Team:Imperial College London/Lab Diaries/XylE team
From 2010.igem.org
m |
|||
| (5 intermediate revisions not shown) | |||
| Line 14: | Line 14: | ||
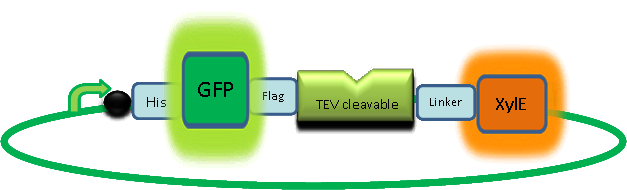
'''Here's a picture of the final construct:''' | '''Here's a picture of the final construct:''' | ||
| - | [[Image:IC_Module3.JPG]] | + | [[Image:IC_Module3.JPG|center]] |
|} | |} | ||
| Line 58: | Line 58: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>mini-prep kit of XylE-transformed E.coli (already overnight grown)</li> | + | <li>mini-prep kit of XylE-transformed ''E.coli'' (already overnight grown)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 73: | Line 73: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>Starting of “Testing expression of XylE in E. | + | <li>Starting of “Testing expression of XylE in ''E.coli''” objective</li> |
<li>1)Annealing EcoRI and speI oligos to J23101 promoter which will be annealed later in front of the RBS-XylE registry gene (overnight) </li> | <li>1)Annealing EcoRI and speI oligos to J23101 promoter which will be annealed later in front of the RBS-XylE registry gene (overnight) </li> | ||
</ul> | </ul> | ||
| Line 88: | Line 88: | ||
'''Thursday, 12-Aug-2010''' | '''Thursday, 12-Aug-2010''' | ||
| - | * | + | * Annealing DNA strands of J23101 promoter in a water bath |
| - | we constructed the standard E.coli promoter J23101 with sticky ends. These ends are complementary to restriction sites made by EcoRI and SpeI enzyme. This promoter will be later used in 3A | + | we constructed the standard ''E.coli'' promoter J23101 with sticky ends. These ends are complementary to restriction sites made by EcoRI and SpeI enzyme. This promoter will be later used in 3A assembly to construct a promoter-RBS-XylE design in a psB1C3 vector. ''E.coli'' will be transformed with this final construct plasmid to assess XylE activity and characterization. It will also be one of the submitted BioBricks. |
| - | * Prepared two overnight cultures of XylE | + | * Prepared two overnight cultures of XylE transformed ''E.coli'' (one 50microliters and one of 450μl) |
| - | these cultures are going to be used tomorrow for mini | + | these cultures are going to be used tomorrow for mini prepping. Mini prep will allow us to isolate ''E.coli'' 's plasmid DNA(which contains the XylE gene). |
'''Friday, 13-Aug-2010''' | '''Friday, 13-Aug-2010''' | ||
| - | * Mini-prep of XylE transformed E.coli | + | * Mini-prep of XylE transformed ''E.coli'' |
| - | Mini | + | Mini prep is usually used to confirm that our gene of interest has not been changed in any way, as the isolated plasmid id sent for sequencing. However, since XylE was taken from the registry, we assume that it is fine and no sequencing is required. The mini prep will later be used for the midi-prep (that gives out higher yields of DNA needed for cloning). |
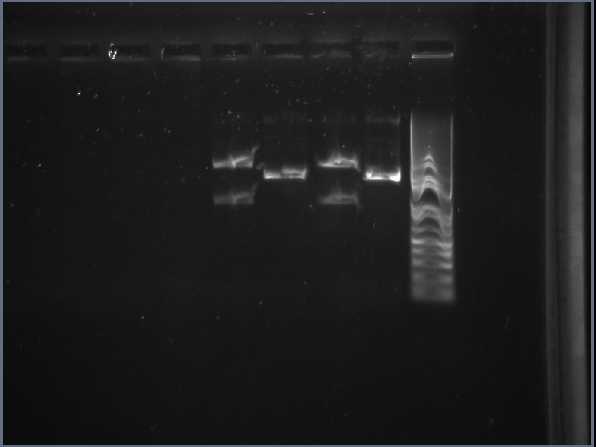
| - | * Gel analysis of plasmid DNA | + | * Gel analysis of plasmid DNA retrieved from mini prep of XylE transformed ''E.coli'', cut with restriction enzymes. From light to the left, 50μg digested DNA : 50 undigested DNA : 450 digested DNA : 450 undigested DNA. In lanes 1 and 3 the smaller band has a size of about 1kB which corresponds to RBS-XylE gene. The bigger bands are the cut vectors. In lanes 2 and 4 is the uncut BioBrick from the registry. It appears smaller on the gel than it actually is as circular DNA travels faster through the pores of agarose gel rather than linearised DNA. |
|} | |} | ||
| Line 129: | Line 129: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>midi | + | <li>midi prep XylE ''E.coli'' (2hrs)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 142: | Line 142: | ||
<ul> | <ul> | ||
<li>3A assemply: make replica plates (overnight)</li> | <li>3A assemply: make replica plates (overnight)</li> | ||
| - | <li>Catechol assay of E.coli </li> | + | <li>Catechol assay of ''E.coli'' </li> |
</ul> | </ul> | ||
</td> | </td> | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>Mini | + | <li>Mini prep XylE ''E.coli''</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 153: | Line 153: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>Midi | + | <li>Midi prep XylE ''E.coli''</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 164: | Line 164: | ||
<ul> | <ul> | ||
<li>restriction digestion of XylE for 3A assemply</li> | <li>restriction digestion of XylE for 3A assemply</li> | ||
| - | <li>gel purification of XylE from | + | <li>gel purification of XylE from restriction digestion</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 170: | Line 170: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>3A | + | <li>3A assembly of vector, XylE and J23101 promoter </li> |
| - | <li>above construct transformed in E.coli</li> | + | <li>above construct transformed in ''E.coli''</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 194: | Line 194: | ||
'''Monday, 16-Aug-2010''' | '''Monday, 16-Aug-2010''' | ||
| - | * Midi | + | * Midi prepped the XylE-transformed ''E.coli''. The DNA yield from the midi prep was 134μg as determined by spectrophotometry. This is the XylE that is going to be used for all further experiments. |
| - | * Restriction digestion of midi | + | * Restriction digestion of midi prepped XylE by Xbal and PstI to prepare it for 3A assembly. (with J23101 promoter and PSB1C3 vector) |
* Gel analysis of the restriction digestion mixture to isolate XylE gene | * Gel analysis of the restriction digestion mixture to isolate XylE gene | ||
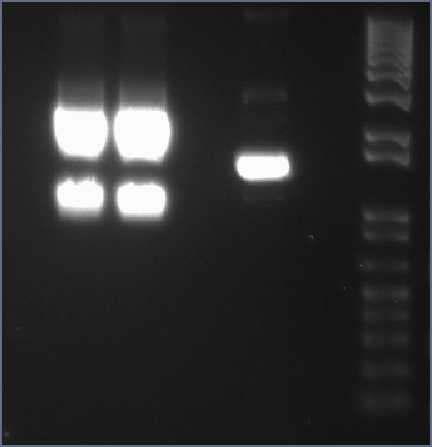
| - | Midi | + | Midi prep XylE digestion with xbaI and PstI The smaller size bands at lanes 2 & 3 are the ones that are going to be cut out and used in gel purification to extract the XylE gene (with sticky ends for XbaL and PstI). |
| - | * I made an overnight culture of Bacillus | + | * I made an overnight culture of ''Bacillus'' |
| - | [[Image:IC_Midi-prep_XylE_digestion_with_xbaI_and_PstI.jpg|thumb|center|400px|Midi | + | [[Image:IC_Midi-prep_XylE_digestion_with_xbaI_and_PstI.jpg|thumb|center|400px|Midi prep XylE digestion with xbaI and PstI The smaller size bands at lanes 2 & 3 are the ones that are going to be cut out and used in gel purification to extract the XylE gene (with sticky ends for XbaL and PstI).]] |
'''Tuesday, 17-Aug-2010''' | '''Tuesday, 17-Aug-2010''' | ||
| - | * | + | * Using the Gel Extraction Kit, we isolated the restriction enzyme cut XylE gene from the agarose gel band. |
| - | * Gel analysis of XylE, J23101 promoter and pSB1C3 vector samples to determine their ratios for 3A | + | * Gel analysis of XylE, J23101 promoter and pSB1C3 vector samples to determine their ratios for 3A assembly ligation |
* 3A assemply of XylE, J23101 promoter and pSB1C3 vector. | * 3A assemply of XylE, J23101 promoter and pSB1C3 vector. | ||
| - | * Transformation of XL-Blue competent E.coli with the above construct. | + | * Transformation of XL-Blue competent ''E.coli'' with the above construct. |
[[Image:IC_XylE-J23101-pSB1C3_Clone_Gel2.JPG|thumb|center|400px|gel analysis of XylE, J23101 promoter and pSB1C3 vector samples to determine the volume ratios of samples to be used for 3A assemply ligation]] | [[Image:IC_XylE-J23101-pSB1C3_Clone_Gel2.JPG|thumb|center|400px|gel analysis of XylE, J23101 promoter and pSB1C3 vector samples to determine the volume ratios of samples to be used for 3A assemply ligation]] | ||
gel analysis of XylE, J23101 promoter and pSB1C3 vector samples to determine the volume ratios of samples to be used for 3A assemply ligation | gel analysis of XylE, J23101 promoter and pSB1C3 vector samples to determine the volume ratios of samples to be used for 3A assemply ligation | ||
| - | * I followed Chris's Bacillus transformation protocol to transform Bacillus with constitutive GFP and RFP DNA as well as a control without DNA | + | * I followed Chris's ''Bacillus'' transformation protocol to transform ''Bacillus'' with constitutive GFP and RFP DNA as well as a control without DNA |
| + | |||
| + | {| style="width:800px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Successful Bacillus transformation! | ||
| + | |- | ||
| + | |align="center"|<html><object width="425" height="344"><param name="movie" value="http://www.youtube.com/v/fFwYCh5wvfc?hl=en&fs=1"></param><param name="allowFullScreen" value="true"></param><param name="allowscriptaccess" value="always"></param><embed src="http://www.youtube.com/v/fFwYCh5wvfc?hl=en&fs=1" type="application/x-shockwave-flash" allowscriptaccess="always" allowfullscreen="true" width="425" height="344"></embed></object></html> | ||
| + | |} | ||
| + | |||
'''Thursday, 19-Aug-2010''' | '''Thursday, 19-Aug-2010''' | ||
| - | * We run the | + | * We run the XylE-GFP1 PCR reaction to construct the His-GFP-Flag and Linker-XylE-Spe construct. |
'''Friday, 20-Aug-2010''' | '''Friday, 20-Aug-2010''' | ||
| - | The J23101 gene in a | + | The J23101 gene in a BioBrick vector containg RFP gene |
| - | * We run a gel on | + | * We run a gel on XylE-GFP1 PCR reaction. Results: GFP was extended successfully, XylE extension FAILED (too much non-specific annealing) |
| - | * Catechol assay on 2hrs bench ligation of promoter, | + | * Catechol assay on 2hrs bench ligation of promoter, XylE and vector failed. |
| - | * Two replica plates of overnight ligated J23101, XylE and pSB1C3 transformed E.coli (one for catechol assay) | + | * Two replica plates of overnight ligated J23101, XylE and pSB1C3 transformed ''E.coli'' (one for catechol assay) |
* The His-GFP-Flag DNA was gel purified | * The His-GFP-Flag DNA was gel purified | ||
| - | * Transformation of XL-1Blue cells with J23101 in J62001 vector from the registry. One more attemp to construct a successful promoter- | + | * Transformation of XL-1Blue cells with J23101 in J62001 vector from the registry. One more attemp to construct a successful promoter-XylE ligation, since we believe that the strand annealed promoter was of bad quality |
|} | |} | ||
| Line 261: | Line 268: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>midi | + | <li>midi prep registry obtained J23101</li> |
<li>gel purification of XbaI-His-GFP-flag-TEVs</li> | <li>gel purification of XbaI-His-GFP-flag-TEVs</li> | ||
</ul> | </ul> | ||
| Line 307: | Line 314: | ||
<ul> | <ul> | ||
<li>gel purification of overnight ligation</li> | <li>gel purification of overnight ligation</li> | ||
| - | <li>transformation of E.coli with overnight ligation product and selection by plating</li> | + | <li>transformation of ''E.coli'' with overnight ligation product and selection by plating</li> |
<li>gel purification of TEVs-linker-XylE-SpeI construct </li> | <li>gel purification of TEVs-linker-XylE-SpeI construct </li> | ||
</ul> | </ul> | ||
| Line 319: | Line 326: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>Gel analysis and gel purification of fusion | + | <li>Gel analysis and gel purification of fusion XylE protein</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 327: | Line 334: | ||
'''Monday, 23-Aug''' | '''Monday, 23-Aug''' | ||
| - | * Set up overnight cultures for midi | + | * Set up overnight cultures for midi prep |
* Gel separation of linker-XylE-Spe DNA <font color = red>'''FAILED'''</font> | * Gel separation of linker-XylE-Spe DNA <font color = red>'''FAILED'''</font> | ||
* PCR reaction for extension of His-GFP-Flag | * PCR reaction for extension of His-GFP-Flag | ||
| - | * Catechol assay on E.coli transformed with overnight ligated J23101, XylE and pSB1C3. <font color = red>'''FAILED'''</font> | + | * Catechol assay on ''E.coli'' transformed with overnight ligated J23101, XylE and pSB1C3. <font color = red>'''FAILED'''</font> |
'''Tuesday, 24th-Aug''' | '''Tuesday, 24th-Aug''' | ||
| - | * Midi | + | * Midi prepped registry obtained J23101. A yield of 130ng/μl of promoter was obtained. The promoter is in a BioBrick vector called J62001. The promoter is upstream of RFP gene. |
*the vector carrying the promoter was digested with SpeI and PstI, while XylE gene was digested with XbaI and PstI. | *the vector carrying the promoter was digested with SpeI and PstI, while XylE gene was digested with XbaI and PstI. | ||
* The promoter and the XylE gene were gel purified. | * The promoter and the XylE gene were gel purified. | ||
* A reaction between the promoter(still on vector) and XylE was set on for overnight ligation | * A reaction between the promoter(still on vector) and XylE was set on for overnight ligation | ||
* PCR purification of GFP2 -> GFP construct ready for full fusion protein. | * PCR purification of GFP2 -> GFP construct ready for full fusion protein. | ||
| - | * Gel purification of XylE lost DNA along the way. Thus PCR XylE1 with gradient for temperature scanning (taq): PCR round 1 included | + | * Gel purification of XylE lost DNA along the way. Thus PCR XylE1 with gradient for temperature scanning (taq): PCR round 1 included 62°C and only rev primer to create pool of successful extensions with with rev primer (60-62°C). PCR round 2 with Fwd primer and temperature scale (68-68°C, 72-74°C). |
'''Wednesday, 25th-Aug''' | '''Wednesday, 25th-Aug''' | ||
| - | Performed gel analysis on the purified XylE and J23101 to obtain ratios for ligation. First gel was scrapped as it produced | + | Performed gel analysis on the purified XylE and J23101 to obtain ratios for ligation. First gel was scrapped as it produced appalling (explanation for Nick: really bad) results, 2nd gel run was successful. |
| - | * Performed a ligation reaction between the vector containing J23101, and XylE(one on bench and one overnight one). | + | * Performed a ligation reaction between the vector containing J23101, and XylE (one on bench and one overnight one). |
| - | * Transformation of the new plasmid into competent E.coli. Successfully transformed colonies can be selected for by loss of RFP expression. | + | * Transformation of the new plasmid into competent ''E.coli''. Successfully transformed colonies can be selected for by loss of RFP expression. |
* XylE-1 PCR with temperature cascade. Gel analysis and purification. | * XylE-1 PCR with temperature cascade. Gel analysis and purification. | ||
'''Thursday, 26th-Aug''' | '''Thursday, 26th-Aug''' | ||
| - | * White colonies from the promoter-XylE transformed E.coli were picked and transferred to new amp plates. One is the replica plate and the other is the catechol assay plate. | + | * White colonies from the promoter-XylE transformed ''E.coli'' were picked and transferred to new amp plates. One is the replica plate and the other is the catechol assay plate. |
* XylE-1, two rounds of PCR/purification were run to obtain a sufficiently clear band. An additional PCR run for XylE-1 was discarded afterwards. | * XylE-1, two rounds of PCR/purification were run to obtain a sufficiently clear band. An additional PCR run for XylE-1 was discarded afterwards. | ||
'''Friday, 27th-Aug''' | '''Friday, 27th-Aug''' | ||
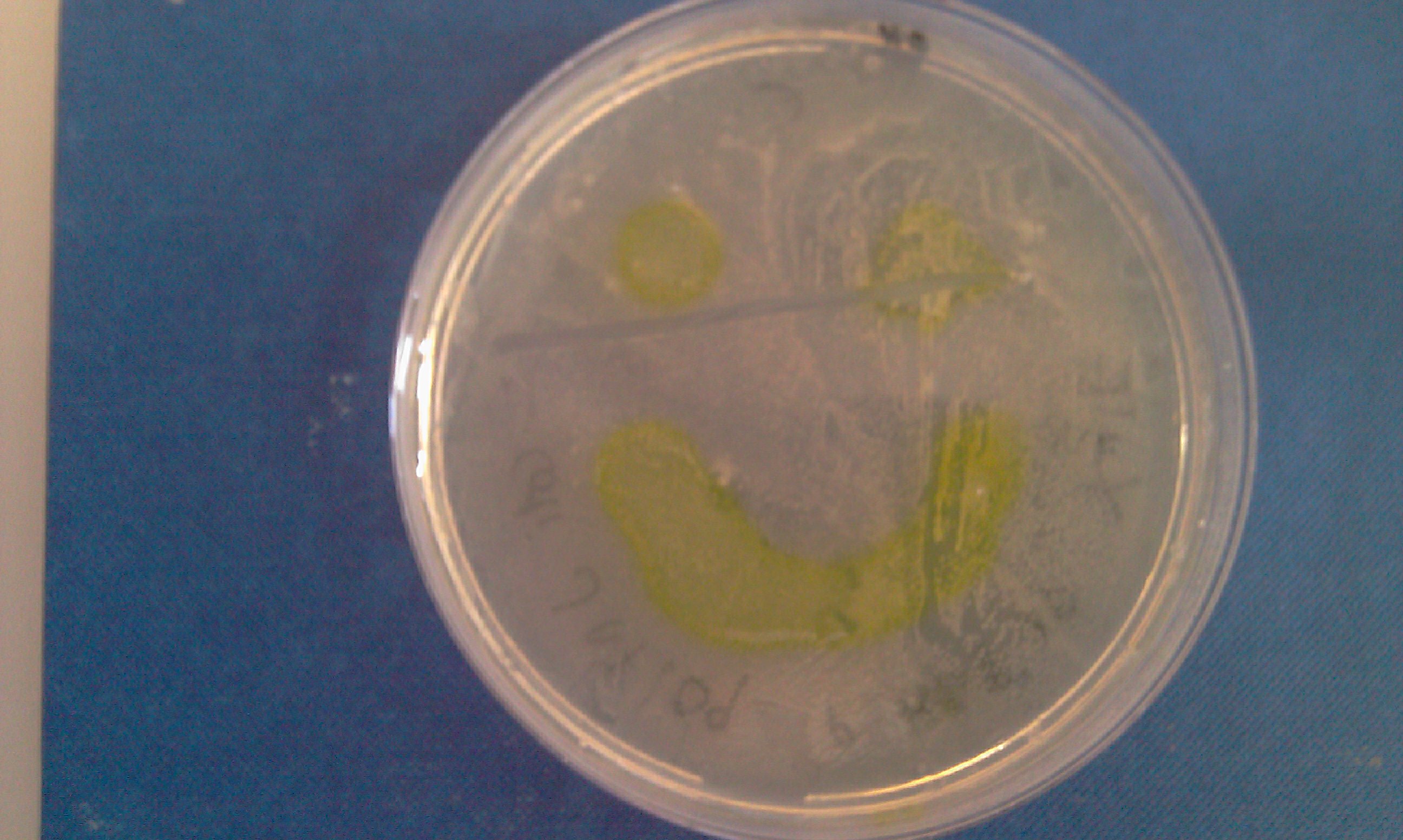
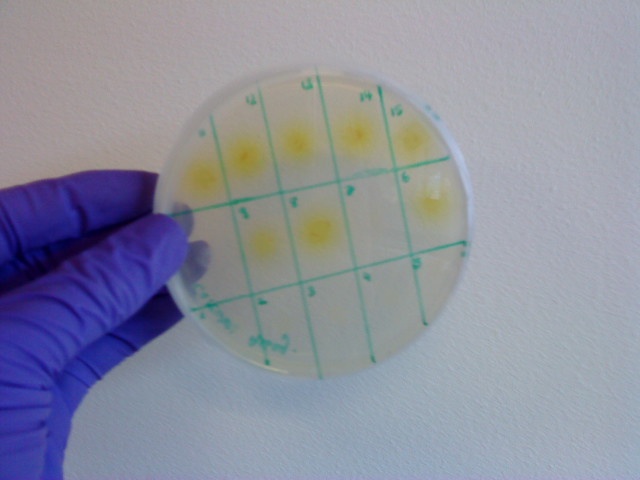
| - | * Catechol assay performed on promoter-XylE transformed E.coli. <font color = green>'''SUCCESSFULL'''</font> | + | * Catechol assay performed on promoter-XylE transformed ''E.coli''. <font color = green>'''SUCCESSFULL'''</font> |
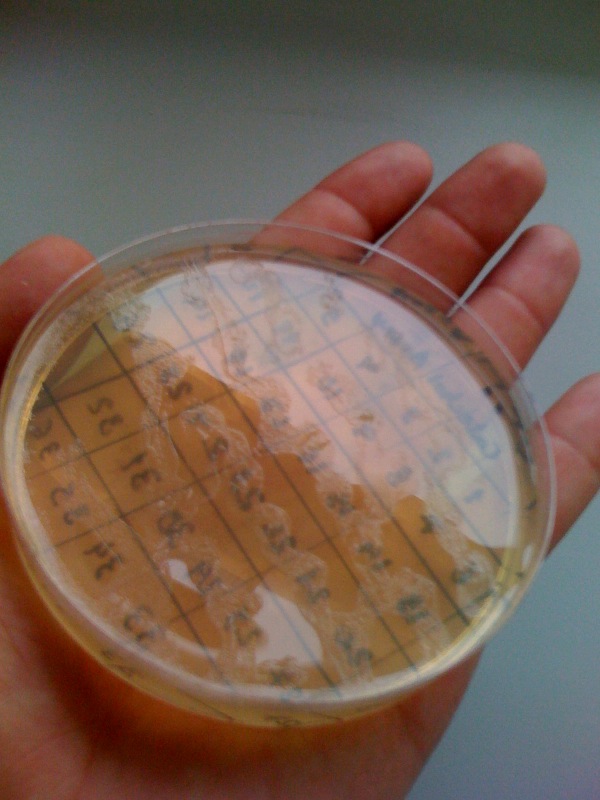
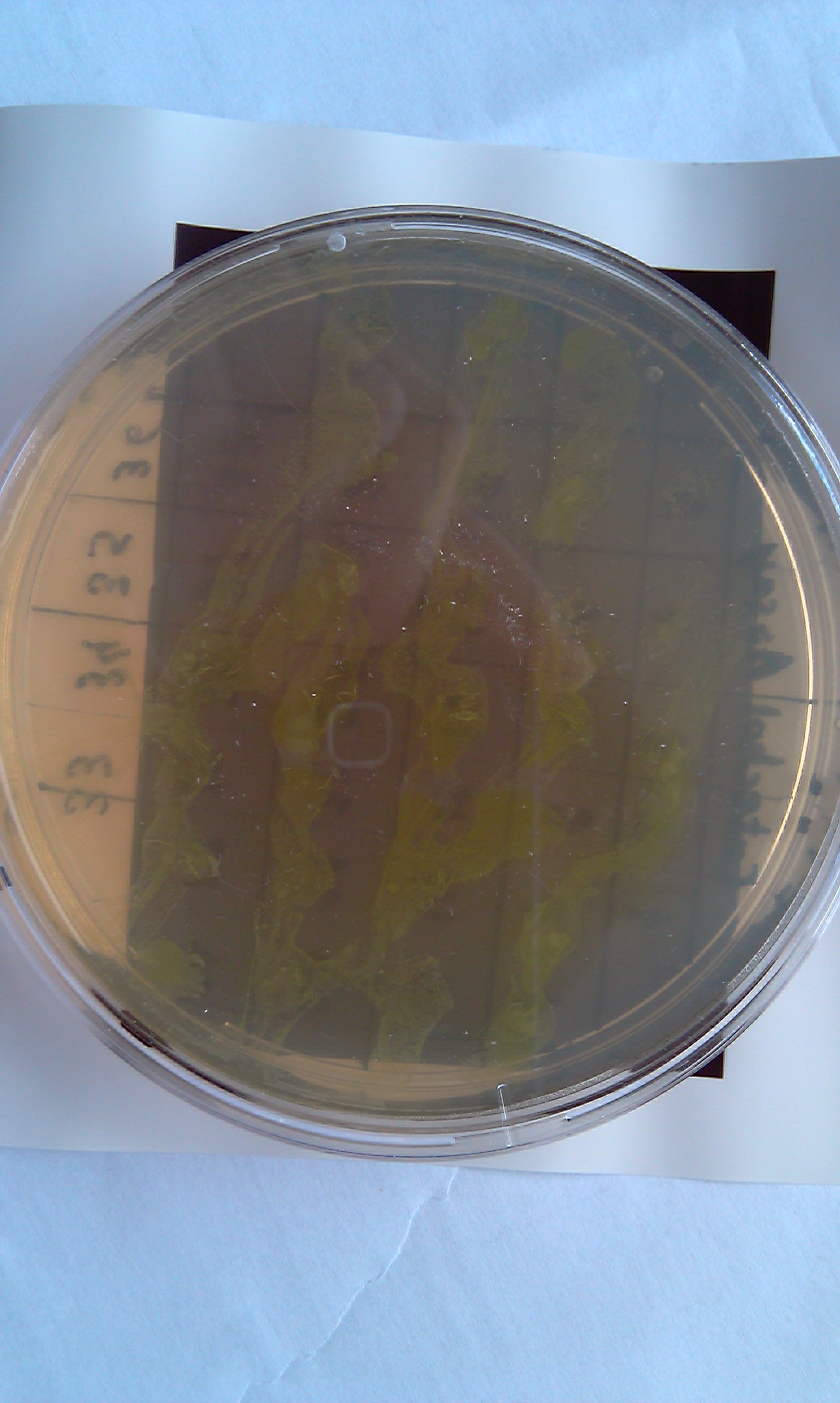
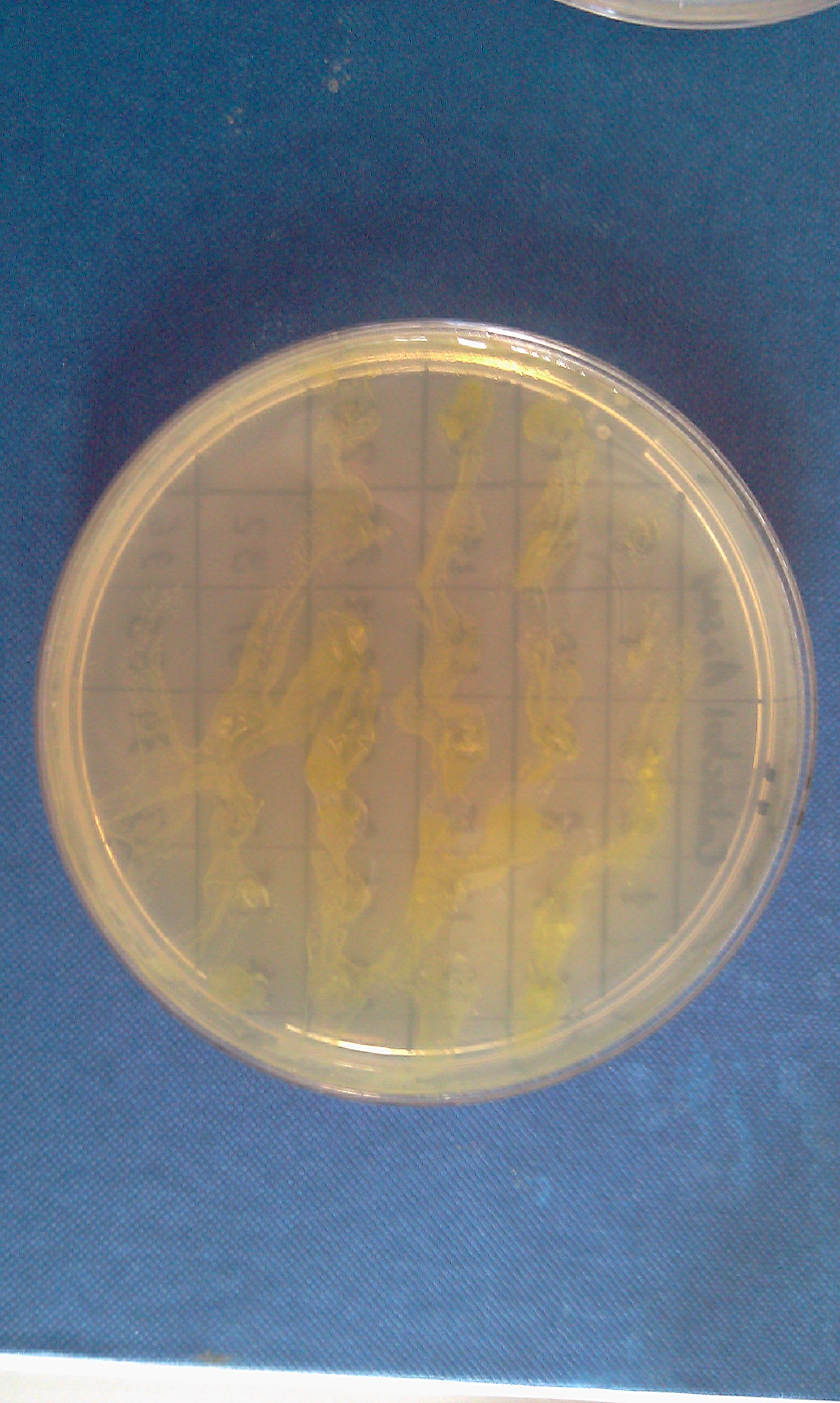
[[Image:IC_Catechol_Assay_before.jpg|thumb|left|Plate before adding catechol assay]] [[Image:IC_Catechol_assay_after_(27-8).jpg|thumb|center|After addition of catechol colonies turn yellow-orange in seconds!!]] | [[Image:IC_Catechol_Assay_before.jpg|thumb|left|Plate before adding catechol assay]] [[Image:IC_Catechol_assay_after_(27-8).jpg|thumb|center|After addition of catechol colonies turn yellow-orange in seconds!!]] | ||
* XylE-2 PCR and gel-purification cycles (2x) to obtain clear band. XylE-2 is now ready for assembly of the GFP-XylE fusion protein. | * XylE-2 PCR and gel-purification cycles (2x) to obtain clear band. XylE-2 is now ready for assembly of the GFP-XylE fusion protein. | ||
| Line 365: | Line 372: | ||
'''Sunday, 29th-Aug''' | '''Sunday, 29th-Aug''' | ||
| - | * Gel analysis of the attempted annealing reaction of GFP-2 XylE-2 showed unsufficiently clear bands for gel-purification. A new reaction is being prepared: 10 rounds of annealing PCR, followed by addition of primers (5' primer for GFP-2 and 3' primer for XylE-2) in order to introduce an amplification step in the reaction. --- Following | + | * Gel analysis of the attempted annealing reaction of GFP-2 XylE-2 showed unsufficiently clear bands for gel-purification. A new reaction is being prepared: 10 rounds of annealing PCR, followed by addition of primers (5' primer for GFP-2 and 3' primer for XylE-2) in order to introduce an amplification step in the reaction. --- Following Kirstin's advice, we are discarding this reaction and wait for the arrival of new primers for XylE-2 (5' + TEV) and GFP-2 (3' +TEV). |
|} | |} | ||
| Line 396: | Line 403: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>mini | + | <li>mini prep of promoter-XylE transformed ''E.coli''</li> |
<li>design of reverse primer for GFP to add the corrected TEV sequence to the construct.</li> | <li>design of reverse primer for GFP to add the corrected TEV sequence to the construct.</li> | ||
</ul> | </ul> | ||
| Line 403: | Line 410: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>midi | + | <li>midi prep of promoter-XylE transformed ''E.coli''</li> |
<li>restriction digestion of pSB1C3+terminator vector DNA</li> | <li>restriction digestion of pSB1C3+terminator vector DNA</li> | ||
<li>restriction digestion of promoter-XylE DNA </li> | <li>restriction digestion of promoter-XylE DNA </li> | ||
| Line 436: | Line 443: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>gel analysis of mini | + | <li>gel analysis of mini prepped promoter-XylE</li> |
<li>cross-check for primer design (GFP-TEV-2) and ordering</li> | <li>cross-check for primer design (GFP-TEV-2) and ordering</li> | ||
</ul> | </ul> | ||
| Line 474: | Line 481: | ||
'''Tuesday, 31st-Aug''' | '''Tuesday, 31st-Aug''' | ||
| - | * Midi prep of colony 24 for XylE-J23101 the final concentration was ~100ng/ | + | * Midi prep of colony 24 for XylE-J23101 the final concentration was ~100ng/μl which wasnt so great but Chris says the protocol produces very poor yields. |
* We are performing the next building step of our vector. PSB1C3 containing terminator B0014 was cut with EcoRI and XbaI. The insert was cut with EcoRI and SpeI and both were incubated for 1.5hrs. Wolf is now running a gel to purify out the insert via gel purification and perform a PCR purification on the vector. | * We are performing the next building step of our vector. PSB1C3 containing terminator B0014 was cut with EcoRI and XbaI. The insert was cut with EcoRI and SpeI and both were incubated for 1.5hrs. Wolf is now running a gel to purify out the insert via gel purification and perform a PCR purification on the vector. | ||
*Advisors have decided it's best not to use Jeremy's tagged XylE due to the 93% difference. Kirsten will be tagging the registry XylE and we shall purify and assay with that instead. | *Advisors have decided it's best not to use Jeremy's tagged XylE due to the 93% difference. Kirsten will be tagging the registry XylE and we shall purify and assay with that instead. | ||
| Line 481: | Line 488: | ||
'''Thursday, 2nd-Sept''' | '''Thursday, 2nd-Sept''' | ||
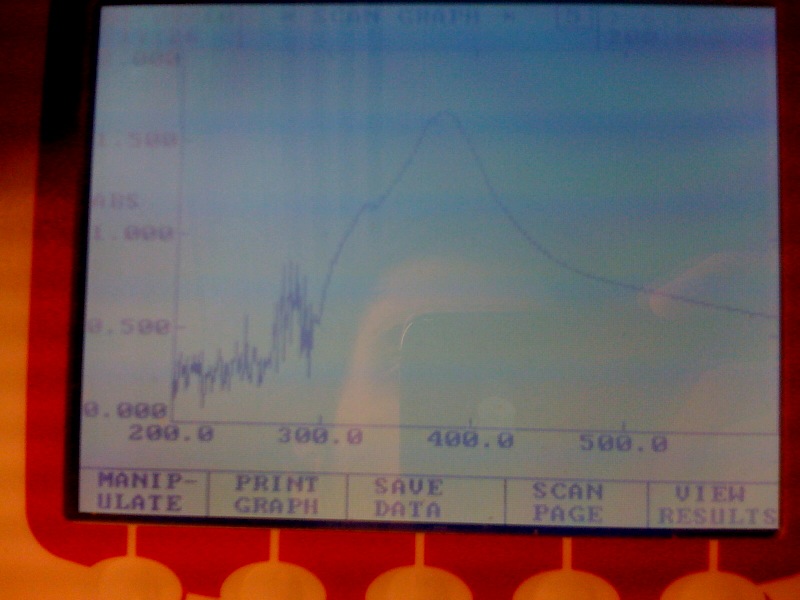
[[Image:IC_200-600nm_spectr.jpg|thumb||center|300px|Spectra of XylE transformed E.coli after addition of catechol assay. The '''broad peak around 380nm wavelength''' arises is due to the presence of the product of the enzymatic reaction involving pyrocatechol and XylE enzyme. This peak if absent if a culture of XylE transformed cells are measured without the addition of catechol]] | [[Image:IC_200-600nm_spectr.jpg|thumb||center|300px|Spectra of XylE transformed E.coli after addition of catechol assay. The '''broad peak around 380nm wavelength''' arises is due to the presence of the product of the enzymatic reaction involving pyrocatechol and XylE enzyme. This peak if absent if a culture of XylE transformed cells are measured without the addition of catechol]] | ||
| - | *Spectrophotometry experiments with XylE transformed E.coli in LB medium (M9 culture was contaminated) reveiled the followings: On catechol assay of the trasformed cells, the '''positive yellow output can be quantitively measured by a broad peak at 380nm.''' | + | *Spectrophotometry experiments with XylE transformed ''E.coli'' in LB medium (M9 culture was contaminated) reveiled the followings: On catechol assay of the trasformed cells, the '''positive yellow output can be quantitively measured by a broad peak at 380nm.''' |
| - | * | + | *Transformation of competent ''E.coli'' cells with promoter-XylE-terminator pSB1C3vector. |
'''Friday, 3rd-Sept''' | '''Friday, 3rd-Sept''' | ||
| Line 521: | Line 528: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>PCR extension of | + | <li>PCR extension of PVeg promoter </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 527: | Line 534: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>Gel purification of extended | + | <li>Gel purification of extended PVeg promoter</li> |
<li>Annealing PCR for GFP-XylE fusion protein</li> | <li>Annealing PCR for GFP-XylE fusion protein</li> | ||
</ul> | </ul> | ||
| Line 534: | Line 541: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>vector with insert tranformed into E.coli and plate to select for successful transformation</li> | + | <li>vector with insert tranformed into ''E.coli'' and plate to select for successful transformation</li> |
</ul> | </ul> | ||
</td> | </td> | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>prepare overnight cultures of pVeg-RBS transformed E.coli </li> | + | <li>prepare overnight cultures of pVeg-RBS transformed ''E.coli'' </li> |
<li>Gel-analysis of amplification-PCR for XylE-2</li> | <li>Gel-analysis of amplification-PCR for XylE-2</li> | ||
</ul> | </ul> | ||
| Line 546: | Line 553: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>midi | + | <li>midi prep of PVeg-RBS vector</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 563: | Line 570: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>restriction digestion and gel purification of thr extended | + | <li>restriction digestion and gel purification of thr extended pVeg promoter</li> |
| - | <li>overnight ligation of the cut | + | <li>overnight ligation of the cut pVeg promoter in a vector </li> |
<li>Gel analysis of GFP-XylE annealing PCR - unsuccessful reaction</li> | <li>Gel analysis of GFP-XylE annealing PCR - unsuccessful reaction</li> | ||
</ul> | </ul> | ||
| Line 589: | Line 596: | ||
'''Monday, 6th-Sept''' | '''Monday, 6th-Sept''' | ||
| - | * PCR extension of | + | * PCR extension of pVeg promoter: EcoRI---pVeg---RBS-SpeI |
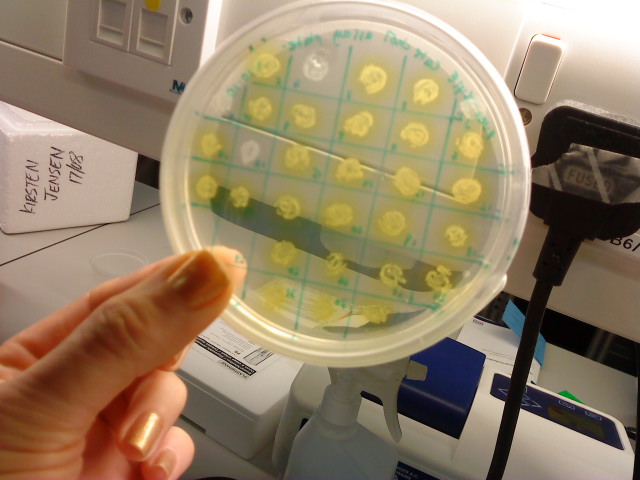
* I performed a catechol assay on the picked transformed colonies to deduce which ones were successfully transformed with the insert plus vector. 1-5 7 and 10 failed to turn yellow (1-5 were background controls)leaving 8 yellow colonies. | * I performed a catechol assay on the picked transformed colonies to deduce which ones were successfully transformed with the insert plus vector. 1-5 7 and 10 failed to turn yellow (1-5 were background controls)leaving 8 yellow colonies. | ||
| Line 600: | Line 607: | ||
* Mini prep of the overnight Colony 8 culture (by Wolf) | * Mini prep of the overnight Colony 8 culture (by Wolf) | ||
* Midi prep of overnight culture | * Midi prep of overnight culture | ||
| - | * Gel purification of PCR product EcoRI-- | + | * Gel purification of PCR product EcoRI--pVeg--RBS--SpeI |
| - | * Overnight restriction digestion of EcoRI-- | + | * Overnight restriction digestion of EcoRI--pVeg--RBS--SpeI with SpeI |
* Started data analysis of plate assay | * Started data analysis of plate assay | ||
* Annealing PCR reaction included 2 samples without and one with additional primers (XbaI-His-GFP Fwd, SpeI-XylE Rev). While the primer-including reaction did not show any clearly identifiable bands, the others showed clear, if very weak bands at 800 and 1000bp, which represent the GFP and XylE constructs respectively. No band was identifiable at the 1.7 kb range, which would have indicated a successful annealing reaction. However, problems with the lense of the gel-analyser were only discovered later to have severely reduced the band brightness. Potentially a PCR-reaction with appropriate primers to amplify an annealed product(XbaI-His-GFP Fwd, SpeI-XylE Rev), could have been successful. However the reaction mixture was disposed of before this became clear. | * Annealing PCR reaction included 2 samples without and one with additional primers (XbaI-His-GFP Fwd, SpeI-XylE Rev). While the primer-including reaction did not show any clearly identifiable bands, the others showed clear, if very weak bands at 800 and 1000bp, which represent the GFP and XylE constructs respectively. No band was identifiable at the 1.7 kb range, which would have indicated a successful annealing reaction. However, problems with the lense of the gel-analyser were only discovered later to have severely reduced the band brightness. Potentially a PCR-reaction with appropriate primers to amplify an annealed product(XbaI-His-GFP Fwd, SpeI-XylE Rev), could have been successful. However the reaction mixture was disposed of before this became clear. | ||
'''Wednesday, 8th Sep''' | '''Wednesday, 8th Sep''' | ||
| - | * A second PCR extension of | + | * A second PCR extension of pVeg promoter to introduce the RBS and cut sites. It was also gel purified and stored in gel lumbs in the freezer. (maybe needed later so keep in mind). |
* Overnight restriction digestion completed. Then, it was run on the gel to check if it worked and then gel purified again. | * Overnight restriction digestion completed. Then, it was run on the gel to check if it worked and then gel purified again. | ||
| - | * Vector PSBI-C3 was digested to remove the terminator and make it sticky for the insert ( | + | * Vector PSBI-C3 was digested to remove the terminator and make it sticky for the insert (pVeg-RBS). Then it was run on the gel to check if restriction has worked, but the gel didn't run far enough to determine easily between undigested and digested vector. |
* For further annealing reactions for GFP-XylE constructs additional XylE(2) template was required -> amplification PCR for XylE(2). | * For further annealing reactions for GFP-XylE constructs additional XylE(2) template was required -> amplification PCR for XylE(2). | ||
'''Thursday, 9th Sep''' | '''Thursday, 9th Sep''' | ||
* Gel with digested and undigested vector PSBI-C3 was run and then the digested vector was purified. | * Gel with digested and undigested vector PSBI-C3 was run and then the digested vector was purified. | ||
| - | * Gel was run to determine the DNA concentration ratio for the ligation of PSBI-C3 and | + | * Gel was run to determine the DNA concentration ratio for the ligation of PSBI-C3 and pVeg-RBS. |
* Vector PSBI-C3 was dephosphorylated. | * Vector PSBI-C3 was dephosphorylated. | ||
| - | * Ligation of PSBI-C3 and | + | * Ligation of PSBI-C3 and pVeg-RBS has been set up overnight. |
* Gel-analysis and gel-purification of the XylE(2) amplification PCR product. | * Gel-analysis and gel-purification of the XylE(2) amplification PCR product. | ||
| - | * The transformation of E.Coli with | + | * The transformation of ''E.Coli'' with pVeg-RBS in PSBI-C3 and PSBI-C3 by itself (to check see how successful dephosphoryaltion of PSBI-C3 was and estimate the percentage of bacteria that contain the insert) was completed |
| - | * Concentration of the midi prep of J23101-XylE-B0014 was determined to be ~600ng/ | + | * Concentration of the midi prep of J23101-XylE-B0014 was determined to be ~600ng/μl (using new protocol) |
* Kyasha kindly digested my midi with EcoRI and SpeI and performed a gel analysis. The results show a potential additional plasmid contaminating my midi however the concentration of DNA was extremely high. NB Chris said that it could be sheared DNA from a midi prep step. | * Kyasha kindly digested my midi with EcoRI and SpeI and performed a gel analysis. The results show a potential additional plasmid contaminating my midi however the concentration of DNA was extremely high. NB Chris said that it could be sheared DNA from a midi prep step. | ||
| - | * The midi prepped plasmid was transformed into testing E.coli strain TOP10. | + | * The midi prepped plasmid was transformed into testing ''E.coli'' strain TOP10. |
'''Friday, 10 sep''' | '''Friday, 10 sep''' | ||
| - | The transformation was a SUCCESS. 2x replica plates were made plate 1# 1-6 plate #2 6-11; colony 6 and colony 9 of plates 1 and 2 respectively were transfered into a 5ml liquid culture + | + | The transformation was a SUCCESS. 2x replica plates were made plate 1# 1-6 plate #2 6-11; colony 6 and colony 9 of plates 1 and 2 respectively were transfered into a 5ml liquid culture + 5μl CmR. These will later be turned into glycerol stocks. After the replica plates have grown up mini preps on a number of colonies shall be performed - this hopefully will eliminate the contaminating plasmid DNA. This will be followed by a midi prep. |
'''Saturday, 11 Sep''' | '''Saturday, 11 Sep''' | ||
| - | The transformation of E.Coli with PSB1-C3 with insert did not work :( | + | The transformation of ''E.Coli'' with PSB1-C3 with insert did not work :( |
|} | |} | ||
| Line 667: | Line 674: | ||
<td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | <td style="background-color:#eeeeee;height:100px;width:200px;text-align:left;color:#555555;"> | ||
<ul> | <ul> | ||
| - | <li>midi | + | <li>midi prep of pVeg-RBS vector</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 726: | Line 733: | ||
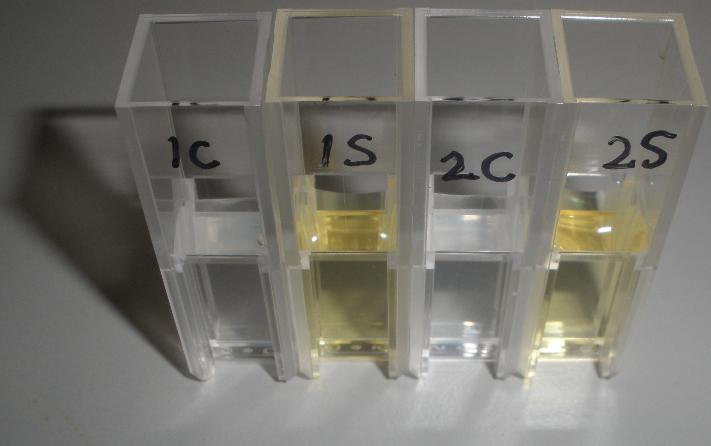
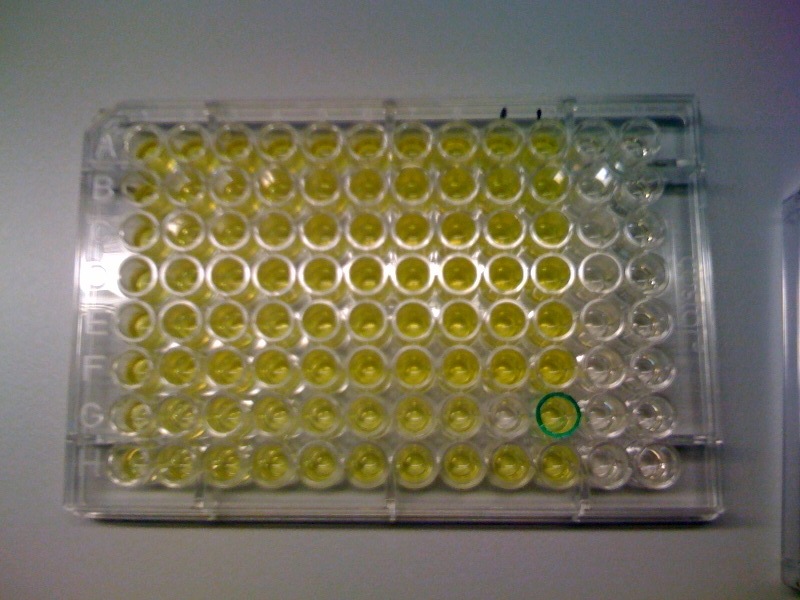
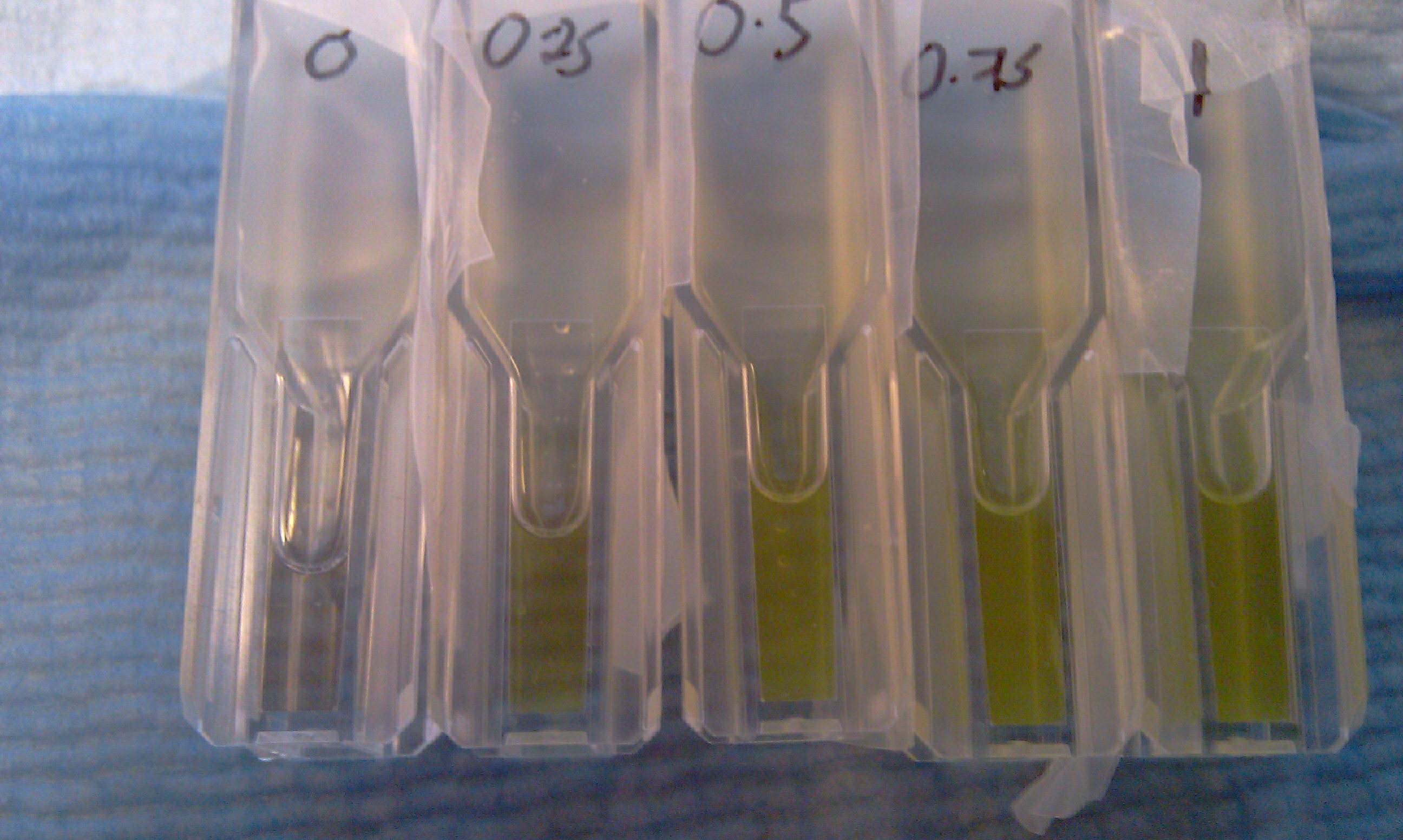
[[Image:CS1.JPG|thumb|right|300px|test confirming that yellow product of XylE enzmymatic reaction is leaks back from the cytosol into the solution.1C and 2C is the XylE producing cells after centrifuging and redissolving of the pellet and 1S and 2S is the supernatant after centrifuging]] | [[Image:CS1.JPG|thumb|right|300px|test confirming that yellow product of XylE enzmymatic reaction is leaks back from the cytosol into the solution.1C and 2C is the XylE producing cells after centrifuging and redissolving of the pellet and 1S and 2S is the supernatant after centrifuging]] | ||
*assay on plate reader of: | *assay on plate reader of: | ||
| - | **0-0.75 initial O.D. of transformed E.coli | + | **0-0.75 initial O.D. of transformed ''E.coli'' |
**0-1mM initial catechol concentration | **0-1mM initial catechol concentration | ||
| - | The assay was carried out with E.coli, top ten spcies, transformed with J23101-XylE-B0014 in pSB1C3 vector.The overnight culture wastransfered in new medium this morning for 4 hrs before assaying. LB medium was used for dilutions and blank. Catechol was diluted in ddH2O. | + | The assay was carried out with ''E.coli'', top ten spcies, transformed with J23101-XylE-B0014 in pSB1C3 vector.The overnight culture wastransfered in new medium this morning for 4 hrs before assaying. LB medium was used for dilutions and blank. Catechol was diluted in ddH2O. |
[[Image:Catechol assay test 2.xls|Data analysis of the assay]] | [[Image:Catechol assay test 2.xls|Data analysis of the assay]] | ||
*Mini preps of XylE 8,10 colonies and 3K3;102 151 260 colonies. Analytical digests were performed using E+S on XylE, E+S AccI and HindIII+E for 3K3 vector respectively. The band patterns correctly identified the vector to be 3k3. Colony 10 of XylE and Colony 260 of the 3k3 vector were set up direct into 100ml LB and so will require til 12pm for midi prepping on weds. Nick performed a second plate reader assay to determine the minimal concentration of Catechol to use for assays. I took 990ul M9 culture containing colony 10 and 10ul catechol was added. This was incubated for 10 mins and then spun down. The supernatant was removed into a cuvette and the cells resuspended in M9 salts. The ODs were then read on the spectrophotometer at 380 and 600nm. It was found that control cuvette of M9 salts had a miniscule reading. M9 + cells ~0.007. Supernatant + cells were ~ 2.5 therefore we could deduce that the coloured catechol breakdown product is exported out of the cells. | *Mini preps of XylE 8,10 colonies and 3K3;102 151 260 colonies. Analytical digests were performed using E+S on XylE, E+S AccI and HindIII+E for 3K3 vector respectively. The band patterns correctly identified the vector to be 3k3. Colony 10 of XylE and Colony 260 of the 3k3 vector were set up direct into 100ml LB and so will require til 12pm for midi prepping on weds. Nick performed a second plate reader assay to determine the minimal concentration of Catechol to use for assays. I took 990ul M9 culture containing colony 10 and 10ul catechol was added. This was incubated for 10 mins and then spun down. The supernatant was removed into a cuvette and the cells resuspended in M9 salts. The ODs were then read on the spectrophotometer at 380 and 600nm. It was found that control cuvette of M9 salts had a miniscule reading. M9 + cells ~0.007. Supernatant + cells were ~ 2.5 therefore we could deduce that the coloured catechol breakdown product is exported out of the cells. | ||
| Line 734: | Line 741: | ||
Piotr: | Piotr: | ||
* Mini prep of 1 sample (6 samples lost due to mistake) of GFP-Xyle fusion protein and its digestion with Spe and Xba (gel to be run on the next day) | * Mini prep of 1 sample (6 samples lost due to mistake) of GFP-Xyle fusion protein and its digestion with Spe and Xba (gel to be run on the next day) | ||
| - | * Preparing E.Coli colonies for the next day for mini prep to be redone | + | * Preparing ''E.Coli'' colonies for the next day for mini prep to be redone |
* PCR of 6 samples of GFP-Xyle fusion | * PCR of 6 samples of GFP-Xyle fusion | ||
| Line 755: | Line 762: | ||
'''Friday 17th September''' | '''Friday 17th September''' | ||
| - | *Catechol assay of transformed E.coli Top ten(J23101-XylE-terminator) in M9 medium + data analysis | + | *Catechol assay of transformed ''E.coli'' Top ten(J23101-XylE-terminator) in M9 medium + data analysis |
**[[Image:M9 catechol concentrations assay alternative.xls|Data analysis]] | **[[Image:M9 catechol concentrations assay alternative.xls|Data analysis]] | ||
**[[Image:Catechol assay in M9.xls| alternative]] | **[[Image:Catechol assay in M9.xls| alternative]] | ||
| Line 770: | Line 777: | ||
Performed ligation again. Errors in the sequencing we received back regarding J23101-XylE-B0014. | Performed ligation again. Errors in the sequencing we received back regarding J23101-XylE-B0014. | ||
One was a log error (in XylE) phew. The other is in PSB1C3 out of the scar site and should be OK. | One was a log error (in XylE) phew. The other is in PSB1C3 out of the scar site and should be OK. | ||
| - | Chris is currently assembling the | + | Chris is currently assembling the pVeg promoter+ RBS in a vector. Once he has finished this, I will combine XylE-GFP and XylE with this promoter for comparison characterization with J23101 (in ''E.coli'' and ''Bacillus'') |
'''Tues 21st Sept''' | '''Tues 21st Sept''' | ||
| Line 794: | Line 801: | ||
*to be followed by a digest. XbaI Pst for FWD (not known for the reverse currently) to insert XylE after it. | *to be followed by a digest. XbaI Pst for FWD (not known for the reverse currently) to insert XylE after it. | ||
*Ligation with XylE and PSB1C3-B0014 possibly. | *Ligation with XylE and PSB1C3-B0014 possibly. | ||
| - | *Transformation of XylE-PSB1C3-B0014 and of XylEe-3k3 into E.coli | + | *Transformation of XylE-PSB1C3-B0014 and of XylEe-3k3 into ''E.coli'' later |
* Hi Nick, hope revision is going well. Love from Team XylE xox | * Hi Nick, hope revision is going well. Love from Team XylE xox | ||
|} | |} | ||
| Line 805: | Line 812: | ||
* Successfull transformations: now have XylE-PSB1C3-B0014 (no promoter) and 3K3-J23101-XylE-B0014. No background plate colonies..many colonies for 3K3-XylE and a few for PSB1C3-XylE. Colonies have been replica plated (plus catechol assay plate for 3K3-XylE) and set up for mini prepping tomorrow. | * Successfull transformations: now have XylE-PSB1C3-B0014 (no promoter) and 3K3-J23101-XylE-B0014. No background plate colonies..many colonies for 3K3-XylE and a few for PSB1C3-XylE. Colonies have been replica plated (plus catechol assay plate for 3K3-XylE) and set up for mini prepping tomorrow. | ||
* Sumo tagged XylE will be given to us by Kirsten at some point this week and we will have to purify it. This can then be used for in vitro testing and comparisons made with the linker-XylE protein. | * Sumo tagged XylE will be given to us by Kirsten at some point this week and we will have to purify it. This can then be used for in vitro testing and comparisons made with the linker-XylE protein. | ||
| - | * pVeg and ComC DE promoter FWD can be added into the promoter-less construct after it has been midi | + | * pVeg and ComC DE promoter FWD can be added into the promoter-less construct after it has been midi prepped and digested accordingly. ComC DE promoter REV can be added onto the reverse XylE construct (PCR'd by Piotr) AFTER it has been put into PSB1C3 and is more manageable. Kirsten is taking care of the GFP-XylE fusion for now, including putting the pVeg promoter plus RBS infront of it (proving difficult.) We still need to receive LacI promoter to create the inducible test version of the fusion XylE-GFP protein. |
'''Tues 28th Sept''' | '''Tues 28th Sept''' | ||
| Line 827: | Line 834: | ||
* Today I peformed ligations for ComCDE FWD promoter and PSB1C3-XylE-B0014 and also PSB1C3-pVeg and XylE-B0014 they will be left overnight and transformed by chris tmro thaaanks. | * Today I peformed ligations for ComCDE FWD promoter and PSB1C3-XylE-B0014 and also PSB1C3-pVeg and XylE-B0014 they will be left overnight and transformed by chris tmro thaaanks. | ||
| - | * I ran the PCR Kirill performed yday to get out blunt ended J23101-XylE on a gel; there was no template DNA! so I've just set up the reaction again... hopefully i'll purify it later for ligation into the final Spec vector that kyasha has maaaaade :))) then we can test XylE in Bacillus. | + | * I ran the PCR Kirill performed yday to get out blunt ended J23101-XylE on a gel; there was no template DNA! so I've just set up the reaction again... hopefully i'll purify it later for ligation into the final Spec vector that kyasha has maaaaade :))) then we can test XylE in ''Bacillus''. |
* I'm currently performing Midi's of my J23101-XylE-B0014 (running low) and of GFP-XylE (3rd time!) | * I'm currently performing Midi's of my J23101-XylE-B0014 (running low) and of GFP-XylE (3rd time!) | ||
|} | |} | ||
| Line 868: | Line 875: | ||
'''Monday 4th Oct''' | '''Monday 4th Oct''' | ||
| - | Did a replica plate and catechol assay plate/ minis 2 6 7 9 12 16. (Successful | + | Did a replica plate and catechol assay plate/ minis 2 6 7 9 12 16. (Successful pVeg-XylE transformation) |
'''Tues 5th Oct''' | '''Tues 5th Oct''' | ||
| Line 877: | Line 884: | ||
* Will perform the same assays already performed by Nick with the 3k3-XylE constructs to compare results. | * Will perform the same assays already performed by Nick with the 3k3-XylE constructs to compare results. | ||
* Just realised Im working with background colony 16!! disregard this after the gel run! | * Just realised Im working with background colony 16!! disregard this after the gel run! | ||
| - | * I'm currently keeping my undigested | + | * I'm currently keeping my undigested pVeg-XylE minis in Florian's DNA box (orange lids and labels) |
| Line 885: | Line 892: | ||
* Niiiiiiiiiiiiiiick!!! you left the country! you were too ugly. good luck babe! | * Niiiiiiiiiiiiiiick!!! you left the country! you were too ugly. good luck babe! | ||
| - | * Midi prepped | + | * Midi prepped pVeg-XylE-B0014 in PSB1C3; will digest out the insert (E+S) and put it into 3K3(E+X). Then both the midi (in PSB1C3) and the ligation 3k3-pVeg-XylE-B0014 can be transformed into TOP10. J23101-XylE-B0014 in 3K3 Vector also needs to be put into TOP10 currently only the PSB1C3 version is in TOP10. We need the rest for comparison testing as TOP10 is the more widely used testing strain. |
| - | * Florian kindly digested | + | * Florian kindly digested pVeg-XylE-B0014-C3 with E+S |
| - | * Ran PCR to get out blunt | + | * Ran PCR to get out blunt pVeg-XylE-B0014 using primers. Did 3 reactions at variations around 60 degrees. |
'''Friday 8th Oct''' | '''Friday 8th Oct''' | ||
| - | * Ran gel analysis of my PCRs 1, 2 and 3* appear to have worked. Samples 1 and 3* show stronger bands and so should be used in the next ligation step into the SPEC vector (which kyasha will be doing) then transformation into Bacillus (meeee.) | + | * Ran gel analysis of my PCRs 1, 2 and 3* appear to have worked. Samples 1 and 3* show stronger bands and so should be used in the next ligation step into the SPEC vector (which kyasha will be doing) then transformation into ''Bacillus'' (meeee.) |
| - | * Chris is purifying digested | + | * Chris is purifying digested pVeg-XylE-B0014-C3 to get the insert out and this will then be ran on a gel alongside cut (E+X) 3k3 to determine a ratio for ligation. |
| - | * Meeting today at 4pm, no one is here!! it'll be me | + | * Meeting today at 4pm, no one is here!! it'll be me Florian, Ben and Piotr attending..yikes |
'''Sunday 10th of the 10th of the 2010!!!!! and im in LAB ''' | '''Sunday 10th of the 10th of the 2010!!!!! and im in LAB ''' | ||
| - | * I did ligations of | + | * I did ligations of pVeg-XylE into 3k3 and of Reverse-XylE into the digested C-Tev LacI vector. |
| - | * I also set up assay cultures for monday 2x | + | * I also set up assay cultures for monday 2x pVegXE 2x XE-C3 2x XE-3K3 and 2x GFP-XylE!!! |
|} | |} | ||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| Line 905: | Line 912: | ||
'''Monday 11th Oct''' | '''Monday 11th Oct''' | ||
*No Nicolas Kylilis, what a douche! | *No Nicolas Kylilis, what a douche! | ||
| - | *Diluted my overnights | + | *Diluted my overnights 250μl into 5ml to grow up to around 0.5 OD |
*Gonna perform assays with them this afternoon after some growing time :))))))) | *Gonna perform assays with them this afternoon after some growing time :))))))) | ||
*Doing data analysis with my last assay results comparing high copy (1C3) vs low copy (3K3)and the effects on XylE activity..boring | *Doing data analysis with my last assay results comparing high copy (1C3) vs low copy (3K3)and the effects on XylE activity..boring | ||
| Line 911: | Line 918: | ||
'''Tues 12th Oct''' | '''Tues 12th Oct''' | ||
| - | *Transformations worked! we now have | + | *Transformations worked! we now have pVeg-XylE in 3k3 and reverse-XylE with lacI promoter (C-TEV vector)i did replica plates, catechol assay plates and overnights for minis. Will mini prep tmro and transform them into TOP10 and we should be good to go with all constructs. |
* Assays messed up, my fault :( 3k3 didnt respond to catechol, so i think that I picked a background colony, i wont pick it again. The results are a bit ridiculous and will probably be chucked. | * Assays messed up, my fault :( 3k3 didnt respond to catechol, so i think that I picked a background colony, i wont pick it again. The results are a bit ridiculous and will probably be chucked. | ||
'''Weds 13th Oct''' | '''Weds 13th Oct''' | ||
| - | mini prepped 3k3- | + | mini prepped 3k3-pVeg-XylE-B0014 and went to the school workshop! set off overnight midi of culture 9 |
'''Thurs 14th Oct''' | '''Thurs 14th Oct''' | ||
| - | Midi prepped 3k3- | + | Midi prepped 3k3-pVeg |
'''Fri 15th''' | '''Fri 15th''' | ||
| - | Transformed all my constructs into T0P10 3k3-J23101/pveg and PSB1C3-J23101/ | + | Transformed all my constructs into T0P10 3k3-J23101/pveg and PSB1C3-J23101/pVeg |
|} | |} | ||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
|style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| Output Photo Gallery | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| Output Photo Gallery | ||
|- | |- | ||
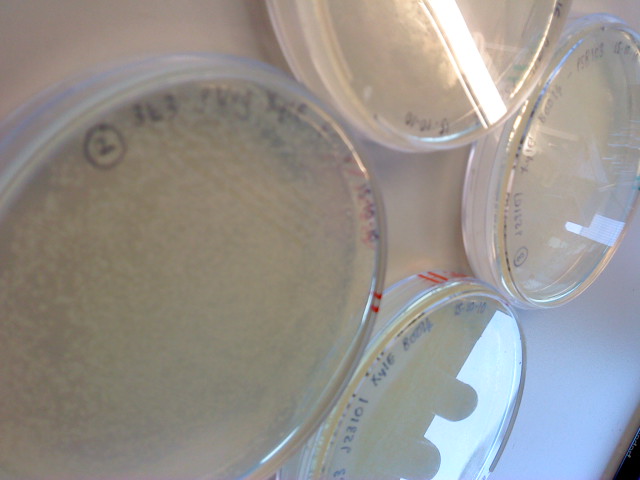
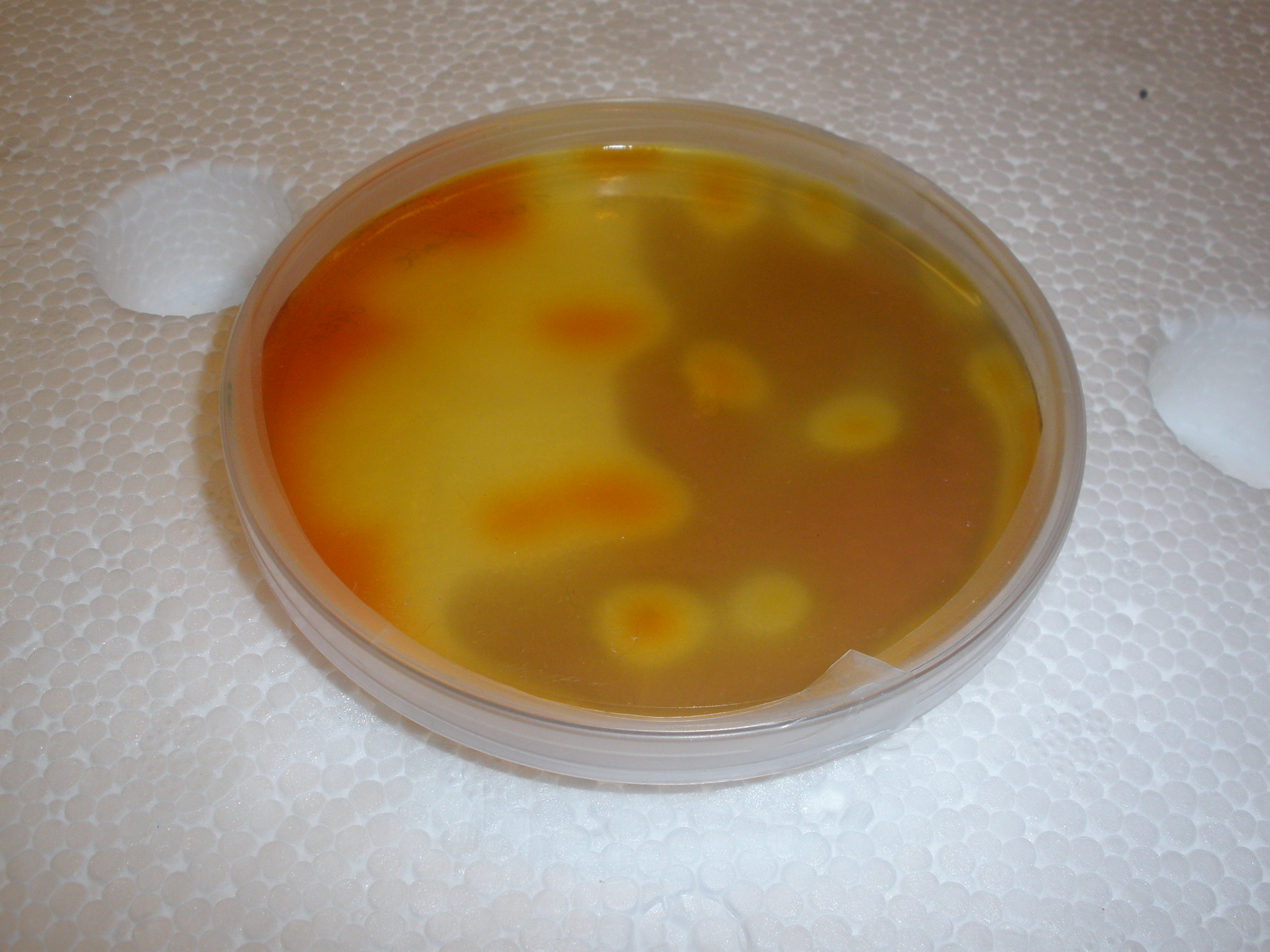
| - | |[[Image:IC_IMAG0310.jpg|thumb| | + | |[[Image:IC_IMAG0310.jpg|thumb|center|300px|First Successful catechol assay :)))]] |
| - | |[[Image:IC_CS1.JPG|thumb| | + | |[[Image:IC_IMAG0313.jpg|thumb|center|300px|First successful catechol assay]] |
| + | |- | ||
| + | |[[Image:IC_CS1.JPG|thumb|center|300px|Experiment which determined the yellow product to be exported out of the cells into the media and not retained by the cells]] | ||
| + | |[[Image:IC_CS2.JPG|thumb|center|300px|Cell vs supernatant assay]] | ||
| + | |- | ||
| + | |[[Image: IC_IMAG0322.jpg|thumb|center|300px|Cuvette assay set up]] | ||
| + | |[[Image:IC_IMAG0315.jpg|thumb|center|300px|:-)]] | ||
|- | |- | ||
| - | |[[Image: | + | |[[Image:miaow.jpg|thumb|center|300px|]] |
| - | + | |[[Image:miaow2.jpg|thumb|center|300px|]] | |
| - | + | ||
| - | |[[Image: | + | |
| - | + | ||
|- | |- | ||
| - | |[[Image:miaow3.jpg|thumb| | + | |[[Image:miaow3.jpg|thumb|center|300px|final batch of transformed XylE constructs into TOP10s]] |
| - | |[[Image:miaow4.jpg|thumb| | + | |[[Image:miaow4.jpg|thumb|center|300px|Fusion GFP-XylE vs XylE cuvette response assay]] |
|- | |- | ||
| - | |[[Image:miaow5.jpg|thumb| | + | |[[Image:miaow5.jpg|thumb|center|300px|View from above]] |
| + | |[[Image:miaow6.jpg|thumb|center|300px|Plate after assay!]] | ||
|} | |} | ||
Latest revision as of 03:25, 28 October 2010
| Lab Diaries | Overview | Surface Protein Team | XylE Team | Vectors Team | Modelling Team |
| Here are the technical diaries for our project. We've split them up into three lab teams and the modelling team. We think it's really important that absolutely anyone can find out what we've been doing. For a really detailed look at what we did, and when, you've come to the right place! | |
| XylE Team |
|
Objectives:
|
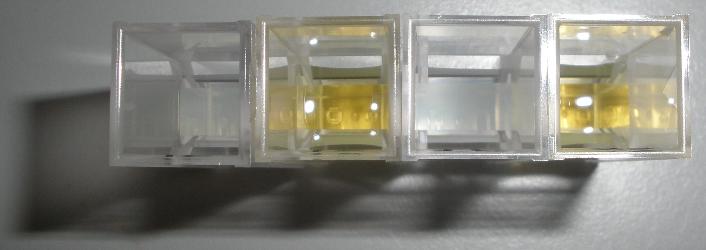
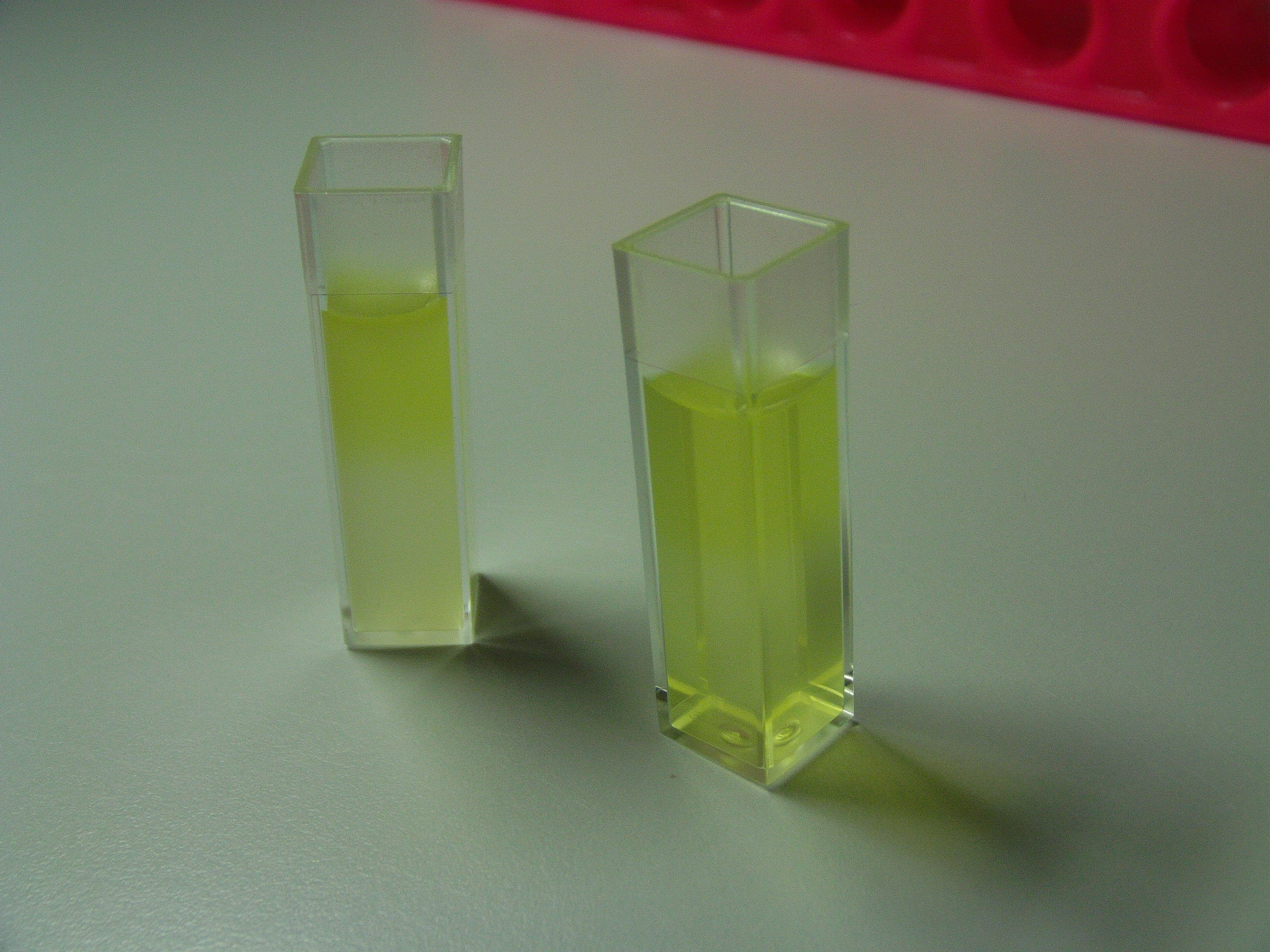
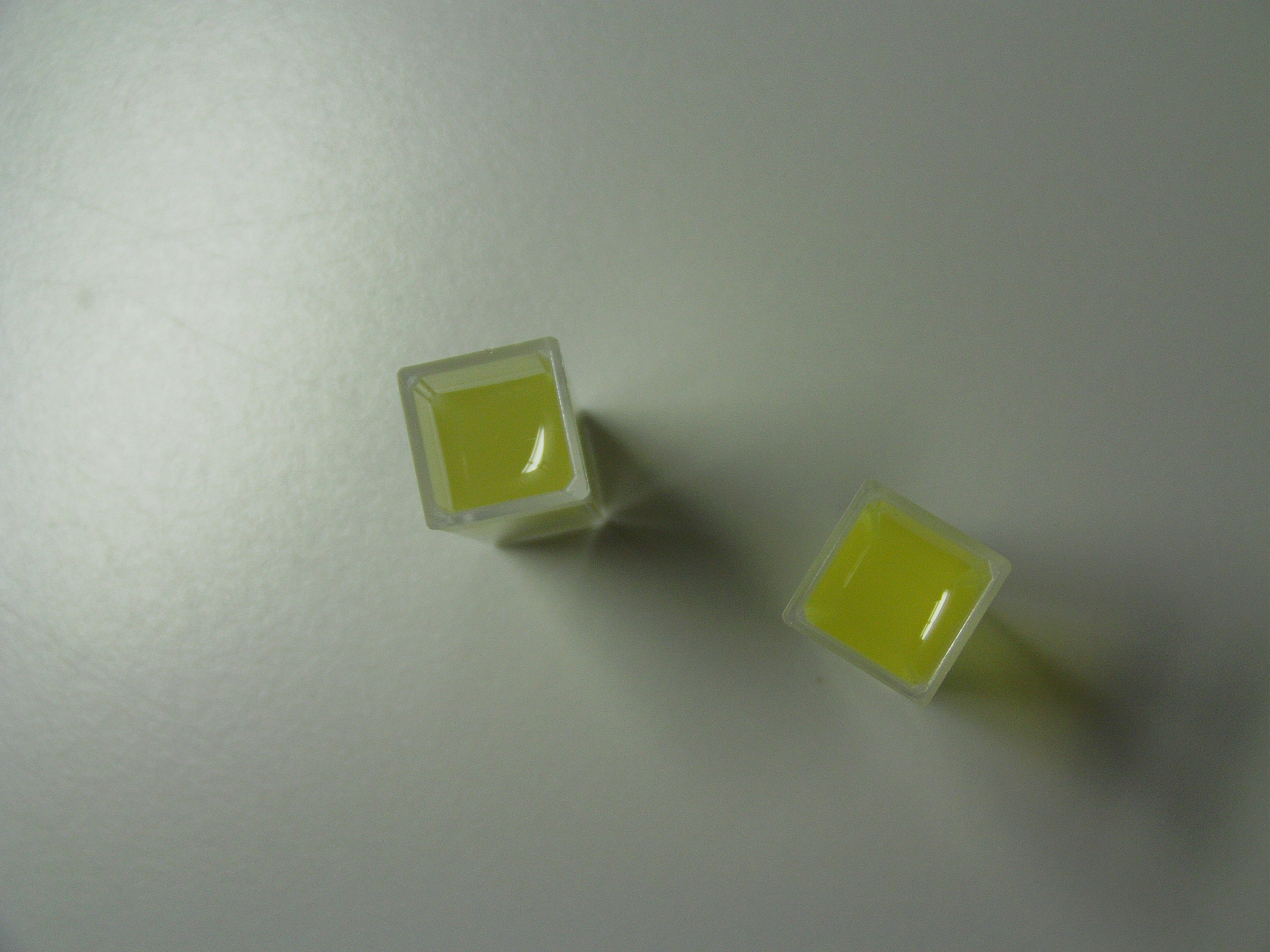
| Typical XylE assay |
| Week 6 | ||||||||||||||||||
|
we constructed the standard E.coli promoter J23101 with sticky ends. These ends are complementary to restriction sites made by EcoRI and SpeI enzyme. This promoter will be later used in 3A assembly to construct a promoter-RBS-XylE design in a psB1C3 vector. E.coli will be transformed with this final construct plasmid to assess XylE activity and characterization. It will also be one of the submitted BioBricks.
these cultures are going to be used tomorrow for mini prepping. Mini prep will allow us to isolate E.coli 's plasmid DNA(which contains the XylE gene). Friday, 13-Aug-2010
Mini prep is usually used to confirm that our gene of interest has not been changed in any way, as the isolated plasmid id sent for sequencing. However, since XylE was taken from the registry, we assume that it is fine and no sequencing is required. The mini prep will later be used for the midi-prep (that gives out higher yields of DNA needed for cloning).
|
| Week 7 | ||||||||||||||||||||
|
Tuesday, 17-Aug-2010
gel analysis of XylE, J23101 promoter and pSB1C3 vector samples to determine the volume ratios of samples to be used for 3A assemply ligation
Friday, 20-Aug-2010 The J23101 gene in a BioBrick vector containg RFP gene
|
| Week 8 | ||||||||||||||||||
|
Monday, 23-Aug
Tuesday, 24th-Aug
Wednesday, 25th-Aug Performed gel analysis on the purified XylE and J23101 to obtain ratios for ligation. First gel was scrapped as it produced appalling (explanation for Nick: really bad) results, 2nd gel run was successful.
Thursday, 26th-Aug
Friday, 27th-Aug
Saturday, 28th-Aug
Sunday, 29th-Aug
|
| Week 10 | ||||||||||||||||||
|
Tuesday 7th Sept
Wednesday, 8th Sep
Thursday, 9th Sep
Friday, 10 sep The transformation was a SUCCESS. 2x replica plates were made plate 1# 1-6 plate #2 6-11; colony 6 and colony 9 of plates 1 and 2 respectively were transfered into a 5ml liquid culture + 5μl CmR. These will later be turned into glycerol stocks. After the replica plates have grown up mini preps on a number of colonies shall be performed - this hopefully will eliminate the contaminating plasmid DNA. This will be followed by a midi prep. Saturday, 11 Sep The transformation of E.Coli with PSB1-C3 with insert did not work :( |
| Week 11 | ||||||||||||||||||
|
Monday 13th september Overnights weren't set up on sunday so they were made up alongside some assay cultures. J23101-XylE-B0014 colonies 8 and 10 of the replica plate #2 were picked. Chris also provided a replica plate containing 3k3 vector colonies. This was over a year old and he was unsure whether it was the correct plasmid or if the cells would grow up. I picked all available colonies 102 150 151 and 260 of kanamycin resistance. Lastly for assays 2x LB 2xM9 cultures were made 5ml +5ul antibiotic. Tues 14th September
The assay was carried out with E.coli, top ten spcies, transformed with J23101-XylE-B0014 in pSB1C3 vector.The overnight culture wastransfered in new medium this morning for 4 hrs before assaying. LB medium was used for dilutions and blank. Catechol was diluted in ddH2O. Data analysis of the assay
Piotr:
Weds 15th September
Thurs 16th September
- digest with XbaI and SpeI - PCR reactions with the following primers added: HIS-GFP, XylE-Xba and the other sample with HIS-GFP and GFP-flag The PCR reactions were not very conclusive on the gels but digests allowed to determine that colonies 1->5 seem to have the right sizes of DNA in them and on Friday they will be prepared to be sent off for sequencing
Friday 17th September
The gel purifications of 3k3 vector and XylE were used for ligation. 3K3 was dephosphorylated and then ligated in a ratio 5:1 with the insert. This was left overnight and Chris tried to transform with it on sunday.. ligation and transformation failed :-( |
| Week 12 |
|
Monday 20th Sept Performed ligation again. Errors in the sequencing we received back regarding J23101-XylE-B0014. One was a log error (in XylE) phew. The other is in PSB1C3 out of the scar site and should be OK. Chris is currently assembling the pVeg promoter+ RBS in a vector. Once he has finished this, I will combine XylE-GFP and XylE with this promoter for comparison characterization with J23101 (in E.coli and Bacillus) Tues 21st Sept
Wed 22nd Sept
Thurs 23rd Sept First day of AUTUMN!!
Fri 24th Sept
|
| Week 13 |
|
Monday 27th Sept
Tues 28th Sept
Weds 29th Sept
Thurs 30th Sept
Friday 1st Oct
ahahahaha awesome. hey you :)
|
|
have a look :) it's from John Hoppkins wiki 2008. There are one or two good ones! Love: Before I heard the doctors tell The dangers of a kiss; I had considered kissing you. The nearest thing to bliss. But now I know biology and sit and sigh and moan; six million mad bacteria and I thought we were alone!
|
| Week 14 |
|
Monday 4th Oct Did a replica plate and catechol assay plate/ minis 2 6 7 9 12 16. (Successful pVeg-XylE transformation) Tues 5th Oct 4 16 colonies were background the rest turned yellow. Mini prepped 2 6 7 9 12 16. Made overnight assay cultures of 3k3-XylE and C3-xylE Weds 6th Oct
Friday 8th Oct
Sunday 10th of the 10th of the 2010!!!!! and im in LAB
|
| Week 15 |
|
Monday 11th Oct
Tues 12th Oct
Weds 13th Oct mini prepped 3k3-pVeg-XylE-B0014 and went to the school workshop! set off overnight midi of culture 9 Thurs 14th Oct Midi prepped 3k3-pVeg Fri 15th Transformed all my constructs into T0P10 3k3-J23101/pveg and PSB1C3-J23101/pVeg |
| Output Photo Gallery | |
 "
"