Team:Lethbridge/Results
From 2010.igem.org
Liszabruder (Talk | contribs) |
Liszabruder (Talk | contribs) |
||
| Line 128: | Line 128: | ||
<BLOCKQUOTE> | <BLOCKQUOTE> | ||
| - | =<font color="white"> | + | =<font color="white">Compartmentalization Parts= |
| - | + | ||
| - | + | ||
| + | One of the sub-projects for the bioremediation of the tailings ponds is to create synthetic <html><a href="https://2010.igem.org/Team:Lethbridge/Project/Compartamentalization"><font color="green"> microcompartments</font></a></html> that we can then use to isolate various pathway within an <i>Escherichia coli</i> cell. To do this we need to have a microcompartment as well as a means to characterize the compartment so that the system can be optimized. Here are the experiments we have performed so far towards the characterization of the microcomparments. | ||
| + | <br><br> | ||
| + | ==<font color="white">Placement of Oligoarginine Tail on Proteins</font>== | ||
| + | <br> | ||
| + | ===<font color="white">Characterized Parts</font>=== | ||
| + | <hr> | ||
<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K249004" target="new"><font color="green" size="+1">BBa_K249004</font></a></html> | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K249004" target="new"><font color="green" size="+1">BBa_K249004</font></a></html> | ||
<br><br> | <br><br> | ||
| Line 139: | Line 143: | ||
<br><br> | <br><br> | ||
<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331031"target="new"><font color="green" size="+1">BBa_K331031</font></a></html> | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331031"target="new"><font color="green" size="+1">BBa_K331031</font></a></html> | ||
| - | + | <br><br> | |
| - | ==<font color="white">Hypothesis</font>== | + | ===<font color="white">Hypothesis</font>=== |
| - | + | <hr> | |
The placement of an oligoarginine sequence at the N-terminus of a protein will destabilize the protein in vivo. | The placement of an oligoarginine sequence at the N-terminus of a protein will destabilize the protein in vivo. | ||
| - | + | <br><br> | |
| - | ==<font color="white">Introduction</font>== | + | ===<font color="white">Introduction</font>=== |
| - | + | <hr> | |
The long term goal of our team is to utilize an oligoarginine tail to specifically target enzymes into amicrocompartment composed of modified lumazine synthase subunits. While conducting background research on the project, we came upon data originally reported by Bachmair <i>et al.</i><sup>1</sup> suggesting that the identity of the amino acid at the N-terminus of a protein is related to its half-life, and mostly notably, that arginine residues at the are destabilizing. This data suggests that by placing an arginine at the N-terminus of a protein to be targeted into a <html><a href="https://2010.igem.org/Team:Lethbridge/Project/Compartamentalization"><font color="green"> lumazine synthase microcompartment</font></a></html> would cause degradation of our protein before it can be moved into the microcompartment. | The long term goal of our team is to utilize an oligoarginine tail to specifically target enzymes into amicrocompartment composed of modified lumazine synthase subunits. While conducting background research on the project, we came upon data originally reported by Bachmair <i>et al.</i><sup>1</sup> suggesting that the identity of the amino acid at the N-terminus of a protein is related to its half-life, and mostly notably, that arginine residues at the are destabilizing. This data suggests that by placing an arginine at the N-terminus of a protein to be targeted into a <html><a href="https://2010.igem.org/Team:Lethbridge/Project/Compartamentalization"><font color="green"> lumazine synthase microcompartment</font></a></html> would cause degradation of our protein before it can be moved into the microcompartment. | ||
<br><br> | <br><br> | ||
We chose to investigate the how the placement of an oligoarginine sequence affects the stability of the protein to which it is fused. | We chose to investigate the how the placement of an oligoarginine sequence affects the stability of the protein to which it is fused. | ||
| - | + | <br><br> | |
| - | ==<font color="white">Method</font>== | + | ===<font color="white">Method</font>=== |
| - | + | <hr> | |
In order to further characterize the C-terminal and N-terminal oligoarginine tag (BioBricks <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K249005" target="new"><font color="green">BBa_K249005</font></a></html> and <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K249004" target="new"><font color="green">BBa_K249004</font></a></html> respectively) and investigate the effect their placement on protein stability, yellow fluorescent proteins (YFP) with the oligoarginine fused to either the C-terminus (<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331023" target="new"><font color="green">BBa_K331023</font></a></html>) or N-terminus (<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331022" target="new"><font color="green">BBa_K331022</font></a></html>) (and preceded by a ribosomal binding site – <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_B0034" target="new"><font color="green">B0034</font></a></html>) were synthesized. We used our <html><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Protocols#Assembly_of_BioBricks_using_the_Red.2FWhite_3-Antibiotic_Assembly_Method"><font color="green"> Red/White 3-Antibiotic assembly method</font></a></html> to add a tetracycline repressible promoter (<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_R0010" target="new"><font color="green">BBa_R0010</font></a></html>) for constitutive expression of the fusion protein. This addition generated BioBricks <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331031" target="new"><font color="green">BBa_K331031</font></a></html> and <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331030" target="new"><font color="green">BBa_K331030</font></a></html> for the C-terminal tagged and N-terminal tagged YFP respectively. | In order to further characterize the C-terminal and N-terminal oligoarginine tag (BioBricks <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K249005" target="new"><font color="green">BBa_K249005</font></a></html> and <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K249004" target="new"><font color="green">BBa_K249004</font></a></html> respectively) and investigate the effect their placement on protein stability, yellow fluorescent proteins (YFP) with the oligoarginine fused to either the C-terminus (<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331023" target="new"><font color="green">BBa_K331023</font></a></html>) or N-terminus (<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331022" target="new"><font color="green">BBa_K331022</font></a></html>) (and preceded by a ribosomal binding site – <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_B0034" target="new"><font color="green">B0034</font></a></html>) were synthesized. We used our <html><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Protocols#Assembly_of_BioBricks_using_the_Red.2FWhite_3-Antibiotic_Assembly_Method"><font color="green"> Red/White 3-Antibiotic assembly method</font></a></html> to add a tetracycline repressible promoter (<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_R0010" target="new"><font color="green">BBa_R0010</font></a></html>) for constitutive expression of the fusion protein. This addition generated BioBricks <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331031" target="new"><font color="green">BBa_K331031</font></a></html> and <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331030" target="new"><font color="green">BBa_K331030</font></a></html> for the C-terminal tagged and N-terminal tagged YFP respectively. | ||
<br><br> | <br><br> | ||
| Line 157: | Line 161: | ||
<br><br> | <br><br> | ||
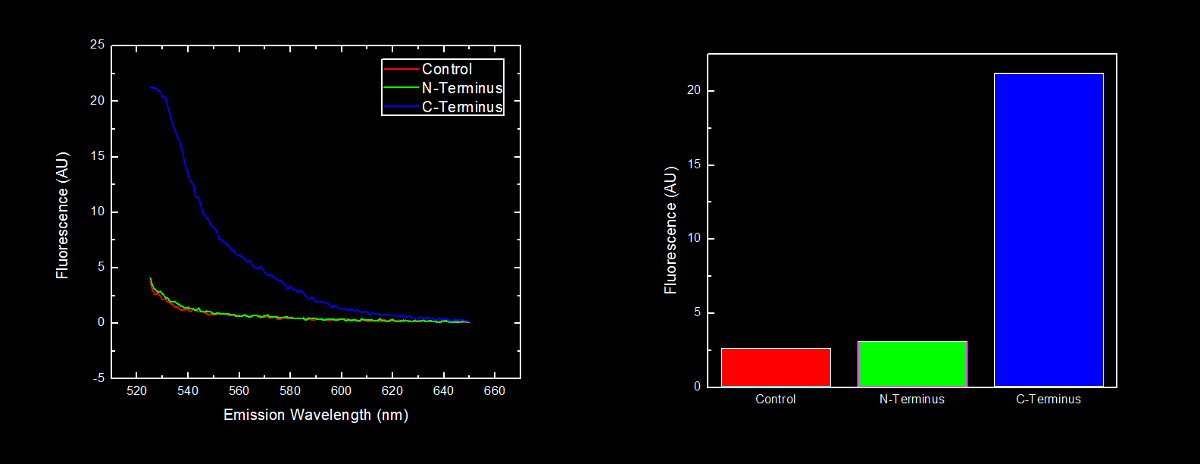
This dilution of cells was excited at 517 nm, and the emission spectra was read from 522 nm to 650 nm. Fluorescence at 524 nm (emission maxima of YFP) of control cells (<i>Escherichia coli</i> DH5α), N-terminal tagged, and C-terminal tagged YFP were compared. | This dilution of cells was excited at 517 nm, and the emission spectra was read from 522 nm to 650 nm. Fluorescence at 524 nm (emission maxima of YFP) of control cells (<i>Escherichia coli</i> DH5α), N-terminal tagged, and C-terminal tagged YFP were compared. | ||
| - | + | <br><br> | |
| - | ==<font color="white">Results</font>== | + | ===<font color="white">Results</font>=== |
| - | + | <hr> | |
N-terminal tagged YFP did not have substantially more fluorescence than control cells. Cells expressing C-terminal tagged YFP had ten times more fluorescence than control cells and cells expressing N-terminal tagged YFP. | N-terminal tagged YFP did not have substantially more fluorescence than control cells. Cells expressing C-terminal tagged YFP had ten times more fluorescence than control cells and cells expressing N-terminal tagged YFP. | ||
[[image:Lethbridge_NvsC-terminalOligoArgBlackfINAL.png|900px]] | [[image:Lethbridge_NvsC-terminalOligoArgBlackfINAL.png|900px]] | ||
| - | + | <br><br> | |
| - | ==<font color="white">Conclusion</font>== | + | ===<font color="white">Conclusion</font>=== |
| - | + | <hr> | |
Our results are consistent with the data reported by Bachmair <i>et al.</i> in that the placement of arginine residues at the N-terminus of our YFP results in no observable fluorescence over control cells. Assuming that transcription of this K331030 and K331031 are equivalent, these data suggest that the N-terminal oligoarginine is reducing the half-life of the protein to which it is fused, ie YFP. | Our results are consistent with the data reported by Bachmair <i>et al.</i> in that the placement of arginine residues at the N-terminus of our YFP results in no observable fluorescence over control cells. Assuming that transcription of this K331030 and K331031 are equivalent, these data suggest that the N-terminal oligoarginine is reducing the half-life of the protein to which it is fused, ie YFP. | ||
| - | + | <br><br> | |
| - | ==<font color="white">Reference</font>== | + | ===<font color="white">Reference</font>=== |
| - | + | <hr> | |
<sup>1</sup>Bachmair A., Finley D., Varshavsky A. <b>In Vivo Half-Life of a Protein Is a Function of Its Amino-Terminal Residue.</b> <i>Science</i> 234. <b>4773</b> 179-186. | <sup>1</sup>Bachmair A., Finley D., Varshavsky A. <b>In Vivo Half-Life of a Protein Is a Function of Its Amino-Terminal Residue.</b> <i>Science</i> 234. <b>4773</b> 179-186. | ||
| - | |||
<br> | <br> | ||
<br> | <br> | ||
 "
"