Team:TU Munich/Lab
From 2010.igem.org
(→Explanations) |
FPraetorius (Talk | contribs) (→In vivo Measurements) |
||
| Line 8: | Line 8: | ||
Our initial idea to prove our concept of antitermination was to use fluorescent proteins as reporters. This approach gives the opportunity to measure the termination and antitermination efficiency of our designed BioBricks ''in vivo'' as well as ''in vitro'', the latter using a translation kit based on <i>E. coli</i> lysate. Later on, we decided to developed an experiment, that relies only on transcription. In this, we used a fluorescent dye, malachite green, that binds a specific RNA aptamer and thus makes it possible to detect transcription activity.<br> | Our initial idea to prove our concept of antitermination was to use fluorescent proteins as reporters. This approach gives the opportunity to measure the termination and antitermination efficiency of our designed BioBricks ''in vivo'' as well as ''in vitro'', the latter using a translation kit based on <i>E. coli</i> lysate. Later on, we decided to developed an experiment, that relies only on transcription. In this, we used a fluorescent dye, malachite green, that binds a specific RNA aptamer and thus makes it possible to detect transcription activity.<br> | ||
==''In vivo'' Measurements== | ==''In vivo'' Measurements== | ||
| - | ''In vivo'' measurements have the highest complexity compared to any other | + | ''In vivo'' measurements have the highest complexity compared to any other experimental set-up. Our system has to deal with several circumstances a cellular environment comes with, such as interaction with other RNAs, degradation by RNases or unspecific interactions. Nevertheless, the measurements are essential, as our switches should finally work inside cells to fulfill our vision of an intracellular logic network. |
{{:Team:TU Munich/Templates/ToggleBoxStart}} | {{:Team:TU Munich/Templates/ToggleBoxStart}} | ||
===Design=== | ===Design=== | ||
| Line 24: | Line 24: | ||
===Construction and Cloning=== | ===Construction and Cloning=== | ||
| - | Our measuring plasmid is based on the BioBrick pSB1A10, A1, distribution 2010. Unfortunately after two months of cloning we had to recognize that the plasmid in use did not work (see also Biobrick validation--> link). So after the first unsuccessful attempts we decided to reclone the system, substituing RFP to mCherry, a dsRED derivative with a | + | Our measuring plasmid is based on the BioBrick pSB1A10, A1, distribution 2010. Unfortunately after two months of cloning we had to recognize that the plasmid in use did not work (see also Biobrick validation--> link). So after the first unsuccessful attempts we decided to reclone the system, substituing RFP to mCherry, a dsRED derivative with a spectrum in the far red, and adding arabinose inducible promoters in front of both fluorescent proteins. |
Revision as of 16:42, 23 October 2010
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
Experiment DesignWe designed different experiment set-ups with different complexity to test RNA signal/switch pairs based on our concept. In vivo MeasurementsIn vivo measurements have the highest complexity compared to any other experimental set-up. Our system has to deal with several circumstances a cellular environment comes with, such as interaction with other RNAs, degradation by RNases or unspecific interactions. Nevertheless, the measurements are essential, as our switches should finally work inside cells to fulfill our vision of an intracellular logic network.
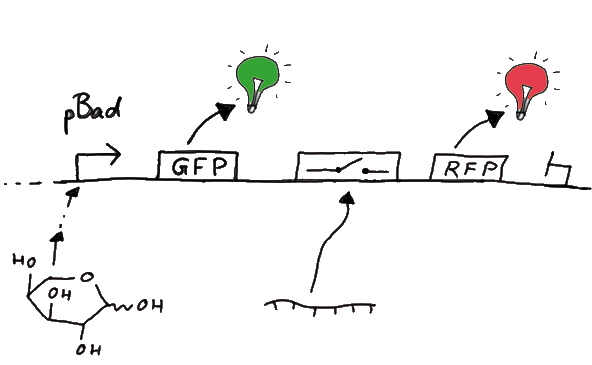
DesignFor the measurements in vivo we decided to use an expression cassette consisting of Green Fluorescent Protein (GFP) coding sequence upstream of the switch and another fluorescent protein coding sequence downstream of it. Both protein coding sequence carry the same ribosome binding site, therefore, the GFP fluorescence can be used as internal control in measurements. Since the spectra should not overlap and to avoid FRET as well as an pure overlap of the spectra, we settled on the usage of red fluorescent protein variants, namely mRFP1 in the first try. While the GFP fluorescence is used to normalize the measurements, the RFP fluorescence is used to detect termination/antitermination.
Upon binding of the signal, the stem loop of the switch would resolve leading to red fluorescence. The GFP fluorescence as internal control carries the advantage that errors in the measurement set can be detected easily. Lack of arabinose or promoter insensitivity can be recognized as well as problems with the fluorescence measurement itself. Plus, we have a way to normalize our measurements and compare different preparations in relation to each other.
Construction and CloningOur measuring plasmid is based on the BioBrick pSB1A10, A1, distribution 2010. Unfortunately after two months of cloning we had to recognize that the plasmid in use did not work (see also Biobrick validation--> link). So after the first unsuccessful attempts we decided to reclone the system, substituing RFP to mCherry, a dsRED derivative with a spectrum in the far red, and adding arabinose inducible promoters in front of both fluorescent proteins.
MeasurementFor switch evaluation, IPTG was added to the cells after about two hours after arabinose induction (baseline). ??? Stimmt das?? A rise of RFP/mCherry emission should be visible in case of a working switch.
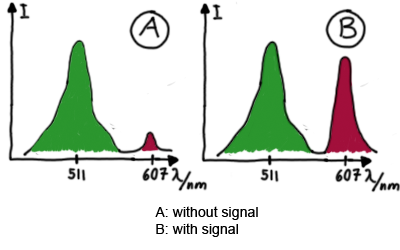
When measuring the termination of our BioBricks and the antitermination by their corresponding signal-RNA, we should be able to observe an increasing RFP emission compared to the GFP emission upon induced signal-RNA production in the cells/in the kit:
With these measurements, it should also be possible to observe differences in efficiency of termination as well as antitermination between our designed switches. In vitro TranslationIn vitro measurements with E. coli lysate make the fluorescence signals independent of cell growth and physical or biological factors, e.g. cell density or growth stadium.
DesignIn this assay we used the same constructs as engineered for the in vivo studies. MeasurementsWe used the cell-free E. coli S30 extract system for circular DNA provided by promega[1], which is prepared by modifications of the Method Zubay et al.[2] described. The characterization of the kit can be obtained from the [http://partsregistry.org/Cell-free_chassis/Commercial_E._coli_S30 Parts Registry]. In vitro TranscriptionAn experiment, in which we detect In vitro transcription, offers an elegant way for a fast and easy prove of principle, since our switch is RNA-based and the whole mechanism takes place on trancriptional level. Most side effects occuring in a complex environment given in a cell or a cell lysate do not arise here.
If measureable effects with our basic concept can be seen in vitro we can use the so gained data to optimize the system in vivo. Since we are working on a totally new principle of trancriptional control, we used this approach for easy variation of different variables like the length of the core unit and the switch to signal ratio.
To study the switches on the transcriptional level gives the advantage, that we would have less interferences and possible artefacts. Also, we are not sure how cellular mechanisms like degradation of RNases or interacting factors as well as molecular crowding influence our systems.
T7 RNA polymeraseThe T7 RNA polymerase is known for satisfying RNA yields together with easy handling. In our approach we had PCR amplified, double stranded switches with an malachitegreen binding aptamer following after the switch (133 bp, see section below) and a single stranded signal with about 30 bp length.
For in vitro expression the T7 RNA Polymerase requires a double stranded promotor region at the beginning of the DNA template but is otherwise capable of handling single stranded DNA, so a sense strain corresponding to the T7 promoter region was added. Transcription is more effective with double stranded DNA as template. Since we ordered the signal sequences we tested we chose the cheaper way in the beginning by using single stranded signals with corresponding sense T7 pieces and switched to double stranded constructs after narrowing down the most promising switch/signal pairs.
E. coli RNA polymeraseIn comparison to the T7 RNA Polymerase the E. coli RNA Polymerase requires slightly more sophisticated proceedings when it comes to the design of switches and handling of the enzyme. The biggest in our case was to store it properly since the only -80°C fridge was in another building. Denaturing Polyacrylamide gel electrophoresisWe also used Polyacrylamide gel electrophoresis (PAGE) for evaluation of termination efficiency of our basic units. Gels containing 15 % acrylamide and 6 M urea were used for separation of 90 (terminated by switch) and 133 bp (continous reading) RNAs.
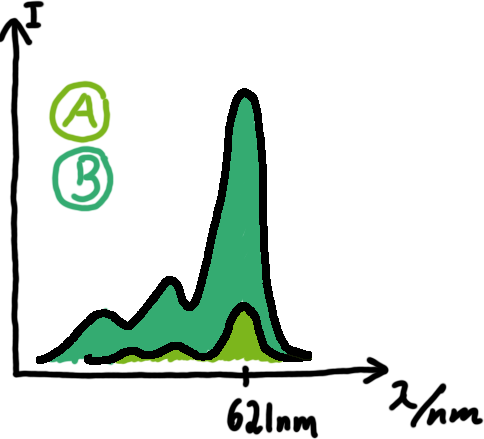
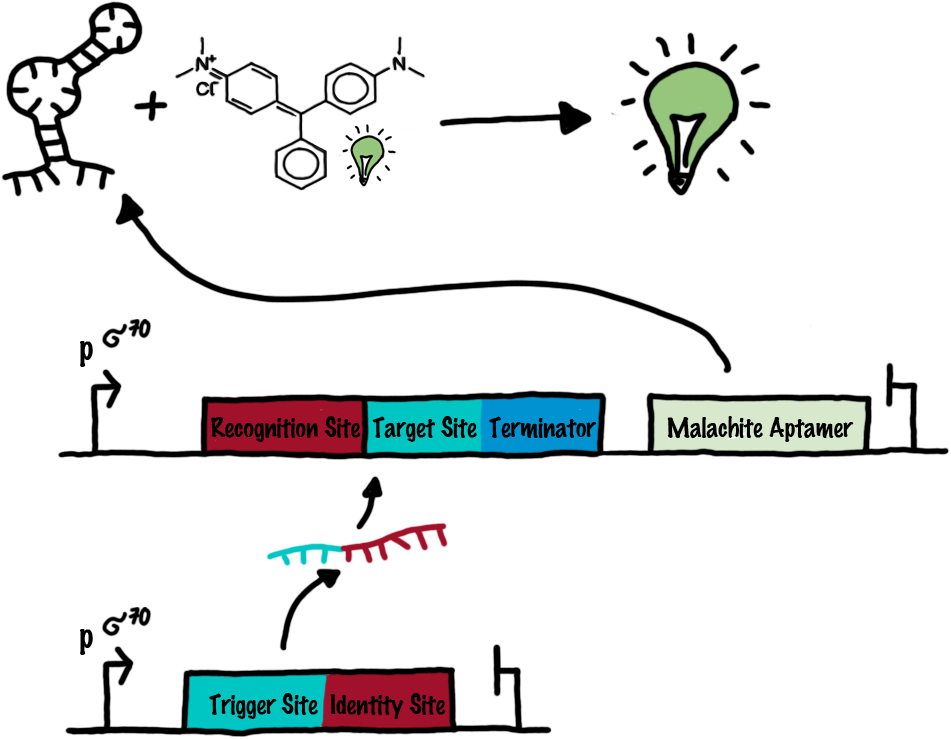
Malachite green assayMalachite-green is a dye with a negligible fluorescence in solution but undergoes a dramatic increase if bound by a RNA -aptamer. Upon binding to the aptamer, the fluorescence of malachite-green increases about 3000 times making it an exceptionel good marker. Since the binding is very specific, transcription in dependence of a signal can be monitored by measuring the fluorescence of malachite-green over time if the aptamer is located behind the switch. Transcription of the aptamer will only take place after anti-termination by a signal. An increase should be visible over time.
OLD: A second possibility to measure parameters of our switches we came up with, was the idea to investigate our system on the transcriptional level only. Therefore, we decided to use malachite green as reporter. Malachite green in a fluorescent dye, whose emission increasing dramaticly (about 3000 times) upon binding of a specific RNA-aptamer. The RNA-aptamer
We made constructs comprising of a sigma(70)-binding promoter followed by a short nonsense sequence, the switches and the aptamer sequence.
References[1] http://www.promega.com/catalog/catalogproducts.aspx?categoryname=productleaf_335&ckt=1 [2] Zubay, G. (1980) Meth. Enzymol. 65, 856–77, Zubay, G. (1973) Ann. Rev. Genet. 7, 267–87. Experimental ResultsIn vivo MeasurementsIn vivo In vitro Measurements
ProtocolsMolecular BiologyPCR
So geht ne PCR CloseDigestion
So geht ne PCR CloseLigation
In vivo Measurement
So haben wir in vivo gemessen CloseIn vitro Translation
So haben wir in vitro gemessen CloseIn vitro Transcription
So haben wir in vivo gemessen CloseLab BookExplanationsIn the following we present an overview regarding our work in the lab. For easier understanding we summarized the work of each week using colored boxes. To get more information about the work and results of a specific week, just click on the according week number. To get a better overview we used the following color code for the boxes:
Chronological Lab Book
Constrcuts for in vivo measurments
08.04.2010Flo & Philipp PCR
09.04.2010Philipp & Flo Gel of PCR products from 08.04.2010
Improved Constructs... Testing
15.04.2010Philipp & Flo
16.04.2010Philipp & Flo
--> worked for R0011, not for B0014
* approximated from the amount used in the digestion beforeClose
Measurements
19.04.2010
* approximated from the amount used in the digestion before 20.04.2010
21.04.2010
--> worked!!!!!
22.04.2010
23.04.2010
Cloning Measurements Measurements Measurements
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close
Cloning Measurements Measurements Measurements
This is what we did in week xxx. Close |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"