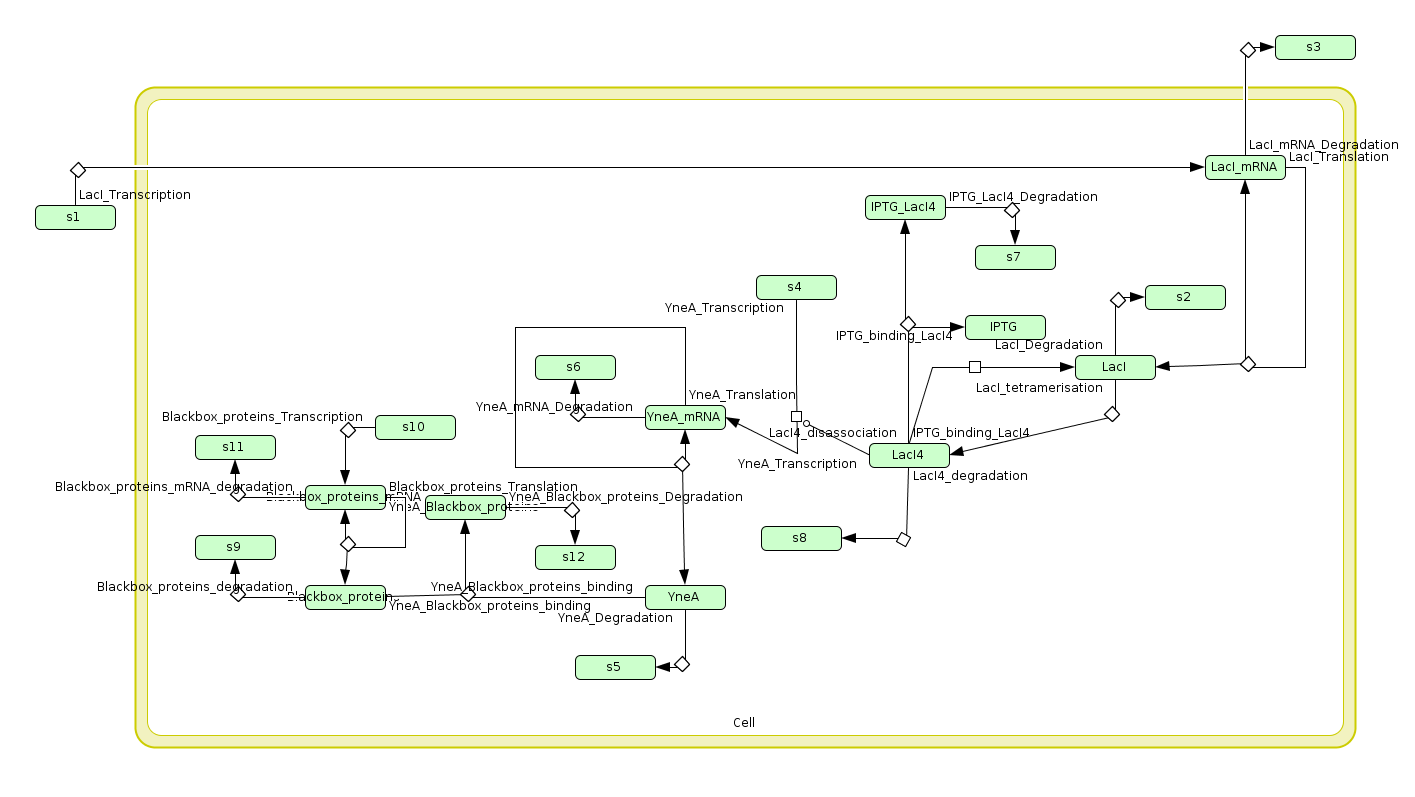
| Visualisation of the model's biochemical network in CellDesigner. |
Downloads:
Cloning strategy
Characterisation
Testing and Characterisation
Selection for integration
To select for integration of the plasmid into the chromosome, B. subtilis will be tested for the ability to hydrolyse starch. Integration of the BioBrick will be done by homologous recombination at the amyE, therefore destroying endogenous expression of amylase. Colonies that are not able to break down starch on agar plate will be selected and cultured for further test. Colonies that do not contain the integrated BioBrick will be able to hydrolyse starch, therefore forming a white halo around the colony as iodine interacts with starch to form blue colour.
The construct we designed has a GFP transcriptional fusion after the yneA coding sequence, so GFP is co-transcribed and acts as our fluorescent marker for transcription of yneA.
Lab work and Results
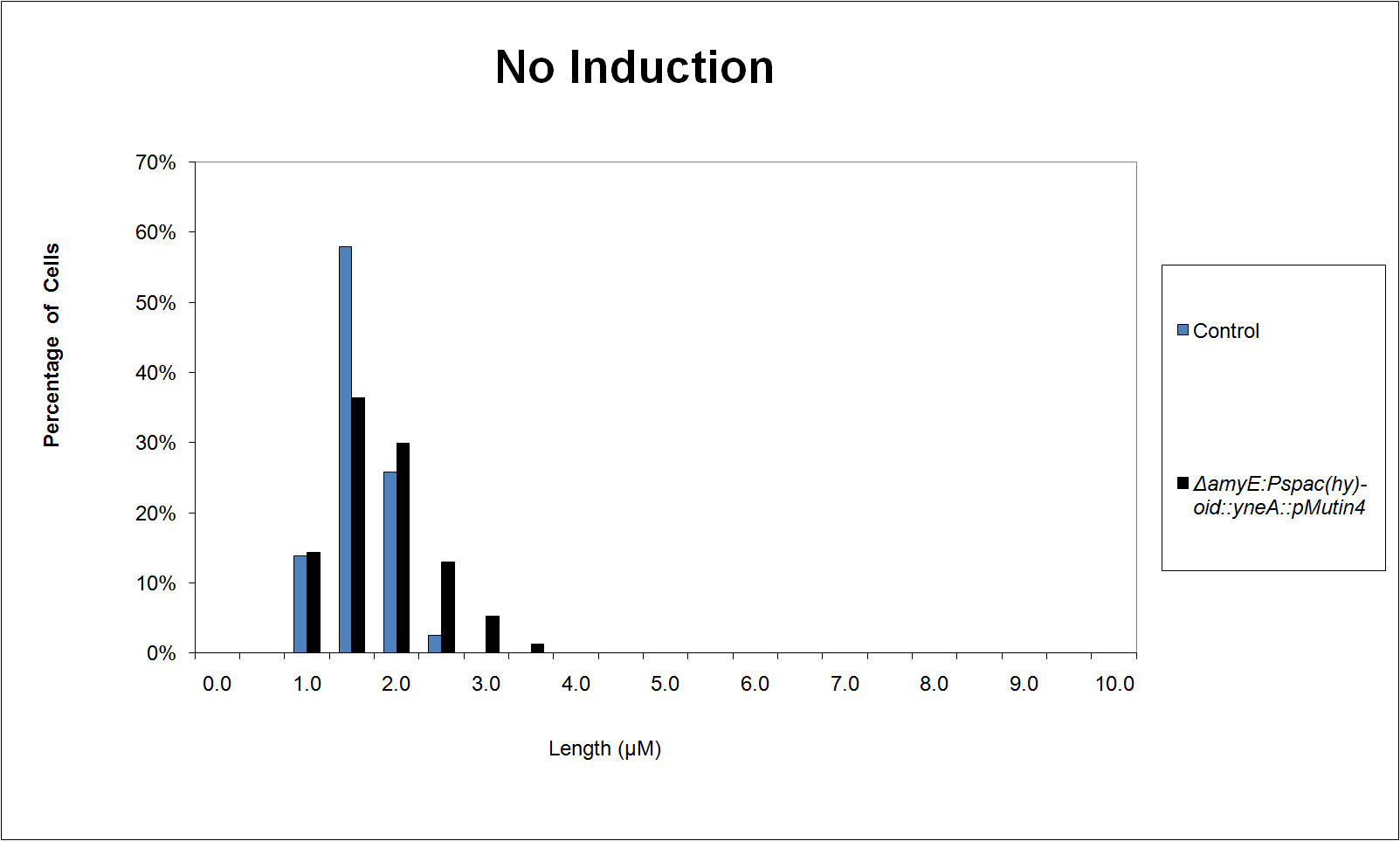
We characterised the part first without, and then with, LacI repression (using the integration vector pMutin4 to integrate lacI into the Bacillus subtilis 168 chromosome.
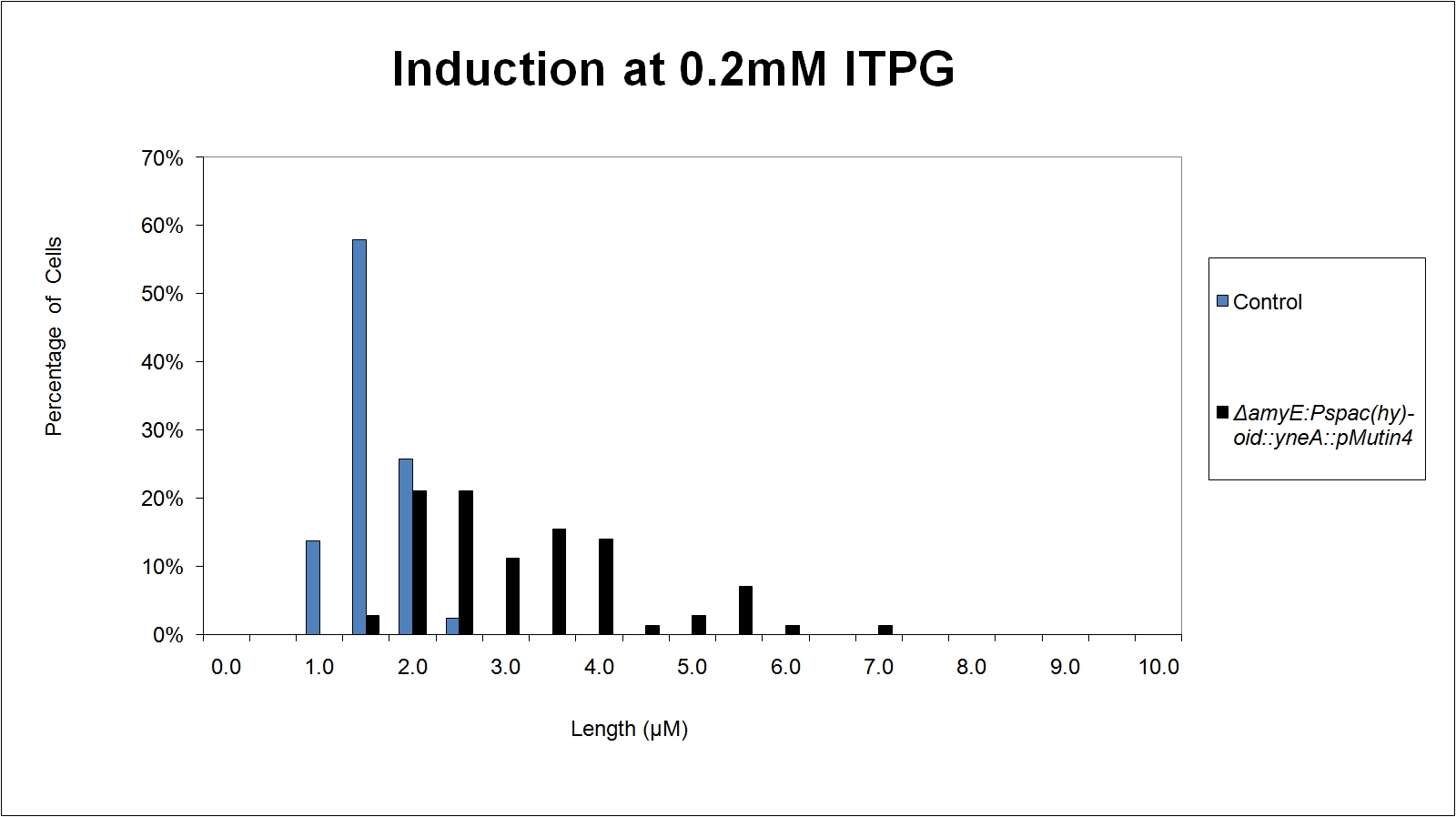
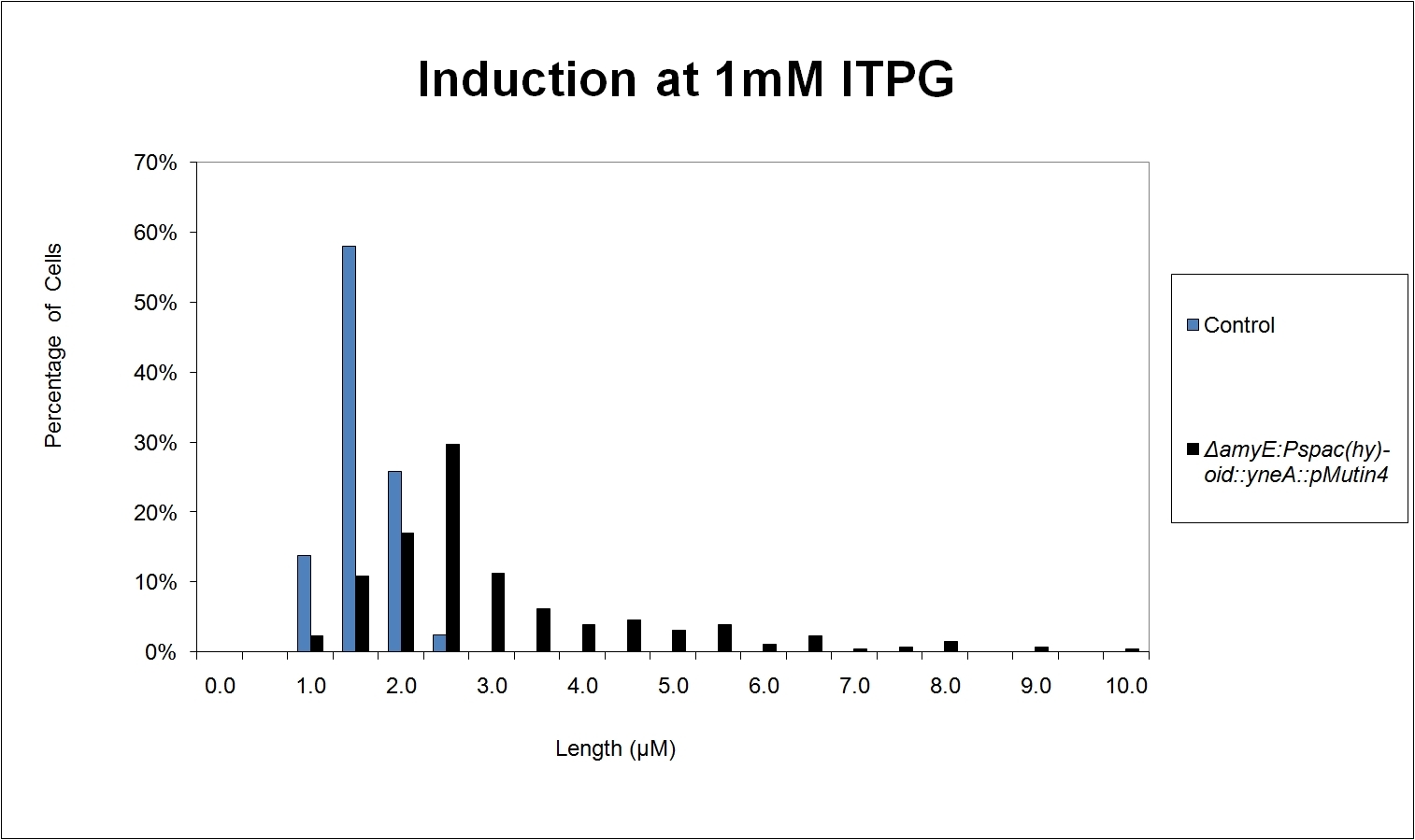
Graphs
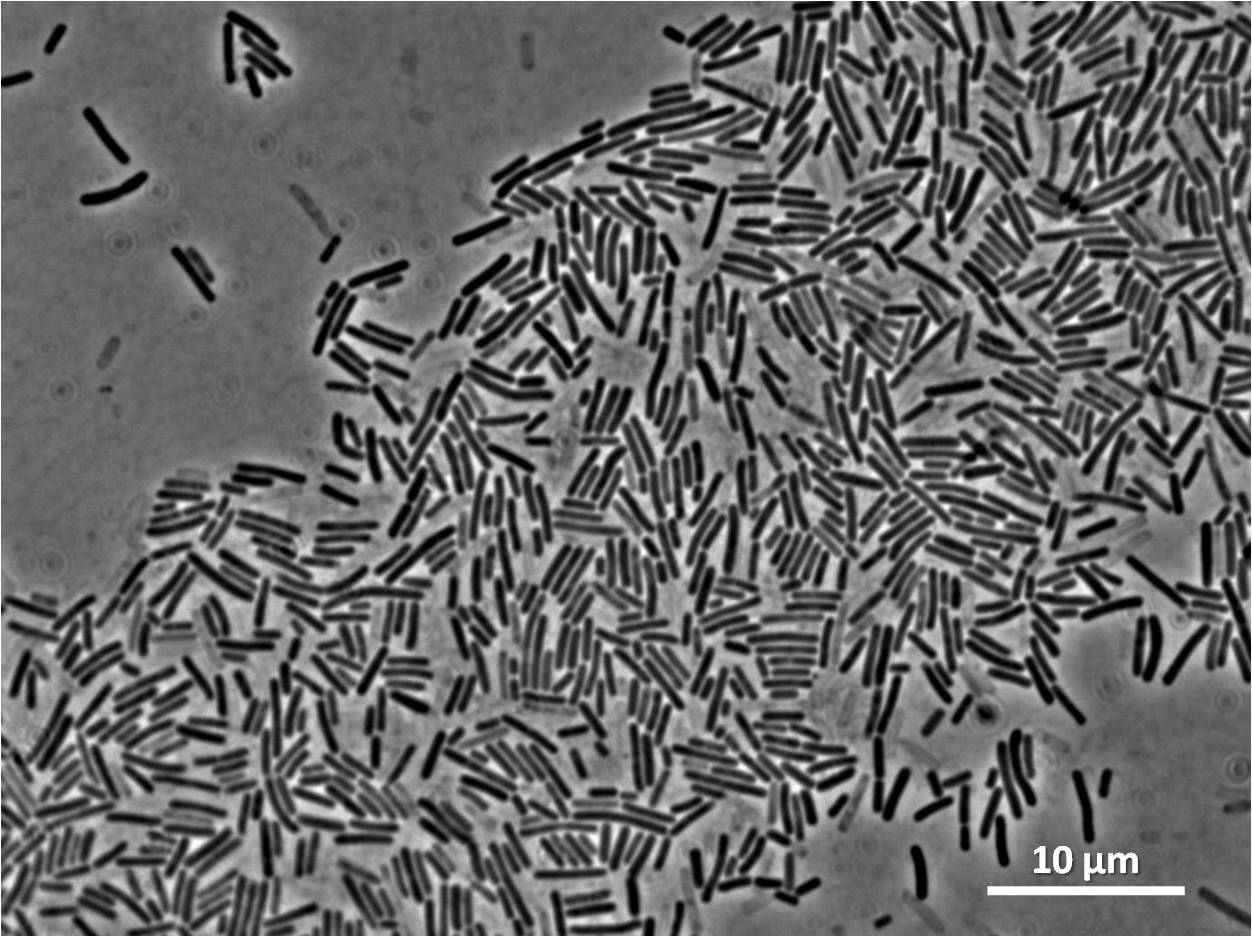
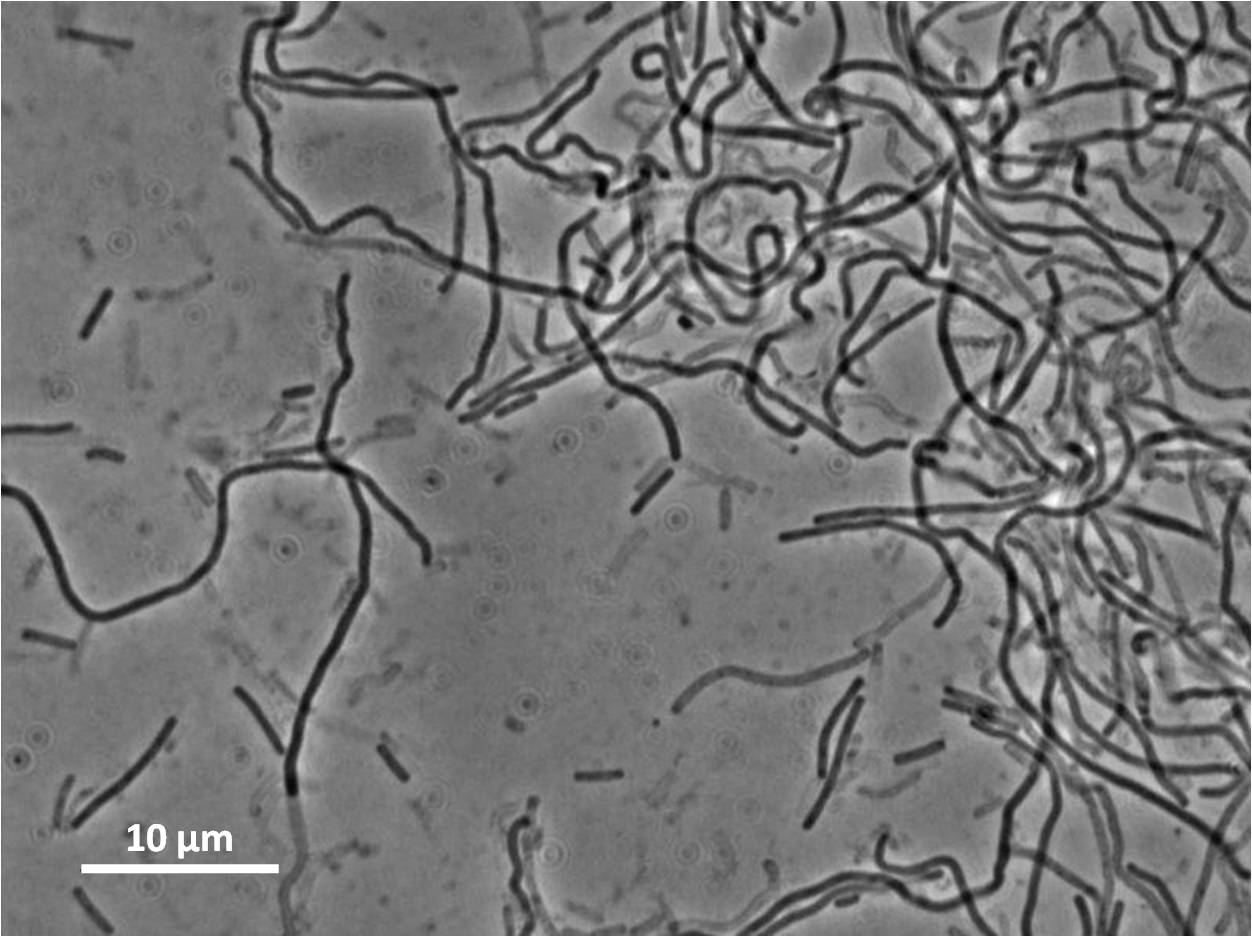
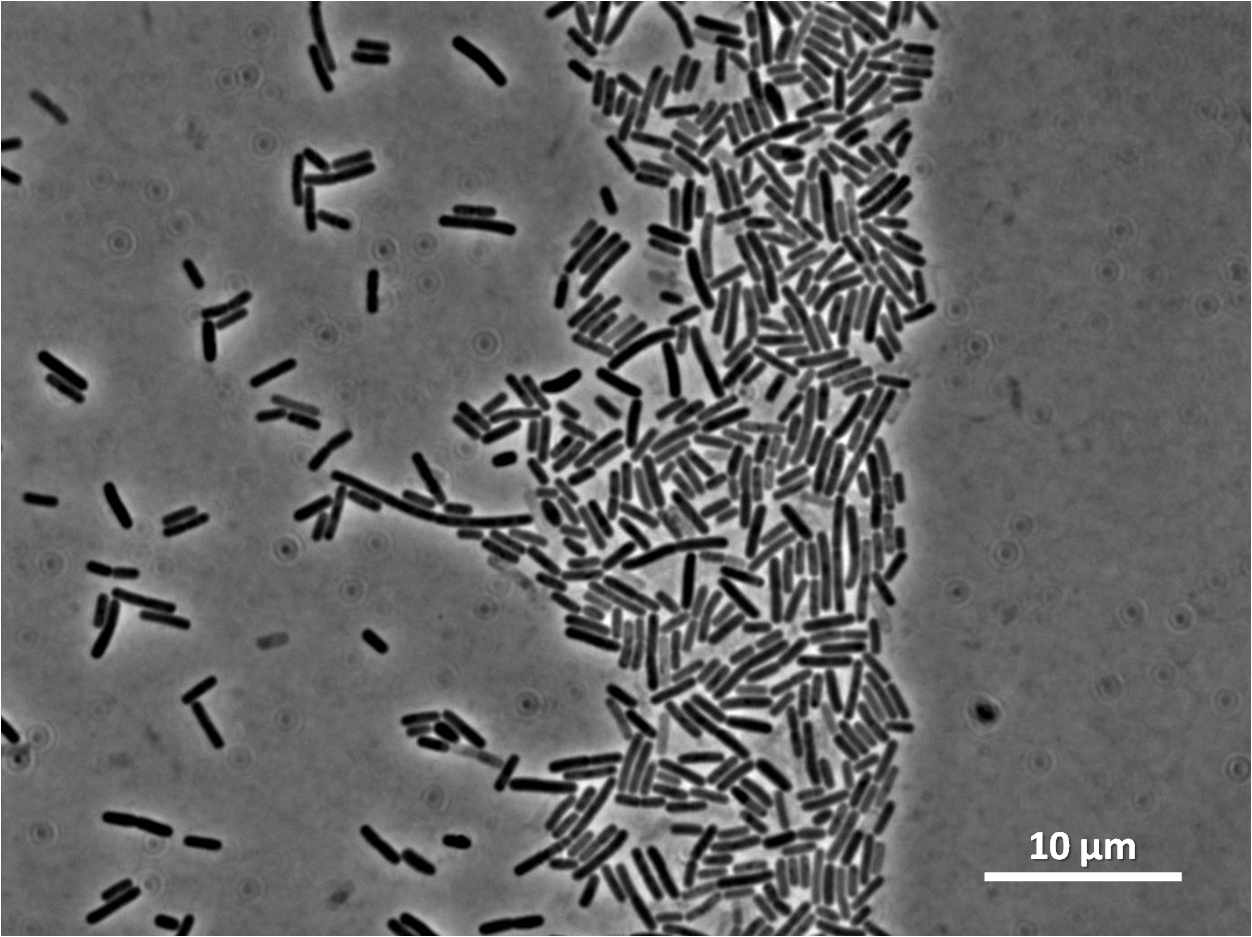
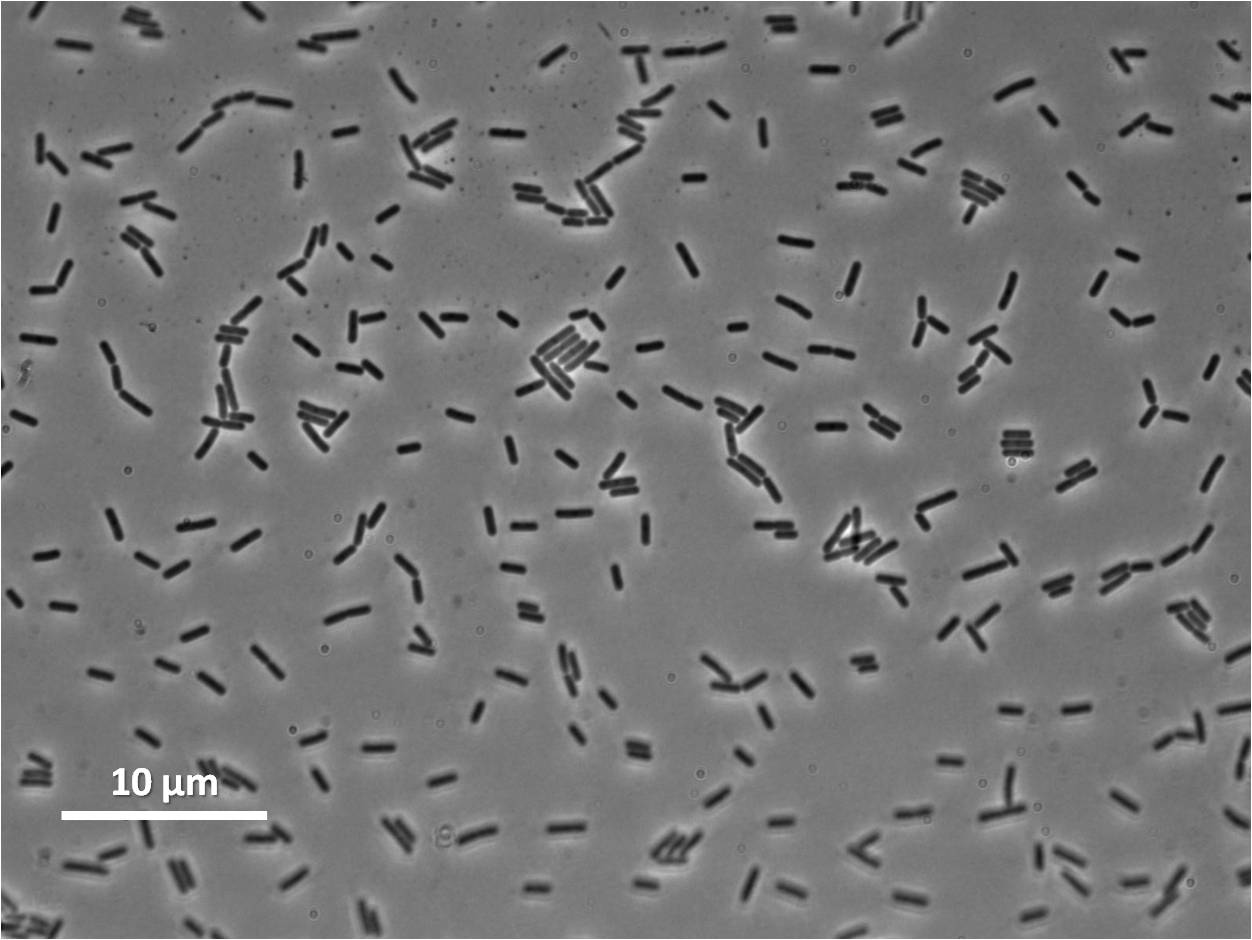
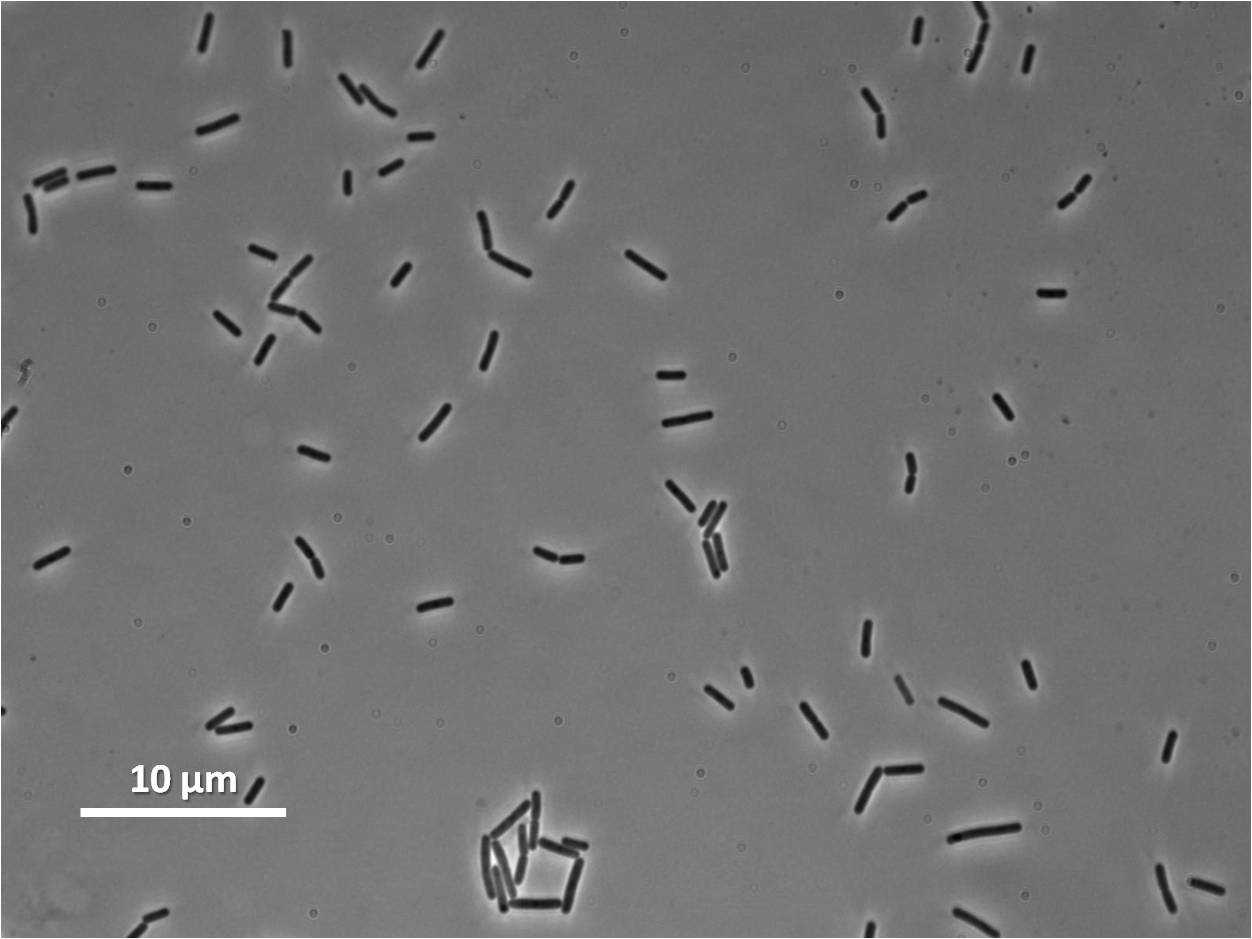
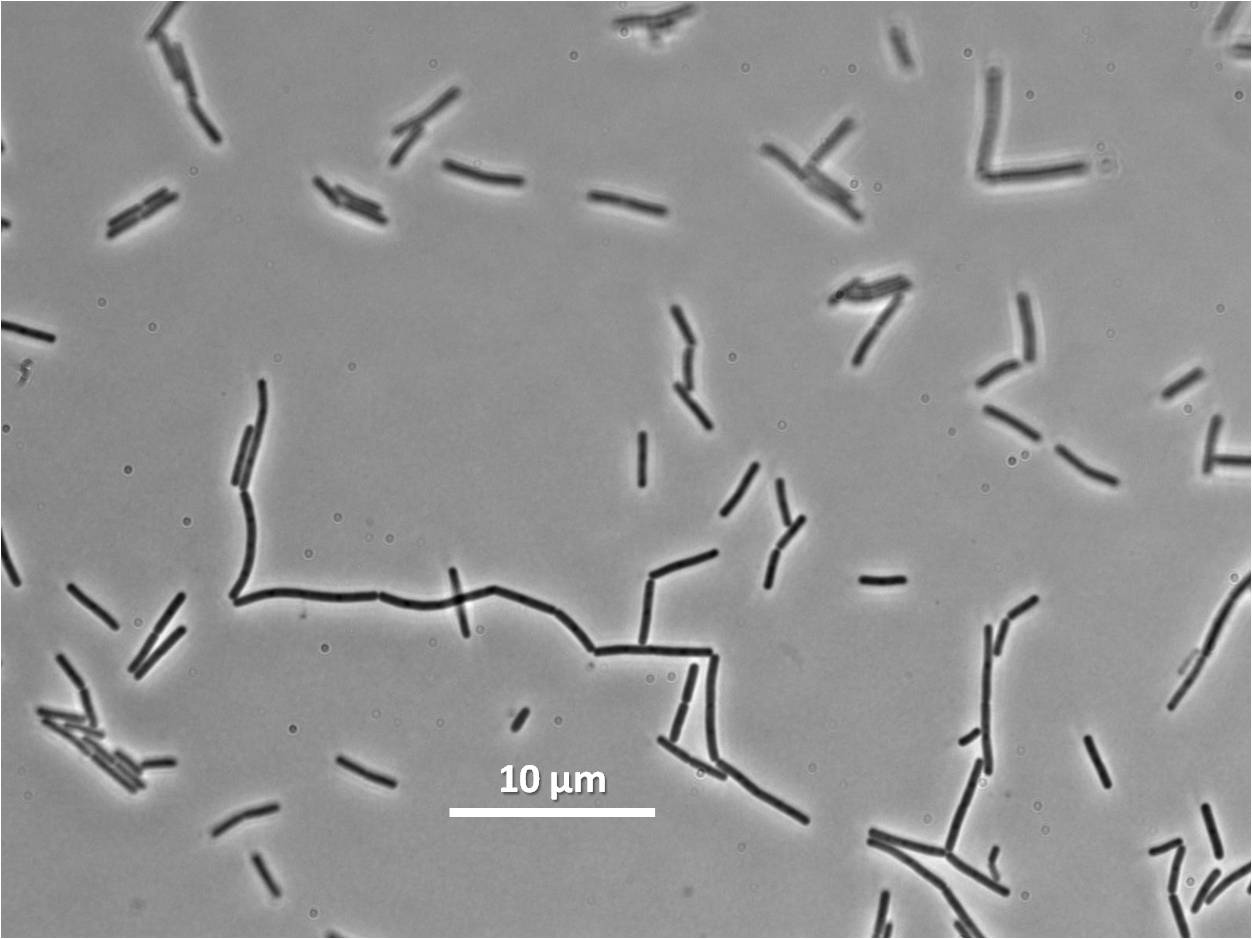
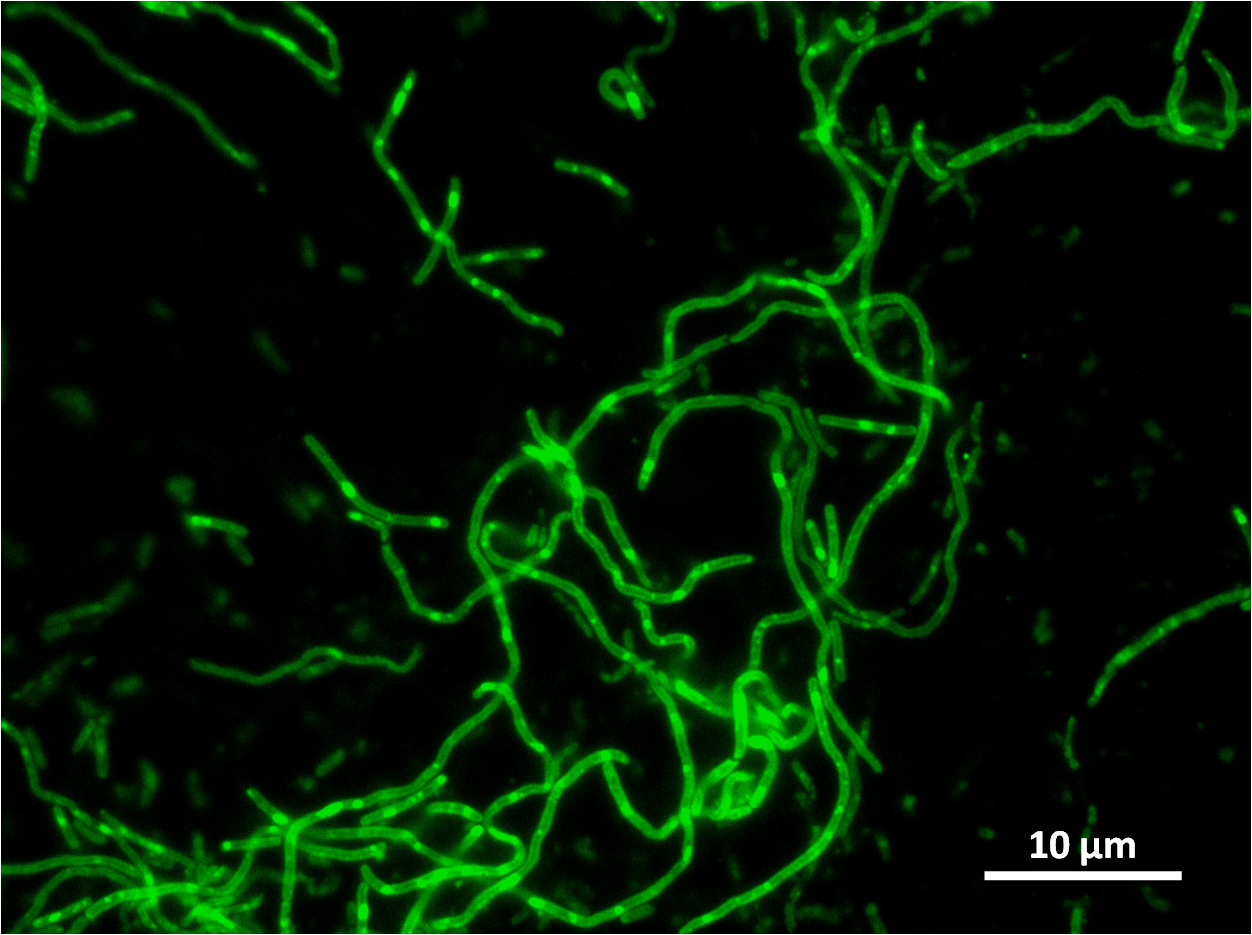
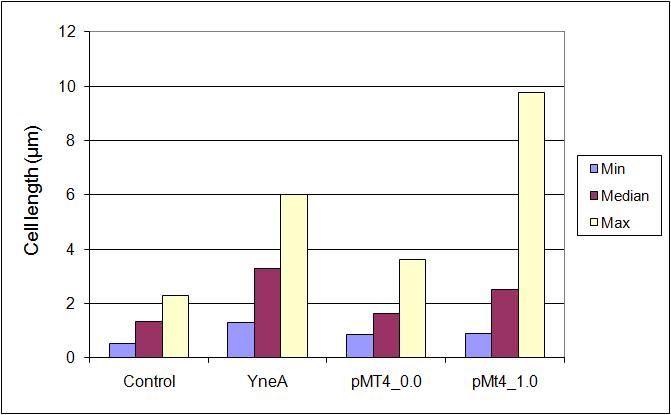
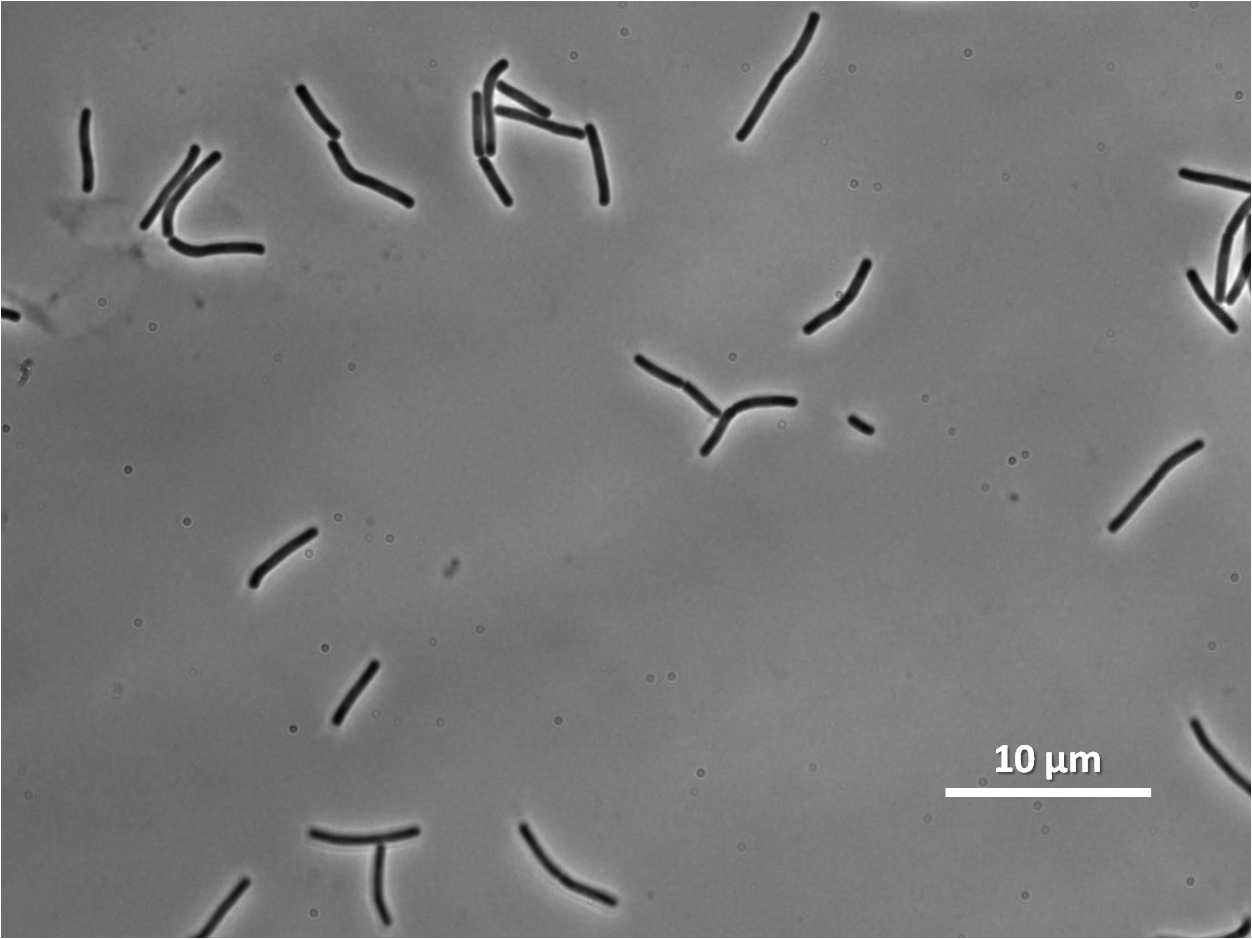
Graphs 2, 3 and 4 show a greater proportion of filamentous cells at a higher concentration of IPTG(1mM IPTG), compared with Bacillus subtilis 168 our control population.
Research
Bacillus subtilis in response to stress such as DNA damage stops the cells from dividing. This is a part of the SOS response initiated by the accumulation of single stranded DNA from DNA damage or stalled replication. Two proteins are vital for this response: RecA and LexA. RecA forms filaments on ssDNA and promotes the autocleavage of LexA. LexA usually represses the SOS operon. dinR is homologous to lexA in E. coli and is transcribed in the opposite direction of yneA.
SOS response is believe to be a universal bacteria phenomenon first studied in E.coli -lexA, recA
In Bacillus subtillis (gram positive) dinR protein is homologous to lexA (Repressor of din-damage inducible genes). din genes include uvrA, uvrB, dinB, dinC dinR and recA. DNA damage inhibits cell division.
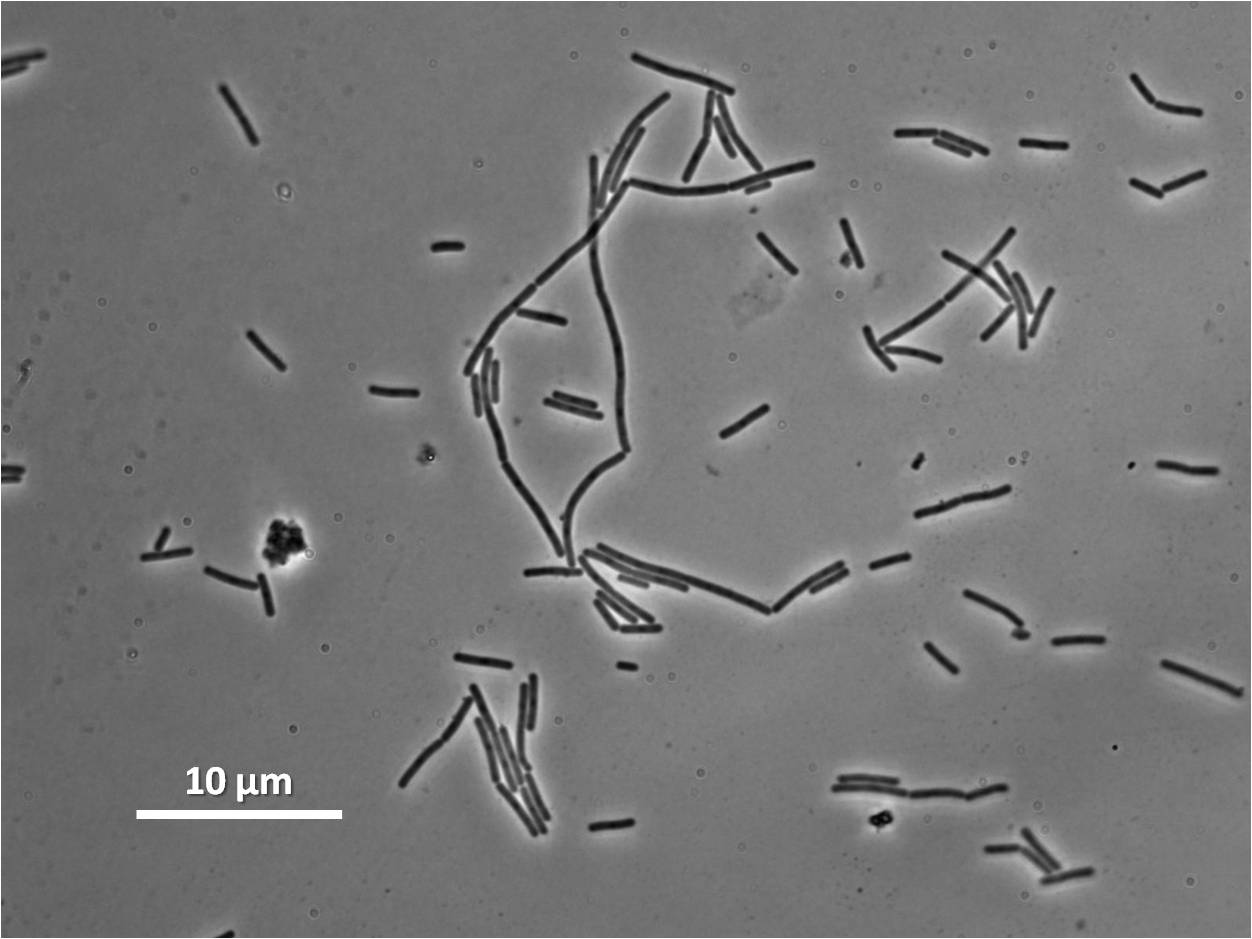
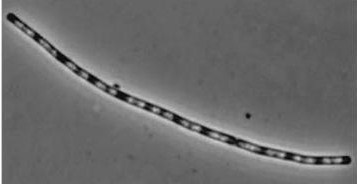
| Wild type Bacillus subtilis | dinRKO Mutant |
|---|---|
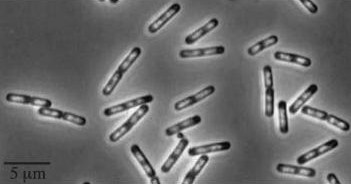
| 
|
Figure1: The images above show Bacillus subtilis Wild type and dinRKO mutant, and the change in cell length. dinR KO mutant over expresses the divergent (opposite direction) transcript for YneA, YneB and YnzC. These genes form the SOS regulon (recA independent SOS response).
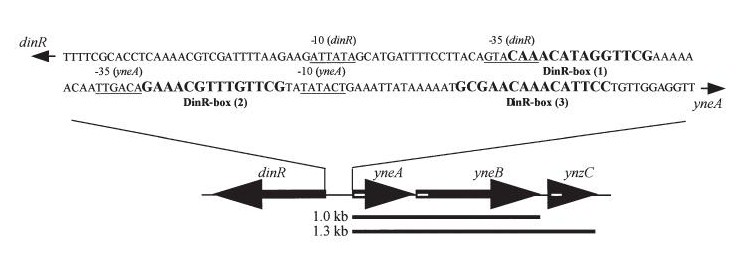
Figure2:The diagram above shows the Coding region for dinR and yneA showing divergent expression.
Expression of YneA from IPTG controlled promoter in wildtype leads to elongation. Disruption of YneA in SOS response leads to reduced elongation. Altering YneB and YnzC expression does not affect cell morphology.
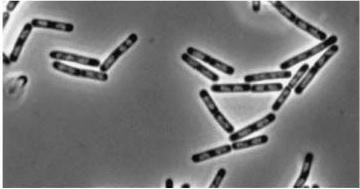
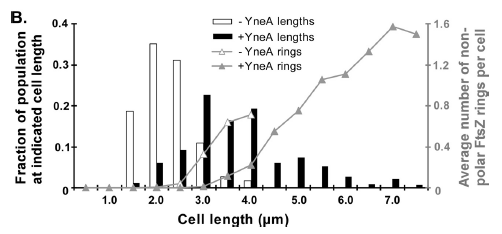
| 
|
Figure3(above left): Shows the double mutant dinR overexpression cancels out the filament formation via over expression of yneA.
Figure4(above right):This graph shows the correllation between reduced FtsZ ring formation, increased cell length and overexpression of yneA.
YneA protein required to suppress cell division and not chromosome replication or segregation.
FtsZ is important for bacterial cell division forming a ring structure at the division site by polymerising assembling other proteins necessary for division at the site.
FtsZ localises to the cell division cycle unless dinR is disrupted or YneA is being induced. YneA suppresses FtsZ ring formation which is proven by 2 hybrid protein association test.
YneA expression by the inactivation of dinR by RecA is important.
References
Kawai, Y., Moriya, S., & Ogasawara, N. (2003). "Identification of a protein, YneA, responsible for cell division suppression during the SOS response in Bacillus subtilis". Molecular microbiology, 47(4), 1113-22.
Mo, A.H. & Burkholder, W.F., (2010). "YneA , an SOS-Induced Inhibitor of Cell Division in Bacillus subtilis , Is Regulated Posttranslationally and Requires the Transmembrane Region for Activity" ᰔ †. Society, 192(12), 3159-3173.
 
|
 "
"