|
|
Experiment Design
We designed different experimental set-ups with varying complexity to test RNA signal/switch pairs based on our concept.
Our initial idea to prove our concept of antitermination was to use fluorescent proteins as reporters. This approach gives the opportunity to measure the termination and antitermination efficiency of our designed BioBricks in vivo as well as in vitro, the latter using a translation kit based on E. coli lysate. Later on, we decided to develop an experiment, that relies only on transcription. In this set-up, we used a fluorescent dye, malachite green, that binds a specific RNA aptamer and thus makes it possible to detect transcription activity.
In vivo Measurements
In vivo measurements have the highest complexity compared to any other experimental set-up. Our system has to deal with several circumstances a cellular environment comes with, such as interaction with other RNAs, degradation by RNases or unspecific interactions. Nevertheless, the measurements are essential, as our switches should finally work inside cells to fulfill our vision of an intracellular logic network.
Read more
Design
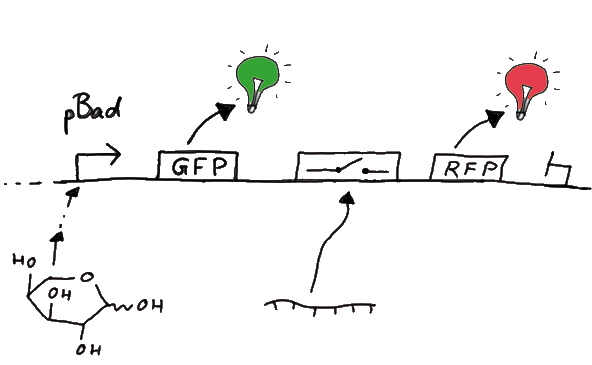
For the measurements in vivo we decided to use an expression cassette consisting of Green Fluorescent Protein (GFP) coding sequence upstream of the switch and another fluorescent protein coding sequence downstream of it. Both protein coding sequence carry the same ribosome binding site, therefore, the GFP fluorescence can be used as internal control in measurements. Since the spectra should not overlap and to avoid FRET as well as an pure overlap of the spectra, we settled on the usage of red fluorescent protein variants, namely mRFP1 in the first try. While the GFP fluorescence is used to normalize the measurements, the RFP fluorescence is used to detect termination/antitermination.
Upon binding of the signal, the stem loop of the switch would resolve leading to red fluorescence. The GFP fluorescence as internal control carries the advantage that errors in the measurement set can be detected easily. Lack of arabinose or promoter insensitivity can be recognized as well as problems with the fluorescence measurement itself. Plus, we have a way to normalize our measurements and compare different preparations in relation to each other.
Construction and Cloning
Our measuring plasmid is based on the BioBrick pSB1A10, A1, distribution 2010. Unfortunately after two months of cloning we had to recognize that the plasmid in use did not work (see also Biobrick validation--> link). So after the first unsuccessful attempts we decided to reclone the system, substituing RFP to mCherry, a dsRED derivative with a spectrum in the far red, and adding arabinose inducible promoters in front of both fluorescent proteins.
To control the expression of the switch, the particular DNA sequence itself is under the control of an IPTG dependent promoter. In the future we want our networks to be able to respond to a variety of external signals like small metabolites, ions or whatever can be found in the parts registry. For a start we went with an established and well-working system like the lac-operon. ---> PLASMID MAP
So upon induction with arabinose a rise of GFP expression can be seen. To monitor changes in gene expression we put our E. coli cells in a fluorimeter and measured fluorescent. It worked quite nicely with living cells in a fluorimeter, the only thing to avoid is too much scattering: the cell density should not exceed an OD600 of ???. Continious stirring and a set temparature at 37°C allowed measuring over severall hours. Cell density was checked in between.
Measurement
For switch evaluation, IPTG was added to the cells after about two hours after arabinose induction (baseline). ??? Stimmt das?? A rise of RFP/mCherry emission should be visible in case of a working switch.
For evaluation of the measuring plasmid itself we incorporated a positive control in every measurement. A random sequence in between GFP and RFP/mCherry was chosen in a corresponding length. An increase in both GFP and mCherry was detectable in the positive control and in the same amount after quantum yield correction, proving that the measuring plasmid is working nicely.
As a negative control we measured the same plasmids as for every switch but without signal. Since our switches are effective terminators if no signal is bound, transcription can not occur and no RFP/mCherry is produced.
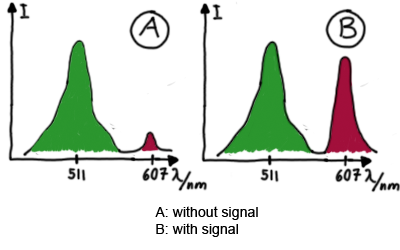
When measuring the termination of our BioBricks and the antitermination by their corresponding signal-RNA, we should be able to observe an increasing RFP emission compared to the GFP emission upon induced signal-RNA production in the cells/in the kit:
With these measurements, it should also be possible to observe differences in efficiency of termination as well as antitermination between our designed switches.
Close
In vitro Translation
In vitro measurements with E. coli lysate make the fluorescence signals independent of cell growth and physical or biological factors, e.g. cell density or growth stadium.
Read more
Design
In this assay we used the same constructs as engineered for the in vivo studies.
Measurements
We used the cell-free E. coli S30 extract system for circular DNA provided by promega[1], which is prepared by modifications of the Method Zubay et al.[2] described. The characterization of the kit can be obtained from the [http://partsregistry.org/Cell-free_chassis/Commercial_E._coli_S30 Parts Registry].
We perform our experiments at 37°C with an amount of approxim. 1 ug plasmid in a reaction volume of 50 uL. Measurements are carried out in a jasco fluorolog, where the fluorescent proteins RFP and GFP are excited and fluorescence is detected, so measurements are done more or less the same as in our in vivo studies. A general problem occuring in the experiments was the low capacity of the kit. The signal intensity is very low, which made it difficult to observe any signal intensity alterations.
Close
In vitro Transcription
An experiment, in which we detect In vitro transcription, offers an elegant way for a fast and easy prove of principle, since our switch is RNA-based and the whole mechanism takes place on trancriptional level. Most side effects occuring in a complex environment given in a cell or a cell lysate do not arise here.
Read more
If measureable effects with our basic concept can be seen in vitro we can use the so gained data to optimize the system in vivo. Since we are working on a totally new principle of trancriptional control, we used this approach for easy variation of different variables like the length of the core unit and the switch to signal ratio.
To study the switches on the transcriptional level gives the advantage, that we would have less interferences and possible artefacts. Also, we are not sure how cellular mechanisms like degradation of RNases or interacting factors as well as molecular crowding influence our systems.
Working with in vitro systems also has the advantage that an input is not needed anymore and the output can also be generated easily. We used two readouts with two different transcription systems to check and investigate our devices: First, we used an malachitegreen-binding aptamer for an fluorescent output (which will be described in detail later) and second, we simply put our reaction educts on an denaturing acrylamide-gel to check for RNA varying in length. As for two different transcription systems we used on the one hand E. coli-RNA Polymerase (RPO) based transcription since the aim is to apply our system in vivo and on the other hand T7 based transcription which is well established through literature.
T7 RNA polymerase
The T7 RNA polymerase is known for satisfying RNA yields together with easy handling. In our approach we had PCR amplified, double stranded switches with an malachitegreen binding aptamer following after the switch (133 bp, see section below) and a single stranded signal with about 30 bp length.
For in vitro expression the T7 RNA Polymerase requires a double stranded promotor region at the beginning of the DNA template but is otherwise capable of handling single stranded DNA, so a sense strain corresponding to the T7 promoter region was added. Transcription is more effective with double stranded DNA as template. Since we ordered the signal sequences we tested we chose the cheaper way in the beginning by using single stranded signals with corresponding sense T7 pieces and switched to double stranded constructs after narrowing down the most promising switch/signal pairs.
E. coli RNA polymerase
In comparison to the T7 RNA Polymerase the E. coli RNA Polymerase requires slightly more sophisticated proceedings when it comes to the design of switches and handling of the enzyme. The biggest in our case was to store it properly since the only -80°C fridge was in another building.
E. coli RPO was ordered saturated with σ70-factor. The switch consists of an ???-promoter, the switch itself and a ???.
Denaturing Polyacrylamide gel electrophoresis
We also used Polyacrylamide gel electrophoresis (PAGE) for evaluation of termination efficiency of our basic units. Gels containing 15 % acrylamide and 6 M urea were used for separation of 90 (terminated by switch) and 133 bp (continous reading) RNAs.
Polyacrylamide gels seperate RNA and DNA according to their size in an electric field. Since the negative charge equals the size of nucelotides in the RNA/DNA, the number of base pairs can be compared between two samples often with one base pair resolution. Since RNA forms three-dimensional structures, the samples are preheated and run in 6 M urea. The polyacrylamide gel is stained in SybrGold afterwards which binds to both single and double stranded DNA and RNA.
Denaturing PAGE is a simple yet elegant way to check for transcription efficiency and termination rates. Since it is a very direct way and it provides a simple yet clear readout, we used it as another method beside the more sophisticated malachitegreen binding assay to evaluate and characterize our switch.
Malachite green assay
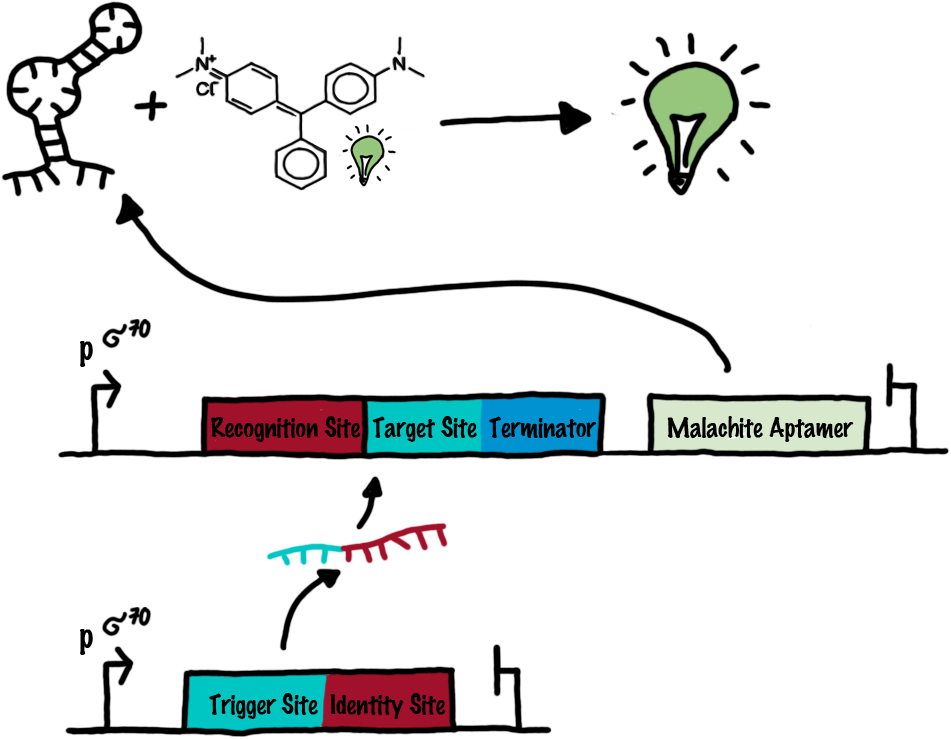
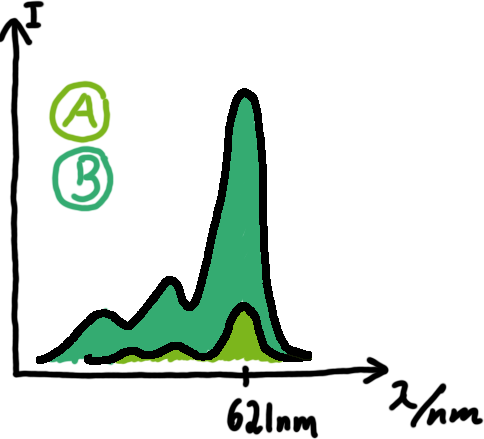 Emission spectra of malachite green; A: without signal-RNA, B: with signal-RNA Malachite-green is a dye with a negligible fluorescence in solution but undergoes a dramatic increase if bound by a RNA -aptamer. Upon binding to the aptamer, the fluorescence of malachite-green increases about 3000 times making it an exceptionel good marker. Since the binding is very specific, transcription in dependence of a signal can be monitored by measuring the fluorescence of malachite-green over time if the aptamer is located behind the switch. Transcription of the aptamer will only take place after anti-termination by a signal. An increase should be visible over time.
For the T7-based measurements we ordered single stranded signals for a first attempt and added matching single strands complementary to the T7 promoter region. The switch was amplified using PCR and consisted of the following elements: Primer-binding site - T7 promotor - switch - malachitegreen binding aptamer. Upon binding of a correct signal to the switch, the stem loop dissolves and transcription is possible.
OLD: A second possibility to measure parameters of our switches we came up with, was the idea to investigate our system on the transcriptional level only. Therefore, we decided to use malachite green as reporter. Malachite green in a fluorescent dye, whose emission increasing dramaticly (about 3000 times) upon binding of a specific RNA-aptamer. The RNA-aptamer
---concept to be described, as well as literature---
<ref>refs</ref>
We made constructs comprising of a sigma(70)-binding promoter followed by a short nonsense sequence, the switches and the aptamer sequence.
Also we made constructs, where the transcription of the signal-RNA is under the control of a sigma(70) promoter. These two linear DNA-constructs, together with the e.coli RNA-polymerase and the right buffer conditions should represent an easy-to-handle measurement kit on the transcriptional level.
Close
References
[1] http://www.promega.com/catalog/catalogproducts.aspx?categoryname=productleaf_335&ckt=1
[2] Zubay, G. (1980) Meth. Enzymol. 65, 856–77, Zubay, G. (1973) Ann. Rev. Genet. 7, 267–87.
Experimental Results
In vivo Measurements
In vivo
In vitro Measurements
Read more
Protocols
Molecular Biology
PCR
Read more
Digestion
Read more
Ligation
In vivo Measurement
Read more
So haben wir in vivo gemessen
Close
In vitro Translation
Read more
So haben wir in vitro gemessen
Close
In vitro Transcription
Read more
So haben wir in vivo gemessen
Close
Lab Book
Explanations
In the following we present an overview regarding our work in the lab. For easier understanding we summarized the work of each week using colored boxes. To get more information about the work and results of a specific week, just click on the according week number. To get a better overview we used the following color code for the boxes:
| This box represents general cloning that were requirde for several measurements. See the protocol section for further details.
|
| The blue box indicates in vivo measurements which are described here.
|
| The green box indicates in vitro measurements relying on in vitro transcription and malachite green measurements. Details can be found here.
|
| The yellow box represents measurements done with an in vitro translation kit and is described in more details here.
|
Chronological Lab Book
Week01
Constrcuts for in vivo measurments
08.04.2010
Flo & Philipp
PCR
- samples:
- R0011_His
- R0011_Trp
- Control
- protocol: protocols
- templates: purified PCR products from 5.2.2010
- primer G1004/1005
- polymerase: Taq
- programm: igempcr
Purification of PCR products with QIAquick PCR purification Kit
- protocol followed. exceptions: DNA-binding/unbinding with 3min 6000rpm followed by 60sec full speed
2% Agarose Gel
09.04.2010
Philipp & Flo
Gel of PCR products from 08.04.2010
- loaded: 10 uL sample+2 uL 6x GLD, 4/2 uL LMW standard
- 110 V, 90min
- stained with Sybrgold, 20min, 1:10.000 dilution in TAE
- Standard - Control - R0011_His - R0011_Trp - Standard(=low molecular weight (see Protocols#Lab_Protocols))
File:100409.JPG
Close
Week02
Improved Constructs... Testing
15.04.2010
Philipp & Flo
[http://web.e14.physik.tu-muenchen.de/igem/index.php/Protocols#PCR PCR] of B0014 and R0011
16.04.2010
Philipp & Flo
- Concentrations measured with nanodrop:
| B0014
| 2.5 ng/uL
|
R0011
| 27.5 ng/uL
|
--> worked for R0011, not for B0014
- PCR of B0014
- Purification with the Zymo Kit, Elution in 20 uL H2O
- Concentration measured with nanodrop, 17.5 ng/uL --> worked
template
| restriction enzymes (biobrick assembly)
|
B0014 (from Christoph, verified PCR products, 21 ng/uL)
| EcoRI, PstI
|
R0011 (from PCR [15042010], 27.5 ng/uL
| SpeI
|
HisSig (1:100 dilution)
| XbaI
|
TrpSig (1:100 dilution)
| XbaI
|
| psB1K3 (with RFP insert, from HiWiPhilipp, 81 ng/uL)
| EcoRI, PstI
|
5 uL template used for each setup. protocol followed
- Gel for purification of the cleaved plasmid
- 2% Agarose in 1x TAE
- 120 V, 90 min
- stained with SybrGold
- digestion, digestion, 1 kb ladder
- Digestion worked (partly). band at 2000 bp (backbone) cut
File:100416.png
- Purification of DNA from Gel
- Ligation of HisSig/TrpSig with R0011in 2 reactions
| used Volume
| approx. concentration*
|
HisSig
| 6 uL
| 7 ng/uL
|
TrpSig
| 6 uL
| 5 ng/uL
|
R0011
| 3 uL
| 6 ng/uL
|
* approximated from the amount used in the digestion before
Close
Week03
Measurements
19.04.2010
- PCR of R0011-TrpSig and R0011-HisSig
- Purification with the Zymo Kit, Elution in 30 uL H2O
- Concentration measured with nanodrop: c(R0011-TrpSig)=20 ng/µL, c(R0011-HisSig)=12.5 ng/µl --> worked
- Gel for analysis of ligation and PCR
- 2% Agarose in 1x TAE
- 110 V, 90 min
- stained with SybrGold 1:10000 20 min
- pure R0011 PCR product used as control
File:GEL 20100419beschriftet.png
LMW
| 4 µl
|
R0011-TrpSig
| 5 µL
|
R0011-HisSig
| 5 µL
|
R0011
| 5 µL
|
Samples seem to have run further than the buffer/dye-Front! But: Ligation Products show bands at shorter lengths than R0011 alone --> Ligation didn't work ?!?
- Ligation of HisSig/TrpSig with R0011in 2 reactions
| used Volume
| concentration
|
pSB1K3
| 5 uL
| 10 ng/µL (nanodrop)
|
B0014
| 3 uL
| 5 ng/µL approx.*
|
* approximated from the amount used in the digestion before
20.04.2010
- Gel for analysis of ligation and PCR (repeat of yesterday's gel)
- 2% Agarose in 1x TAE
- 130 V, 75 min
- stained with SybrGold 1:10000 60 min
- pure R0011 PCR product used as control
- Excision and purification of marked bands at 200bp using QIA Kit, elution in 30 µl H2O
File:Gel100420marked.png
- PCR of excised and purified bands of R0011-TrpSig and R0011-HisSig
- complete samples (30 µl) used as templates
- Purification with the Zymo Kit, Elution in 30 uL H2O
- Concentrations of PCR-products: 0.5-1 ng/µl --> Gel excision or PCR didn't work
- Transformation (Woehlke-Lab)
- 8 µl of ligation product pSB1K3-B0014 to 50 µl XL-10 competent cells
- 200 µl plated on a Kanamycin-containing Plate
- remaining 800 µl stored @4°C in S1-lab
21.04.2010
- Gel for analysis of ligation and PCR (repeat of yesterday's gel)
- 2% Agarose in 1x TAE
- 110 V, 90 min
- stained with SybrGold 1:10000 80 min
- pure R0011 PCR product used as control
- Excision and purification of marked bands at 200bp using Zymo 5 Kit, elution in 20 µl H2O
File:100421beschriftet.gif
- PCR of excised and purified bands of R0011-TrpSig and R0011-HisSig
- complete samples (20 µl) used as templates
- Purification with the Zymo Kit, Elution in 25 uL H2O
- Concentrations of PCR-products:
- R0011-TrpSig: 22.5 ng/µl
- R0011-HisSig: 9.5 ng/µl
--> worked!!!!!
- 'Colony PCR
- 7Colonies picked and resuspended in 20 µl LB+Kana (each)
- PCR of 2 µl of each sample, 2 µl as negative control (Program: ColonyPCR, modified)
- 15 µl of each sample mixed with 3 µl GLPn and loaded to Gel
File:100421colony.png
- Overnight cultures:
- remaining 18 µl of samples 1, 3, 6, and 7 added to 5 ml LB + kanamycin
- 37°C on Shaker
22.04.2010
- Gel for purification of PCR products R0011-TrpSig and R0011-HisSig (yesterday's result)
- 2% Agarose in 1x TAE
- 110 V, 90 min
- stained with SybrGold 1:10000 30 min
- pure R0011 PCR product used as control
- Excision and purification of marked bands at 200bp using Zymo 5 Kit, elution in 20 µl H2O
File:100422beschriftet.png
- Miniprep
- Result: about 4 µg Plasmid
23.04.2010
template
| template volume
| restriction enzymes
| Buffer
|
HisSig (1:100 dilution)
| 5 µl
| EcoRI, SpeI
| NEB4
|
TrpSig (1:100 dilution)
| 5 µl
| EcoRI, SpeI
| NEB4
|
| psB1K3-B0014 from Miniprep (No 7, 35 ng/µl)
| 5 µl
| EcoRI, XbaI
| NEB4
|
Incubated 90 min @ 37°C
- Gel for purification of the cleaved plasmid
- 2% Agarose in 1x TAE (leftover from yesterday)
- 140 V, 90 min
- stained with SybrGold 40 min
- 4 µl 1 kb ladder, 10 µl purified digestion + 2 µl GLPn, 10 µl purified digestion + 2 µl GLPn
- Digestion worked (partly). band at 2400 bp cut out
File:100423beschriftet.png
- Purification of DNA from Gel
- A260/A230 and A260/A280 values were strange (see labbook)
Close
Week04
Cloning Measurements Measurements Measurements
26.04.2010
- Digestion of pSB1K3-B0014 with EcorI and XbaI
- 10 µl template (No1, 50 ng/µl)
- 5 µl BSA, 5 µl Buffer NEB#4
- 1 µl EcoRI, 1 µl XbaI
- 28 µl H2O
- 1.5 h @ 37°C
- Purification with Zymo5 Kit, elution in 20 µl H2O
- Ligation of Signals and PSB1K3-B0014
- 3 µl of each sample, end volume 20 µl
- Preparation of Measurement Plasmid from Folder, Transformation
- Plate 1022, Spots 1E, 1G, 2A: pSB1A10 with different Inserts, all inserts are Zinc-finger constructs with about 1,6 kb
- Transformation of XL10 with Ligation Products (8 µl each) and pSB1A10 (2 µl each)
- Preparation of Measurement Plasmid from Folder, Transformation
- Plate 1022, Spots 1E, 1G, 2A: pSB1A10 with different Inserts, all inserts are Zinc-finger constructs with about 1,6 kb
- growing over night cultures of remaining PSB1K3-B0014-transformed cells
27.04.2010
- Plenty of cultures on both HisSig and TrpSig Ligation plates, but nothing on pSB1A10 plates! --> repeat DNA extraction, ask Prof. Simmel for new Distribution
- Colony PCR
- 7Colonies picked and resuspended in 20 µl LB+Kana (each)
- PCR of 2 µl of each sample, 2 µl as negative control (Program: ColonyPCR, modified)
- 10 µl of each sample mixed with 2 µl GLPn and loaded to Gel
File:100427beschriftet.png
Many colonies with pSB1K3-B0014, not one with pSB1K3-Sig-B0014
- Miniprep of PSB1K3-B0014
- Samples I and II: 5 ml overnight cultures, centrifuged 10 min @ 3200 g, resuspended in 600 µl of the same culture
- Samples III and IV: 600 µl overnight cultures
- all samples mixed with 100 µl lysis buffer, Miniprep with Zyppy kit, each sample eluted in 50 µl H2O
- Concentration measured (Nanodrop, LP=1mm, Factor 10, 4 µl sample)
- cI=61.5 ng/µl
- cII=33.5 ng/µl
- cIII=103 ng/µl
- cIV=108 ng/µl
--> Better results for 600 µl cultures without centrifuging!!!
- Preparation of Measurement Plasmid from Folder, Transformation
- Plate 1022, Spots 1E, 1F, 1G, 1H, 2A: pSB1A10 with different Inserts, all inserts are Zinc-finger constructs with about 1,6 kb
28.04.2010
- No colonies on plates from Yesterday's transformations, but on the older plates (from monday) some colonies appeared
- 7 Colonies picked and resuspended in 20 µl LB0 (each)
- 1&2 from plate "1E", 3&4 from plate "1G", 5,6&7 from plate "2A", 8 LB0
- Colony PCR
- PCR of 2 µl of each sample, 2 µl as negative control (Program: ColonyPCR, modified)
- 15 µl of each sample mixed with 3 µl GLPn and loaded to Gel
- 1% Agarose in 1xTAE, 95 V, after 50 minutes changed to 110 V
File:100428beschriftet.png
- Digestion of pSB1K3-B0014 with EcorI and XbaI
- 10 µl template (No1, 50 ng/µl)
- 5 µl BSA, 5 µl Buffer NEB#4
- 1 µl EcoRI, 1 µl XbaI
- 28 µl H2O
- 1.5 h @ 37°C
- heat inactivation 5min @60°C
- Dephosphorylation of restricted vector
- Purification with Zymo5 Kit, elution in 20 µl H2O
- loaded on gel (with 4 µl GLPn) (Gel shown above)
- Gel excision with Zymo Kit
29.04.2010
- Digestion of pSB1K3-B0014 with EcorI and XbaI
- 10 µl template (NoIV, 108 ng/µl)
- 5 µl BSA, 5 µl Buffer NEB#4
- 1 µl EcoRI, 1 µl XbaI
- 28 µl H2O
- 1.5 h @ 37°C
- heat inactivation 5min @60°C
- Dephosphorylation of restricted vector
- Purification with Zymo5 Kit, elution in 20 µl H2O
- loaded on gel (with 4 µl GLPn)
File:X
- Gel excision with Zymo Kit
- Digestion of R0011 with SpeI
- 10 µl template (R0011, X ng/µl)
- 5 µl BSA, 5 µl Buffer NEB#4
- 1 µl SpeI
- 29 µl H2O
- 1.5 h @ 37°C
- Ligation
- 5 µl R0011 (S-digested) with 12 µl TrpSig or HisSig, respectively (X-digested)
- complete ligation (20 µl) loaded on Gel (with 4 µl GLPn)
File:100429beschriftet.png
- Gel excision with Zymo Kit, eluted in 42 µl H2O
- Transformation
- 50 µl XL-10 transformed with 2 µl of pSB1A10 prepared from IGem 2009 Distribution (13 µl left in pink Box @-20°C)
-
30.04.2010
PCR R0011-HisSig and R0011-TrpSig --> 13 ng/µl x 20 µl
Close
Week05
Cloning Measurements Measurements Measurements
04.05.2010
- Digestion of pSB1K3-B0014 with EcorI and XbaI
- 10 µl template (NoIII, 103 ng/µl)
- 5 µl BSA, 5 µl Buffer NEB#4
- 1 µl EcoRI, 1 µl XbaI
- 28 µl H2O
- 1.5 h @ 37°C
- Purification with Zymo5 Kit, elution in 15 µl H2O
- loaded on gel (with 3 µl GLPn)
File:100504beschriftet.png
- Gel excision with Zymo Kit
- Digestion of R0011-HisSig and R0011-TrpSig with EcorI and SpeI
- 10 µl template (PCR-product)
- 5 µl BSA, 5 µl Buffer NEB#4
- 1 µl EcoRI, 1 µl SpeI
- 28 µl H2O
- 1.5 h @ 37°C
- Purification with Zymo5 Kit, elution in 20 µl H2O
- Ligation
- 4 µl R0011-Signal (E/S-digested) with 4 µl pSB1K3-B0014 (E/X-digested)
- 15 min @ RT, 20 min heat inactivation @ 65°C
- Overrnight liquid cultures of pSB1A10-RFP made from
--> each in 600 µl LB+Carbenicillin (=Ampicillin) @37°C
05.05.2010
- Miniprep of pSB1A10; 4 samples (1, 2; 3a; 3b)
- eluted in 50 µl H2O each
- Concentrations:
- c1=37,5 ng/µl
- c2=56,5 ng/µl
- c3a=46,5 ng/µl
- c3b=30 ng/µl
- Digestion of pSB1A10 with EcorI and PstI, 4 samples (1, 2; 3a; 3b)
- 15 µl template
- 5 µl BSA, 5 µl Buffer NEB#3
- 1 µl EcoRI, 1 µl PstI
- 23 µl H2O
- 1.5 h @ 37°C
- heat inactivation 5min @60°C
- Purification with Zymo5 Kit, elution in 15 µl H2O
- loaded on gel (with 3 µl GLPn)
File:100505beschriftet.png
Insert @ 1 kb as expected, but vector @ 2kb and not @ 5kb as expected!!!!
--> Wrong Plasmid! Comparison to the [http://partsregistry.org/cgi/assembly/plate_egel.cgi?id=615 Gel in the registry] shows: The Distribution contains the wrong plasmid!
- Digestion of HisTerm and TrpTerm with EcorI and PstI
- 5 µl template
- 5 µl BSA, 5 µl Buffer NEB#3
- 1 µl EcoRI, 1 µl PstI
- 33 µl H2O
- 1.5 h @ 37°C
- Clones picked: 7 from each Plate (pSB1K3-R0011-TrpSig-Boo14 and pSB1K3-R0011-HisSig-Boo14)
- Colony PCR
- PCR of 2 µl of each sample, 2 µl as negative control (Program: ColonyPCR)
- 15 µl of each sample mixed with 3 µl GLPn and loaded to Gel
- 2% Agarose in 1xTAE, 130 V, 90 min
File:100505bbeschriftet.png
06.05.2010
- Digestion of pSB1K3 with EcorI and XbaI
- 20 µl template (sample III, 103 ng/µl)
- 5 µl BSA, 5 µl Buffer NEB#3
- 1 µl EcoRI, 1 µl XbaI
- 18 µl H2O
- 1.5 h @ 37°C
- heat inactivation 5min @60°C
- loaded on gel (with 10 µl GLPn) in 4 lanes
File:100506beschriftet.png
- Gel excision with Zymo Kit (lanes 1&2) and with Qiaquick Kit (lanes 3&4)
- c1=4.5 ng/µl
- c2=3.5 ng/µl
- c3=2 ng/µl
- c1=7 ng/µl
- A260/A230 and A260/A280 values were strange (see labbook)
- Ligation
- 4 µl R0011-Signal (E/S-digested) with 10 µl pSB1K3-B0014 (E/X-digested, from 23.04.)
- 15 min @ RT, 20 min heat inactivation @ 65°C
07.05.2010
- Clones picked: 7 from each Plate (pSB1K3-R0011-TrpSig-Boo14 and pSB1K3-R0011-HisSig-Boo14)
8=======o
---Too damn stupid to do a PCR!!!---
8=======o
- replated picked clones on new plates, incubated at RT
Close
Week06
Cloning Measurements Measurements Measurements
10.05.2010
- Colony PCR of picked clones from Fr 07.05.2010
- PCR of 2 µl of each sample, 2 µl as negative control (Program: ColonyPCR, modified)
- 15 µl of each sample mixed with 3 µl GLPn and loaded to Gel
- 2% Agarose in 1xTAE, 120 V, 110 min
- stained in SybrSafe 50 min
File:100510beschriftet.png
Interpretation:
Colonies contain an Insert with Prefix and Suffix, length is 200 bp. This is too short for the desired R0011-Signal-B0014 (245 or 247 bp) construct, but longer than B0014 (136 bp) which was the Insert in the digested vector.
Possible explanation: Ligation worked, but not with R0011-Signal-construct but with R0011 or Signal. Which?
| fragment
| length without Prefix/Suffix
| length with Prefix/Suffix
|
| R0011
| 55 bp
| 96 bp
|
| B0014
| 95 bp
| 136 bp
|
| TrpSig/HisSig
| 34 bp/32 bp
| 75 bp/73 bp
|
| TrpSig-B0014/HisSig-B0014
| 135 bp/ 133 bp
| 176 bp/ 174 bp
|
| R0011-TrpSig-B0014/R0011-HisSig-B0014
| 206 bp/ 204 bp
| 247 bp/ 245 bp
|
| R0011-B0014
| 156 bp
| 197 bp
|
| R0011-TrpSig/R0011-HisSig
| 95 bp/93 bp
| 136 bp/134 bp
|
Prefix: 20 bp; Suffix: 21 bp; X-S-scar: 6 bp
--> it looks as if R0011 is ligated to B0014, which makes the whole construct wothless. The R0011-HisSig control looks more like R0011 alone as well.
11.05.2010
- Ligation
- 4 µl Signal (E/S-digested; from ) with 5 µl pSB1K3-B0014 (E/X-digested; from)
- 15 min @ RT
12.05.2010
- Colony PCR of picked clones from Tu 12.05.2010
- PCR of 2 µl of each sample, 2 µl as negative control (Program: ColonyPCR, modified)
- 15 µl of each sample mixed with 3 µl GLPn and loaded to Gel
- 2% Agarose in 1xTAE, 120 V, 110 min
- stained in SybrSafe 60 min
File:100512beschriftet.png
| fragment
| length without Prefix/Suffix
| length with Prefix/Suffix
| length after PCR
|
| R0011
| 55 bp
| 96 bp
| 104 bp
|
| B0014
| 95 bp
| 136 bp
| 154 bp
|
| TrpSig/HisSig
| 34 bp/32 bp
| 75 bp/73 bp
| 93 bp/91 bp
|
| TrpSig-B0014/HisSig-B0014
| 135 bp/ 133 bp
| 176 bp/ 174 bp
| 194 bp/ 192 bp
|
R0011-TrpSig-B0014/
R0011-HisSig-B0014
| 206 bp/ 204 bp
| 247 bp/ 245 bp
| 265 bp/ 263 bp
|
| R0011-B0014
| 156 bp
| 197 bp
| 215 bp
|
| R0011-TrpSig/R0011-HisSig
| 95 bp/93 bp
| 136 bp/134 bp
| 154 bp/152 bp
|
Prefix: 20 bp/29 bp after PCR; Suffix: 21 bp/30 bp after PCR; X-S-scar: 6 bp
14.05.2010
- Digestion of pSB1K3 with EcorI and XbaI
- 10 µl template (sample III, 103 ng/µl)
- 2 µl BSA, 2 µl Buffer NEB#4
- 1 µl EcoRI, 1 µl XbaI
- 4 µl H2O
- 1 h @ 37°C
- loaded on gel (with 4 µl GLPn) in 1 lane
File:100514beschriftet.png
- Gel excision with Zymo Kit
- Digestion of HisSig and TrpSig with EcorI and SpeI
- 10 µl template ("1:100")
- 2 µl BSA, 2 µl Buffer NEB#3
- 1 µl EcoRI, 1 µl SpeI
- 4 µl H2O
- 1.5 h @ 37°C
- Purification with Zymo 5
- or heat inactivated (20 min @ 80°C)
- Transformation
- 50 µl XL-10 transformed with 10 µl of Ligation mix
- 50 µl untransformed cells plated on Kana-plate as control
Close
Week07
Cloning Measurements Measurements Measurements
17.05.2010
- Plates from Friday:
- plenty colonies on control plate --> XL10 cells are impure!
- use DH5α from now on!!!!!
- Transformation
- 50 µl DH5α transformed with 10 µl of Friday's Ligation mix
- plated on Kana-Plates; Overnight @ 37°C
- DNA Isolation from BioBrick Distribution 2010
- 10 µl H2O added to Well 1A of plate 1 containing pSB1A10 with RFP-insert
- 2 µl used for Transformation of 50 µl DH5α-cells
- plated on Carbenicillin (=Amp-analogon)-plates, Overnight @ 37°C
18.05.2010
- Colony PCR of picked clones
- PCR of 2 µl of each sample, 2 µl as negative control (Program: ColonyPCR)
- 10 µl of each sample mixed with 10 µl Formamide loading buffer and loaded to Polyacrylamide Gel
Samples:
LMW|R0011|HisSig|TrpSig|HisSig E/S-Dig|TrpSig E/S-Dig|B0014|LMW2|Colony PCR His1|His2|Trp1|Trp2|control|HisTerm|TrpTerm|HisTerm E/P-Dig|TrpTerm E/P-Dig
- 5 µl of each samples mixed with 5 µl formamide loading dye and loaded to gel(except Ladder and colonyPCR)
- LMW: 3 µl LMW (Korbinian) + 3 µl Formamide loading Dye
- LMW2: 5 µl LMW Quickload (with GLP) + 10 µl Formamide loading Dye
- stained in SybrSafe
Important Mistake! See below Gel!
File:100518beschriftet.png
IMPORTANT MISTAKE: DENATURING GELS NOT USEFUL FOR dsDNA!!!
REPEAT WITH NATIVE GEL, IGNORE INTERPRETATION!!!
(*R0011 and B0014 look normal
- ColonyPCR: bands that look like B0014 in all clones (and in control???? strange!) --> Religation?
- Signals at the wrong size: should be about 75 bp, look like 200 bp!!!
- terminators completely strange: should be around 100 bp!
--> are all of our sequences just wrong???
What are we going to do? Order everything new?)
19.05.2010
- Gel from Korbinian
- 5 µl of each samples mixed with 5 µl formamide loading dye and loaded to gel(except Ladder and colonyPCR)
- LMW: 3 µl LMW (Korbinian) + 3 µl Formamide loading Dye
- colonyPCR: from Tuesday, 8 µl sample with 8 µl Formamide loading Dye
- stained in SybrSafe 20 min
File:100519paabeschriftet.png
- Colony PCR
- 4 colonies picked from each Plate (Ligations from yesterday; Signal-B0014)
- 15 µl of each Sample mixed with 3 µl GLPn and loaded to Gel:
- 3% Agarose in 1x TBE, 130 V
File:100519beschriftet.png
20.05.2010
- Digestion
- template
- 2 µl BSA
- 2 µl Buffer
- 1 µl of each enzyme
- water to reach 20 µl
Template
| Enzymes
| NEB Buffer #
|
HisTerm & TrpTerm (10 µl)
| EcoRI, PstI
| 3
|
HisSig & TrpSig (10 µl)
| EcoRI, SpeI
| 4
|
B0014 (5 µl)
| XbaI, PstI
| 3
|
pSB1A10_RFP (14 µl)
| EcoRI, PstI
| 3
|
pSB1K3_RFP (14 µl)
| EcoRI, PstI
| 3
|
pSB1K3_B0014 N° 4 (14 µl)
| EcoRI, XbaI
| 4
|
- incubated @37°C for 1.5 h
- digested inserts heat inactivated (20 min @ 80°C)
- digested plasmids loaded on gel (with 4 µl GLPn) in 1 lane
File:100520beschriftet.png
- Gel excision with Zymo Kit
- c(pSB1A10, I)=4.5 ng/µl
- c(pSB1A10, II)=1.5 ng/µl ?!?!?!
- c(pSB1K3 E/P)=7 ng/µl
- c(pSB1K3_B0014 E/X)=2.5 ng/µl
- Gel of PCR products
- 3% Agarose in 1x TBE; 2h @130 V
File:100520bbeschriftet.png
- Ligation
- templates
- 2 µl T4-buffer 10x
- 1 µl T4-Ligase
- Water to reach 20 µl
Vector
| Insert
|
psB1A10 (E/P; sample I) (10 µl)
| TrpTerm (E/P) (4 µl)
|
psB1A10 (E/P; sample I) (10 µl)
| HisTerm (E/P) (4 µl)
|
psB1K3_B0014 (E/X) (12 µl)
| HisSig (E/S) (2 µl)
|
psB1K3_B0014 (E/X) (12 µl)
| TrpSig (E/S) (2 µl)
|
psB1K3 (E/P) (8 µl)
| HisSig (E/S)(2 µl) + B0014 (X/P)(1.5 µl)
|
psB1K3 (E/P) (8 µl)
| TrpSig (E/S)(2 µl) + B0014 (X/P)(1.5 µl)
|
- Transformation
- 50 µl DH5a transformed with 10 µl of Ligation mix
- 50 µl DH5a transformed with 2 µl of pSB1K3_B0014
- 50 µl DH5a transformed with 2 µl of pSB1K3_RFP
- 50 µl DH5a transformed with 2 µl of pSB1A10_RFP
21.05.2010
- Colony PCR
- 4 colonies picked from each Plate (pSB1K3_HisSig_B0014, pSB1K3_TrpSig_B0014, pSB1K3_HisSig_B0014 double ligation, pSB1K3_TrpSig_B0014 double ligation)
- each clone resuspended in 20 µl LB0, 3 µl used as template for PCR
- 15 µl of each Sample mixed with 3 µl GLPn and loaded to Gel:
- 3% Agarose in 1x TBE, 130 V
-
File:100521beschriftet.png
Close
Week08
Cloning Measurements Measurements Measurements
25.05.2010
- Colony PCR
- 4 colonies picked from each Plate
- pSB1K3_HisSig_B0014
- pSB1K3_TrpSig_B0014
- pSB1K3_HisSig_B0014 double ligation
- pSB1K3_TrpSig_B0014 double ligation
- pSB1A10_TrpTerm
- pSB1A10_HisTerm
- each clone resuspended in 20 µl LB0, 2 µl used as template for PCR
- 15 µl of each Sample mixed with 3 µl GLPn and loaded to Gel:
- 3% Agarose in 1x TBE, 220 V (double Gel, 35 cm)
- stained in SybrSafe
File:100525beschriftet.png
File:100525bbeschriftet.png
- overnight cultures made of
- HisSig 3, DL1, DL4
- TrpSig DL2, DL4
- HisTerm/TrpTerm 1,2,3
26.05.2010
- Miniprep of cultures set up 25.05.2010
- HisSig 3, DL1, DL4
- TrpSig DL2, DL4
- HisTerm/TrpTerm 1,2,3
- Restriction
- analytical: E/P HisSig(3, DL2); TrpSig(DL2, DL4): 1.5 h 37 °C
- prep: E/X HisSig(3, DL2); TrpSig(DL2, DL4): 1.5 h 37 °C
- prep: E/S R0011: 1.5 h 37 °C, inactivation 20 min @ 80 °C
- total volume each 20 u, 10 uL template
- Gel: 1% Agarose, TAE - 1,5 h 110 V
File:100526beschriftet.png
- Ligation:
- HisSig_3 (E/X) with R0011 (E/S)
- HisSig_DL2 (E/X) with R0011 (E/S)
- TrpSig_DL2 (E/X) with R0011 (E/S)
- TrpSig_DL4 (E/X) with R0011 (E/S)
- batches
- total volume 20 uL
- 2 uL R0011 (E/S)
- 2 uL T4 buffer
- 1 uL T4 Ligase
- 15 uL vector
- PCR
- His and TrpSig
- His and TrpTerm
- R0011
- B0014
- 50 uL total volume
- 1 uL template
- 1 ul G1004
- 1 uLG1005
- 0.2 uL Taq
- 5 uL Taq standard buffer
- rest water
27.05.2010
- Colony PCR
- 3 colonies picked from each Plate
- pSB1K3_R0011_HisSig_B0014 (N° 3 from yesterday)
- pSB1K3_R0011_HisSig_B0014 (N° DL1 from yesterday)
- pSB1K3_R0011_TrpSig_B0014 (N° DL2 from yesterday)
- pSB1K3_R0011_TrpSig_B0014 (N° DL4 from yesterday)
- "N6"
- "N15"
- each clone resuspended in 20 µl LB0, 2 µl used as template for PCR
- 15 µl of each Sample mixed with 3 µl GLPn and loaded to Gel:
- 3% Agarose in 1x TBE, 220 V (double Gel, 35 cm)
- stained in SybrSafe
File:100527beschriftet.png
| fragment
| length without Prefix/Suffix
| length after PCR
|
| R0011
| 55 bp
| 104 bp
|
| B0014
| 95 bp
| 154 bp
|
| TrpSig/HisSig
| 34 bp/32 bp
| 93 bp/91 bp
|
| TrpSig-B0014/HisSig-B0014
| 135 bp/ 133 bp
| 194 bp/ 192 bp
|
R0011-TrpSig-B0014/
R0011-HisSig-B0014
| 206 bp/ 204 bp
| 265 bp/ 263 bp
|
|
|
|
|
| I712074 ("N6")
| 46 bp
| 105 bp
|
| I719005 ("N15")
| 23 bp
| 82 bp
|
Prefix: 29 bp after PCR; Suffix: 30 bp after PCR; X-S-scar: 6 bp
- overnight cultures made of
- HisSig 3_1, DL1_3
- TrpSig DL4_1, DL4_3
- N15 1&2
- HisTerm/TrpTerm (picked colonies from yesterdays plates #2 each)
- Purification of yesterday's PCR
- elution in 50 µl H2O
- c(HisSig)=5.5 ng/µl
- c(TrpSig)=10 ng/µl
- c(HisTerm)=6.5 ng/µl
- c(TrpTerm)=6.5 ng/µl
- c(R0011)=13 ng/µl
- c(B0014)=12.5 ng/µl
- 5 µl loaded on gel with 1 µl GLPn; 3% Agarose in 1x TBE, 220 V (double Gel, 35 cm)
File:100527bbeschriftet.png
28.05.2010
- Miniprep of cultures from 27.05.2010, Elution in 50 uL nuclease free water
- (1)HisSig 3_1
- (2)HisSig DL1_3
- (3)TrpSig DL4_1
- (4)TrpSig DL4_3
- (7)HisTerm#1
- (8)HisTerm#2 (7 ml culture)
- (9)TrpTerm#1
- (10)TrpTerm#2 (7ml culture)
- (5)N15-1 (=BBa_I719005)
- (6)N15-2 (=BBa_I719005)
sample
| DNA concentration (ng/uL)
|
HisSig 3_1
| 6
|
HisSig DL1_3
| 11
|
TrpSig DL4_1
| 16
|
TrpSig DL4_3
| 21.5
|
HisTerm#1
| 27
|
HisTerm#2
| 66
|
TrpTerm#1
| 34
|
TrpTerm#2
| 29.5
|
N15-1 (=BBa_I719005)
| 19.5
|
N15-2 (=BBa_I719005)
| 18.5
|
- analytical digest (E/P)
- of all samples. total volume 20 uL, 5 uL template used for Term-constructs, 10 uL teplate for all others
- Agarose Gel
- 3% broad range agarose in TBE. Run in TBE, 140 V, 1.50 h
- stained with SybrGold, 45 min
- signals look fine
- terminators also (without pre/suffix 97/104 bp)
- T7 promoter without pre/suffix has a length of 23 bp + cut pre/szffix ca at 60 bp -->buffer?
File:100528 beschriftet.png
Close
Week09
Cloning Measurements Measurements Measurements
31.05.2010
- Digestions
- 15N-1 (BBa_I719005, 19.5 ng/uL) with S/P
- HisSig/TrpSig (6.5/10 ng/uL) with X/P
- 2 h 37 °C
- heat inactivation of insert-digestions
- Gel: 1% Agarose in 1x TAE
- 1 h 25 min, 115 V
- stained with SybrGold, 40 min
- File:100531.png
- Band at ~2100b cut und purified using the zymo kit
- ligation
- HisSig (4 uL of digest) with purified plasmid (with BBa_I719005)
- TrpSig (2 uL of digest) with purified plasmid -=-
- reason: concentration of His Sig before digest was 1/2 of TrpSig
- of DH5a with Ligation batches, HisSig1-3, HisSig3-1, TrpSig DL4-1, TrpSig 4-3
01.06.2010
- of Ligations transformed into DH5a yesterday
- 4 Colonies of each Ligation
- Gel: 3% broad range Agarose in 1xTBE
- 1.5 h 140 V
- stained with SybrGold
File:100601.png
- calculation for the expected size of the fragments
| part
| size (bp)
|
| HisSig
| 32
|
| TrpSig
| 34
|
| T7 promoter
| 23
|
| prefix
| 20
|
| suffix
| 21
|
| X/S scar
| 6
|
- in PCR we get additional bp due to the primers - +9 at pre/suffix=+18 bp
- overall size of the fragments expected to come out of the PCR: T7_HisSig: 120 bp, T7_TrpSig: 122 bp
- 5 ml cultures of pSB1K3_R0011_HisSig_B0014 (1_3 & 3_1) and pSB1K3_R0011_TrpSig_B0014 (DL4_1 & DL4_3)
- 1 ml cultures of each colony monitored in Colony PCR
02.06.2010
- Miniprep of yesterdays cultures using Zymokit, elution by nuclease-free water
- Concentration determination
- analytic digestion
- results on gel:
JobNr. Barcode Last change Date/Time Last message / Files 6549287 AE2739 02.06.2010 / 13:51:12 HisSig 1-3-forward G1004
We just received your order. Many thanks.
6549288 AE2738 02.06.2010 / 13:51:12 HisSig 3-1-forward G1004
We just received your order. Many thanks.
6549289 AE2737 02.06.2010 / 13:51:12 TrpSig DL4-1-forward G1004
We just received your order. Many thanks.
6549290 AE2736 02.06.2010 / 13:51:12 TrpSig DL4-3-forward G1004
We just received your order. Many thanks.
6549291 AE2735 02.06.2010 / 13:51:12 HisTerm-forward G1004
We just received your order. Many thanks.
6549292 AE2734 02.06.2010 / 13:51:12 TrpTerm-forward G1004
We just received your order. Many thanks.
- Gel 3% broad range agarose in 1x TBE
- File:100602 beschr.png
Close
Week10
Cloning Measurements Measurements Measurements
07.06.2010
- Sequenbcing results from GATC
- HisSig DL1-3 is ok
- HisSig 3-1 is ok
- TrpSig DL4-1 is ok
- TrpSig DL4-3 is ok
- TrpTerm + HisTerm bad runs... --> new sequencing order with Primer 100 bp upstream (within GFP)
- Files can be found stored in our GATC account
- Sequencing@GATC: both Term-constructs with primer pGFP-FP provided by GATC
- Restrictions
- psB1A10_TrpTerm/HisTerm with Nsi1, Aat2
- pSB1K3_R0011_HisSig/TrpSig_B0014 with Pst1, Aat2
- T7 bb with Spe1, Pst1
- PCRProducts: HisSig/TrpSig with Pst1, Xba1, 2 h @37°C
- all plasmid digests done sequential as enzymes do not have 100% activity in the same buffer, each reaction 1.5 h@37°C
- psB1A10_TrpTerm/HisTerm and T7 bb dephosphorylated the last 30 min
liquid culture (10 ml) of pSB1K3_R0011_HisSig/TrpSig_B0014
08.06.2010
- Sequencing results from GATC: His/TrpTerm with pGFP-FP primer
- HisTerm worked
- TrpTerm worked
- checked with blast2seq
- 8 colonies from each plate of T7His, T7Trp, MonsterHis, MonsterTrp as ligations resulted in many colonies
- for all colony PCR reactions
File:100608 beschriftet.png
File:100608-2 beschriftet.png
Interpretation/Info: The R0011_Sig_B0014 construct was cut with PstI, but not ligated into a PstI site but instead into the NsiI site --> even if sticky end are compatible, the bases in the 3` direction are different --> primer lags 7 bp compared to standard procedure --> we didn´t expect to find the signal construct by colony PCR --> control digestion tomorrow
- T7-Signal constructs seem to have worked, expected size was 23 bp (T7)+ 32 (HisSig)/34 (TrpSig) bp + 30+29 (PCRPre+Suf)=114/116 bp
09.06.2010
- Miniprep of 4 Monster_His, 4 Monster_Trp, 3 T7_His and 3 T7_Trp cultures
- Analytical digestion of plasmids mentioned above
- gel of Monster_Plasmid digestion
File:100609 Monster inverse.jpg
gel didn´t work at all --> even after > 2h, bands were not separated correctly, even the 1kb ladder was "stacked" in the gel-pockets, the 100 bp ladder should show EQUAL distances between the lines [http://www.neb.com/nebecomm/productfiles/778/images/N3231_fig1_v1_000034.gif see here], it looks like the gel was "more dense" at the pockets---> no idea what happened --> repeat Monster-digestion tomorrow?
- gel of T7_Plasmid digestion
File:100609 T7 sig invers.jpg
T7_Trp E + S digestion 107 bp and T7_His 105 bp --> worked for all picked colonies. (regard that there is an excess of plasmid DNA-basepairs of factor >30 --> thats why the inserts are much weaker than the plasmid signals.
occured trouble:
- Ladders and loading dye´s empty --> i used those of eike, BUT: eikes 1 kb ladder is different --> compare [http://www.neb.com/nebecomm/products/productN3272.asp here] and his loading dye was much more diluted, even if there was also 6x Sac GLP written on it -> i hope this won´t cause any trouble
10.06.2010
- Promega E.coli S30 in vitro transcription/translation kit
- Spe1, Aat2 from NEB, 500 U each
- of Ligation colonies from MonsterHis/trp 1-3 and T7His/Trp 1,2,5/1,2,3
- 2h digestion
- Monster: 6 uL DNA template with Aat2/Spe1 in Buffer 4/Bsa
- T7-Signal: 6 uL DNA template with E/P in Buffer 3
- used standards: lmw, 2-log click here
- Gel1: 1% Agarose in 1xTBE for Digestions of Monsterplasmid
- run in big chamber @ 200 V for 1 h 20 min
- File:100610 t7sig.png
- Gel2: 3% Agarose (broad range) in 1xTBE for Digestions of T7-Signal
- run in small chamber @140 V for 1 h 35 min
- File:100610 monster.png
- Conclusions:
- all T7-Signal ligations loaded on the gel worked
- monsterplasmid didn't work? bands at 800 bp, 900 bp, 1.3 kbp, 2.2 kbp, 3 kbp, we SHOULD expect to see our Insert, wich is Prefix+R0011_Signal_B0014_small Suffix, which should run around 300-400 bp...
11.06.2010
- Gel: large 1% Agarose in TAE. Load: The rest of N/A cut Messplasmids from 07.06.2010. Run @220 V for 3.5 h
- fragments expected are 5087 and 176. original size of plasmid is 5263. This is a Try to differ between 5087 and 5263 bp
- File:100611.png
- Band @ 5000 bp of Trp_Term purified, obviously digestion was 100%. Bad oint is that HisTerm includes an Nsi1 cleavage site...
Close
Week11
Cloning Measurements Measurements Measurements
14.06.2010
- 10 uL pSB1A10_TrpTerm Aat2/Nsi1 0.5 ng/uL
- 1 uL R0011_TrpSig_B0014 Aat2/Psb1 11 ng/uL
- 2 uL T4 ligase buffer
- 1 uL T4 Ligase
- 6 uL H2O
- 10 min RT
- Transformation of DH5a with
- Ligation
- T7His#1
- T7Trp#1
- pSB1K3_R0011_TrpSig_B0014
- pSB1K3_R0011_HisSig_B0014
- pSB1A10_TrpTerm
- pSB1A10_HisTerm
15.06.2010
- Over night cultures
- Aliquots of the Promega in vitro expressions kit from e. coli s30 extract:
- 40 uL with aa mix including all aa.
16.06.2010
Fluoresence measurements using in vitro kit
- in vitro kit sample
- adding psBA1A10 Trp_Term --> constant over time, no significant changes compared to kit alone --> high efficiency of AraC
- adding L-(+)Arabinose (final concentration 2%) --> after approx. 10 min significant GFP production --> measuring for xxx min --> RFP is slightly increased (to proof if correlated to GFP peak --> crossdetection)
- adding psB1K3 R0011_TrpSig_B0014
Cell culture
5 ml culture for
- psBA1A10 Trp_Term/HisTerm
- psB1K3 R0011_TrpSig/Hissig_B0014
17.06.2010
Cloning
Digestion of Trp-Sig with E/P and psB1A10 Trp_Term with E/P
- Gel purification of psB1A10 Trp_Term E/P cut
File:100617 pSB1A10 EPcut dunkler.jpg
- Heat inactivation of Trp-Sig E/P cut
- Ligation for 10 min @ RT and Transformation in DH5-a cells
in vitro measurements
18.06.2010
cloning
- Transformation (about 20 colonies) --> picking 5 colonies
- colony PCR
- Gel
2 % broad range agarose, 1 h 120 V File:100618 psb1A10-Trp sig colonypcr.jpg
Sample 2, 4, 5 shows probably Trp-Signal + Pre/Suffix --> send sample 2 for sequencing!
- control digestion of all 10 picked psB1A10-TrpSig in 1% broad range agarose, > 3 h, 120 V
File:100618 psb1A10-Trp sig controlverdau.jpg
--> digestions worked, but again, no insert can be found, despite gel was at maximum resolution ( 3h 120 V, see LMW)
in vitro measurements
f$%&&§ s%§$! again, nothing worked! despite we saw an increasing "GFP signal" comparable to 16.06.10, taking spectra suggested we DONT see significant GFP-production! We used new water for preparing the samples, cleaned cuvettes with "new water", used other DNA-samples etc. Somehow, it seems like we dont express GFP (we compared Christophs results! One should see a really significant spectrum!
Next steps:* Try in vivo measurements, just using psb1A10_xTerm without Signal (thus just measuring plasmid) to proof if kit or measuring plasmid causes this problem!
Close
Week12
Cloning Measurements Measurements Measurements
This is what we did in week xxx.
Close
Week13
Cloning Measurements Measurements Measurements
This is what we did in week xxx.
Close
Week14
Cloning Measurements Measurements Measurements
This is what we did in week xxx.
Close
Week15
Cloning Measurements Measurements Measurements
This is what we did in week xxx.
Close
Week16
Cloning Measurements Measurements Measurements
This is what we did in week xxx.
Close
|
 "
"