Team:Newcastle/Filamentous Cells
From 2010.igem.org
(Difference between revisions)
RachelBoyd (Talk | contribs) |
RachelBoyd (Talk | contribs) (→Filamentous cells genes list) |
||
| Line 2: | Line 2: | ||
=Filamentous Cells= | =Filamentous Cells= | ||
==Filamentous cells genes list== | ==Filamentous cells genes list== | ||
| - | + | ||
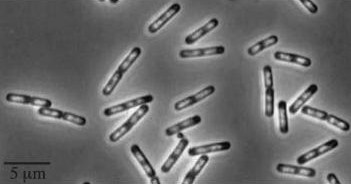
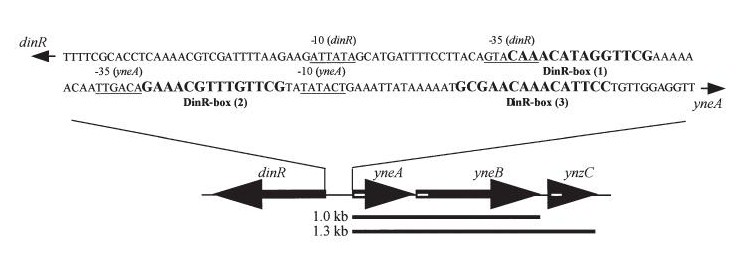
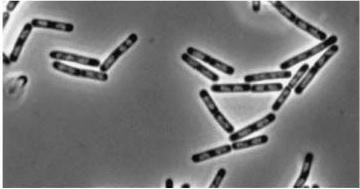
| - | + | *''yneA'' (Transcribed with ''yneB'', ''ynzC'', is an analogue of ''sulA'' in ''E.coli'') * our biobrick is designed to over express this gene r educing cell division possibly by inhibiting FtsZ ring formation or constriction. | |
| - | + | *''dinR'' (Homologue of ''lexA'' in ''E.coli'' transcribed in the opposite direction) | |
| - | + | *''ftsZ'' (Involved in the recruitment of other proteins to the divisisome for cytokinesis, strangely over expression results in disruption of Zring formation as well as reduced expression) | |
| - | + | *''secA'' (Involved in the secretion of extracellular proteins and the insertion of transmembrane proteins) | |
| - | + | * ''recA'' (Involved in SOS response removing the repressor DinR (LexA)) | |
| - | + | *''wpr'' and ''epr'' produce extracellular proteases that cleave the signal peptide/transmembrane domain of YneA | |
| - | + | *''ezr'' produces a protein which sequesters FtsZ monomer by binding its C terminal domain and also inhibits GTP binding; however overexpression does not result in filamentation. | |
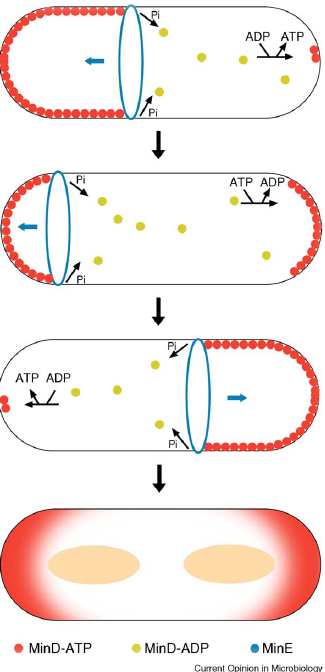
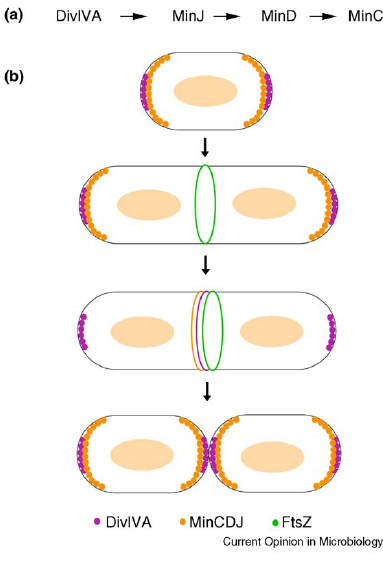
| - | + | * ''min C,D ,J and divIVA'' prevent polar cell division . | |
| - | + | *Positive regulators of FtsZ: ''ftsA, zapA, zipA, ftsL and divIC'' | |
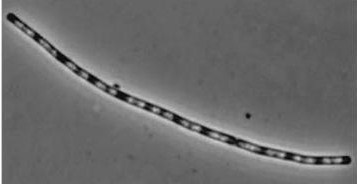
| - | + | *Inhibitors of Daughter cell separation: ''lytC,D,E,F and cwlS'' *Chains rather than filaments, ''yneA'' is also reported to increase the time spent in chains well into the stationary phase of bacterial growth. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Revision as of 12:05, 24 June 2010

| |||||||||||||
| |||||||||||||
Contents |
Filamentous Cells
Filamentous cells genes list
- yneA (Transcribed with yneB, ynzC, is an analogue of sulA in E.coli) * our biobrick is designed to over express this gene r educing cell division possibly by inhibiting FtsZ ring formation or constriction.
- dinR (Homologue of lexA in E.coli transcribed in the opposite direction)
- ftsZ (Involved in the recruitment of other proteins to the divisisome for cytokinesis, strangely over expression results in disruption of Zring formation as well as reduced expression)
- secA (Involved in the secretion of extracellular proteins and the insertion of transmembrane proteins)
- recA (Involved in SOS response removing the repressor DinR (LexA))
- wpr and epr produce extracellular proteases that cleave the signal peptide/transmembrane domain of YneA
- ezr produces a protein which sequesters FtsZ monomer by binding its C terminal domain and also inhibits GTP binding; however overexpression does not result in filamentation.
- min C,D ,J and divIVA prevent polar cell division .
- Positive regulators of FtsZ: ftsA, zapA, zipA, ftsL and divIC
- Inhibitors of Daughter cell separation: lytC,D,E,F and cwlS *Chains rather than filaments, yneA is also reported to increase the time spent in chains well into the stationary phase of bacterial growth.
YneA
Kawai, Y., Moriya, S., & Ogasawara, N. (2003). Identification of a protein, YneA, responsible for cell division suppression during the SOS response in Bacillus subtilis. Molecular microbiology, 47(4), 1113-22. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/12581363.
 "
"