Team:Heidelberg/Project/miMeasure
From 2010.igem.org
(→Abstract) |
(→Analysis of endogenous miRNA) |
||
| (53 intermediate revisions not shown) | |||
| Line 4: | Line 4: | ||
{{:Team:Heidelberg/Side_Top}} | {{:Team:Heidelberg/Side_Top}} | ||
[[Image:MiMeasure.png|frameless|250px|miMeasure Plasmid]] | [[Image:MiMeasure.png|frameless|250px|miMeasure Plasmid]] | ||
| + | |||
| + | '''With miMeasure you can:''' | ||
| + | |||
| + | *Characterize synthetic/ natural binding sites | ||
| + | **Regulate EGFP with customed BS | ||
| + | **Use EBFP2 as reference | ||
| + | |||
| + | *Clone BS behind the EGFP with the BBb standard | ||
| + | |||
| + | *Characterize miRNA expression in cell populations or at single cell level by measuring differences in the relative EGFP intensities with standard methods like FACS or microscopy | ||
| + | |||
| + | |||
| + | |||
{{:Team:Heidelberg/Side_Bottom}} | {{:Team:Heidelberg/Side_Bottom}} | ||
=miMeasure= | =miMeasure= | ||
| - | |||
| - | + | ||
| - | + | ||
| - | + | <h3> The '''miMeasure''' standard plasmid has been engineered to enable the easy input of synthetic microRNA binding sites behind one of two fluorescent proteins while the second is used for normalization. Expression of regulated reporter and control from a bidirectional CMV promter guarantee faithful and reproducible measurements in any kind of cell. The fluorescence readout can be used to quantify the regulatory efficiency of the binding site in knockdown percentage. Once the properties of a synthetic binding site are elucidated, they can be used to manipulate and accurately fine-tune gene expression in vitro and in vivo. </h3> | |
==Introduction== | ==Introduction== | ||
| - | Micro RNAs | + | Micro RNAs mainly regulate the translation of their target genes by interacting with regions in the 3' untranslated region (UTR) of their target mRNA. Base-pairing with the miRNA binding site (BS) causes formation of diverse miRNA-mRNA duplexes {{HDref|reviewed by Fabian et al., 2010}}. The BS consists of a seven nucleotide long seed region that is perfectly matched to the miRNA, and surrounding regions that matched partially. The seed region is defined as being the minimal required base-pairing that can regulate the mRNA. Apart from the seed region, binding can be unspecific, creating mismatches and bulges. The position and properties of the bulges seem to play a central role in miRNA binding and therefore knockdown efficiency {{HDref|reviewed by Bartel et al., 2009}}. Since we were going to use synthetic miRNA BS in our genetherapeutic approach, we had to find a way to study their effects in a standardized manner that would be comparable and reproducible. <br> |
| - | One goal of the iGEM Team Heidelberg 2010 was to test the effects of changes in BS sequences on expression control. Thereby miRNA BS should be characterized. To standardize our measurements of knockdown according to BS specificity, we had to come up with a new standard that is independent from the endogenous cell machinery. We decided to bring synthetic miRNAs into play, hence we engineered BS for them creating an artificial regulatory circuit<!--, simulating naturally occurring miRNAs and miRNA BS without having to worry about the effect of endogenous targets-->. Of course there are also differences that arise through the availability of the enzymes involved in the miRNA pathway that may differ slightly from cell to cell. Therefore, | + | |
| - | The main purpose of our measurement standard, miMeasure, is to express two nearly identical but discernible proteins: one of them tagged with a BS, the other one unregulated (even though the possibility exists to clone in a reference binding site). The two reporters are expressed by a bidirectional CMV promoter to make sure their transcription rate is comparable. We used a destabilized version of GFP, dsEGFP and a dsEBFP2 that was derived from the same sequence ([https://2010.igem.org/Team:Heidelberg/Project/miMeasure#References Ai et al., 2007]). Thus, we could make sure that both proteins exhibit the same synthesis and degradation properties, making them directly comparable. Hereby, we can also neglect the difference between mRNA and protein knockdown and can take the fluorescence of the marker protein as a direct, linear output of mRNA down-regulation. We included a BBb standard site into our plasmid, which allows to clone BS behind the GFP. If co-transfected with the corresponding shRNA miR, GFP will be down-regulated, while BFP expression is maintained. The ratio of GFP to BFP expression can be used to conclude the knockdown efficiency | + | One goal of the iGEM Team Heidelberg 2010 was to test the effects of changes in BS sequences on expression control. Thereby miRNA BS should be characterized. To standardize our measurements of knockdown according to BS specificity, we had to come up with a new standard that is independent from the endogenous cell machinery. We decided to bring synthetic miRNAs into play, hence we engineered BS for them creating an artificial regulatory circuit<!--, simulating naturally occurring miRNAs and miRNA BS without having to worry about the effect of endogenous targets-->. Of course there are also differences that arise through the availability of the enzymes involved in the miRNA pathway that may differ slightly from cell to cell. Therefore, constructs containing changes in BS sequences has to be compared to construct containing no binding sites and containing perfect binding sites. Ideally, the miRNA would be stably expressed in the cell line, but a uniform co-transfection also leads to an even distribution of synthetic shRNA-like miRNAs (shRNA miRs). Additionally, miRNA levels can be adjusted by differing transfection ratios. <br> |
| + | |||
| + | The main purpose of our measurement standard, miMeasure, is to express two nearly identical but discernible proteins: one of them tagged with a BS, the other one unregulated (even though the possibility exists to clone in a reference binding site). The two reporters are expressed by a bidirectional CMV promoter to make sure their transcription rate is comparable. We used a destabilized version of GFP, dsEGFP and a dsEBFP2 that was derived from the same sequence ([https://2010.igem.org/Team:Heidelberg/Project/miMeasure#References Ai et al., 2007]). Thus, we could make sure that both proteins exhibit the same synthesis and degradation properties, making them directly comparable. Hereby, we can also neglect the difference between mRNA and protein knockdown and can take the fluorescence of the marker protein as a direct, linear output of mRNA down-regulation. We included a BBb standard site into our plasmid, which allows to clone BS behind the GFP. If co-transfected with the corresponding shRNA miR, GFP will be down-regulated, while BFP expression is maintained. The ratio of GFP to BFP expression can be used to conclude the knockdown efficiency of the BS. Having destabilized marker proteins with a turnover time of two hours enables us not only to avoid accumulation of marker proteins, which would make the knockdown harder to observe, but also to conduct time-lapse experiments. In the future, this could be for example a way to observe dynamic activity patterns of endogenous miRNAs. | ||
<html> | <html> | ||
<div class="backtop"> | <div class="backtop"> | ||
| Line 28: | Line 42: | ||
===Analysis of Randomized Binding Sites Against Synthetic miRNA=== | ===Analysis of Randomized Binding Sites Against Synthetic miRNA=== | ||
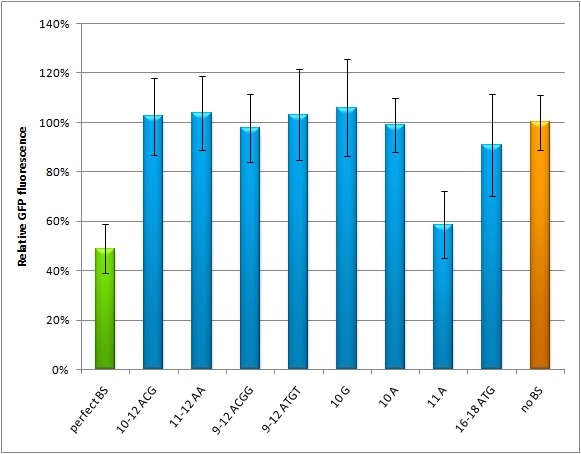
| - | + | We modified binding sites for the synthetic miRNA by introducing random basepairs surrounding the seed region as described above, thereby changing the knockdown efficiency. In figure 1 we plotted the knockdown percentage of our constructs. The miMeasure construct and negative control were [https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Transfection co-transfected] with either shRNA miRsAg expressed from a CMV promoter on pcDNA5 or an inert synthetic RNA as control in 1:4 ratio, respectively. | |
| - | + | <br><br> | |
| - | + | ||
| - | + | ||
| - | + | ||
{| class="wikitable sortable" border="0" align="center" style="text-align: left" | {| class="wikitable sortable" border="0" align="center" style="text-align: left" | ||
| Line 60: | Line 71: | ||
|- | |- | ||
|} | |} | ||
| + | <br><br> | ||
| + | |||
| + | ====Flow cytometry measurements==== | ||
| + | |||
| + | Hela cells transfected with the constructs (described above) are taken for flow cytometry. Each measurement contains around 10000 cells. The cells are plotted on a logarithmic scale in relation to EGFP and EBFP2 intensity. Each dot intensity represents the count of fluorescent cells for each EGFP/EBFP2 intensity pair. The dots are colour coded, so that the orange dots represent cells cotransfected with different miMeasure constructs and the miRsAg and the blue one represent cells cotransfected with non-specific miRNA (miR-155), respectively. So the blue set of measurements represent the negative control. Before the real measurements, cells transfected with EGFP or EBFP2 alone were measured to establish the gain of the detectors and compensate for fluorescent bleedthrough, especially of EBFP2 into the EGFP channel . This ensured that EGFP and EBFP2 alone shows a perfectly horizontal and vertical distribution respectively (data not shown). For the miMeasure constructs, both population of dots make up a oblique line on the logarithmic scale, which shows the correlation of EGFP and EBFP2 very well. If the two different coloured dots overlap, they become white. Thus both lines overlap almost completely in the negative control containing no binding site, whereas the orange line shifts to the left for the miMeasure construct with the perfect binding site. All the other constructs are like the negative control. | ||
| + | As the difference is subtle on the logarithmic scale, we also observed the cell distribution on a linear scale. All coloured distributions appeared more scattered on the linear scale, but the shifting of the orange dots was more visible for the construct containing the perfect binding site. again all the other constructs have the same range of scattering as the negative control. | ||
| + | |||
| + | [[Image:Flow1.jpg|thumb|610px|center|'''Figure 1: GFP/BFP correlation of single transfected Hela cells according to flow cytometry analysis on a logarithmic scale''' different binding sites for miRsAg cotransfected with miRsAg or with mi-R155, respectively. The orange dots represent the cotransfected cells with miRsAg and the blue dots the cotransfected cells with miR-155. Hela cells were used.]]^ | ||
| + | |||
| + | [[Image:Flow-linear-result.jpg|thumb|610px|center|'''Figure 2: GFP/BFP correlation of single transfected Hela cells according to flow cytometry analysis on a linear scale''' different binding sites for miRsAg cotransfected with miRsAg or with mi-R155, respectively. The orange dots represent the cotransfected cells with miRsAg and the blue dots the cotransfected cells with miR-155. Hela cells were used.]] | ||
| + | |||
| + | Flow cytometry offers an efficient way to estimate the efficiency of expression regulation. Here we could observe that while the perfect binding site generate a partial knockdown, all modified binding site constructs looked similar to the negative control. To characterize potential differences that cannot be identified by flow cytometry, we next imaged the same cells and precisely quantified their fluorescence intensity using laser scanning confocal microscopy at high resolution. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ====Confocal microscopy measurements==== | ||
| + | |||
| + | We used microscopy analysis to determine the EGFP expression in relation to EBFP2. EBFP2 serves as a normalization for transfection efficiency. Nine miMeasure constructs with different binding sites were designed. Those were cloned downstream of EGFP behind the miMeasure construct, whereas the EBFP2 expression stays unaffected. The GFP/BFP-ratio stand for the level of GFP-expression normalized to one copy per cell. | ||
| + | |||
| + | |||
| + | |||
| + | [[Image:M12-M22_HeLa_daten.jpg|thumb|500px|center|'''Figure 3: GFP/BFP ratio normalized by the negative control''' The data are generated by confocal microscopy of Hela cells, which were transfected with different miMeasure constructs M12-M22 including the negative control (miMeasure construct without binding site). The negative control equals to 1.]] | ||
| Line 83: | Line 118: | ||
</div> | </div> | ||
</html> | </html> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
===Analysis of miRaPCR Generated Binding Sites Against a Natural miRNA=== | ===Analysis of miRaPCR Generated Binding Sites Against a Natural miRNA=== | ||
| Line 99: | Line 125: | ||
The [https://2010.igem.org/Team:Heidelberg/Notebook/BSDesign/July miRaPCR] generates binding sites out of rationally designed fragments. These are aligned with each other by chance, whereby different spacer regions are inserted randomly in between. It has been suggested that having more than one binding site of for the same miRNA behind each other can lead to stronger down-regulation than a single one. If imperfect binding sites are aligned, it is also supposed to be stronger than a single one. This is what we tested using MiRaPCR for effortless assembly of binding site fragments. | The [https://2010.igem.org/Team:Heidelberg/Notebook/BSDesign/July miRaPCR] generates binding sites out of rationally designed fragments. These are aligned with each other by chance, whereby different spacer regions are inserted randomly in between. It has been suggested that having more than one binding site of for the same miRNA behind each other can lead to stronger down-regulation than a single one. If imperfect binding sites are aligned, it is also supposed to be stronger than a single one. This is what we tested using MiRaPCR for effortless assembly of binding site fragments. | ||
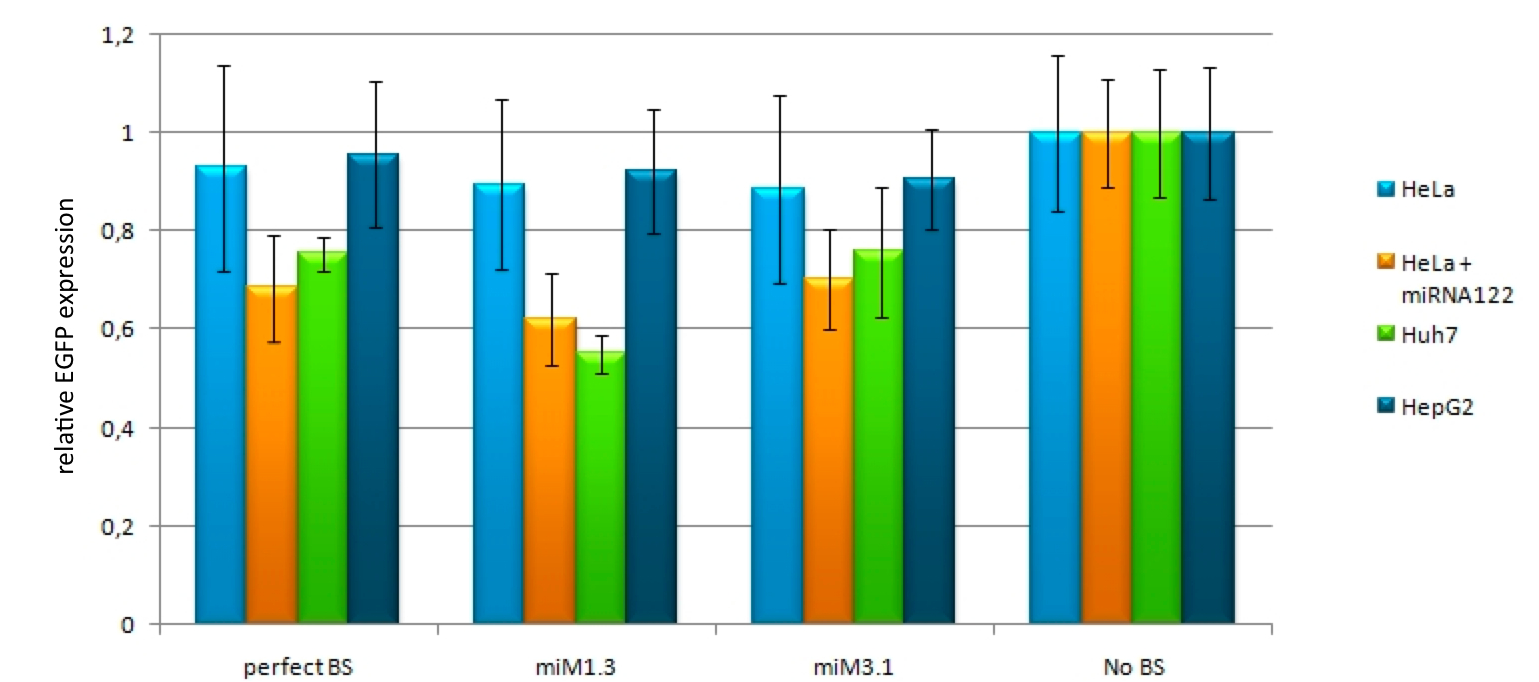
For our experiments, we took advantage of the high abundance of miRNA 122 in liver cells and tested different combinations of binding sites created by miRaPCR. We transfected HeLa and HuH7 cells with the constructs described in table 2. Since HuH7 cells express miR-122, the constructs will be affected in the HuH7 cells without cotransfecting any miRNAs, whereas miR-122 were cotransfected for the HeLa cells in 2:1 ratio. The result shows the ratio between GFP and BFP normalized to the negative control (miMeasure constructs without binding sites). The miMeasure constructs were also compared to the expression of miMeasure containing one perfect binding site for miRNA 122. | For our experiments, we took advantage of the high abundance of miRNA 122 in liver cells and tested different combinations of binding sites created by miRaPCR. We transfected HeLa and HuH7 cells with the constructs described in table 2. Since HuH7 cells express miR-122, the constructs will be affected in the HuH7 cells without cotransfecting any miRNAs, whereas miR-122 were cotransfected for the HeLa cells in 2:1 ratio. The result shows the ratio between GFP and BFP normalized to the negative control (miMeasure constructs without binding sites). The miMeasure constructs were also compared to the expression of miMeasure containing one perfect binding site for miRNA 122. | ||
| + | <br><br><br><br> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
{| class="wikitable sortable" border="0" align="center" style="text-align: left" | {| class="wikitable sortable" border="0" align="center" style="text-align: left" | ||
|-bgcolor=#009be1 | |-bgcolor=#009be1 | ||
| Line 134: | Line 142: | ||
|} | |} | ||
| - | |||
| + | ====Flow cytometry==== | ||
| - | The | + | The same constructs in Hela cells were analyzed by flow cytometry, too. Here the orange dots also represent the miMeasure construct transfected with the specific miRNA and the blue dots make up the negative control. The orange dots from the construct containing the perfect and the aligned binding sites have lower EGFP expression compared to EBFP2 expression, since the EGFP trend doesn't correspond to the EBFP2 trend, but shifts to the laft. The orange dots rise with the fluorescence intensity and collapses to zero, when the EBFP2 intensity is high. For the other constructs the EGFP range fully corresponds with the EBFP2 range. |
| + | The linear plot shows the orange and blue dots in more distinct lines. The orange dots from the construct containing the perfect and aligned binding sites assemble along the y-axis, where the EGFP fluorescence intensity is zero. For the other construct the EGFP line fully overlap with the EBFP2. | ||
| + | [[Image:Flow_miR122.jpg|thumb|620px|center|'''Figure 4:EGFP2/EBFP correlation of single transfected Hela cells according to flow cytometry analysis''' different binding sites for miR122 cotransfected with miR-122 or with miR-155, respectively. The orange dots represent the cotransfected cells with miR122 and the blue dots the cotransfected cells with miR-155. Hela cells were used.]] | ||
| + | [[Image:Flow_miR122_linear.jpg|thumb|620px|center|'''Figure 5: EGFP/EBFP2 correlation of single transfected Hela cells according to flow cytometry analysis''' different binding sites for miR122 cotransfected with miR-122 or with miR-155, respectively. The orange dots represent the cotransfected cells with miR122 and the blue dots the cotransfected cells with miR-155. Hela cells were used.]] | ||
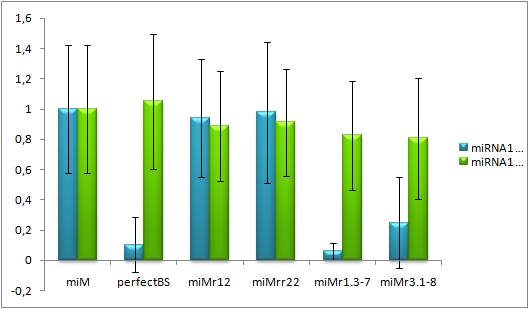
| + | Each cell in the images has a position in the x- and y-axis. So the intensity and number of cells with a certain intensity can be used for [https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Flow_cytometry calculation] of the mean EGFP/EBFP2 ratio. The calculated data show, that there is a significant EGFP knock-down in the cells cotransfected with miR-122 and the contruct with the perfect binding site or the aligned binding sites, respectively. | ||
| + | [[Image:Hela_miRNA122_ratio.jpg|thumb|620px|center|'''Figure 6:EGFP/EBFP2 correlation of single transfected Hela cells according to flow cytometry analysis''' different binding sites for miR122 cotransfected with miR-122 or with miR-155, respectively.]] | ||
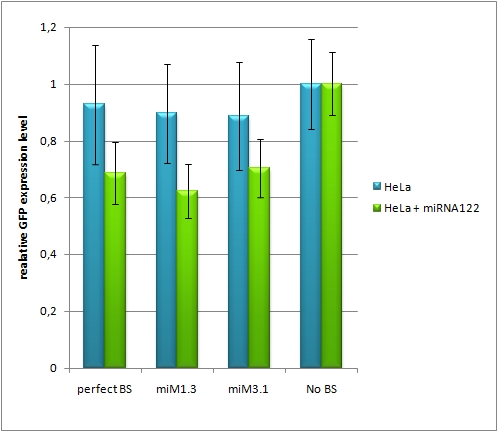
| + | ===Confocal microscopy=== | ||
| - | + | [[Image:HeLa_miR122.jpg|thumb|500px|center|'''Figure 7: different binding sites for miR122, HeLa cotransfected with miR122 expression plasmid''']] | |
| - | The | + | The EGFP-expression normalized to the EGFB2 expression is set to 100% for the miMeasure construct transfected with the non-matching miRNA (in this case miR-155). The knock-down efficiency of one perfect binding site is around 30%, which also accounts for the three aligned perfect binding sites and the two aligned imperfect ones. The binding site with bulges from position 9-12 and 9-22 don't show any knock-down. |
| - | + | ||
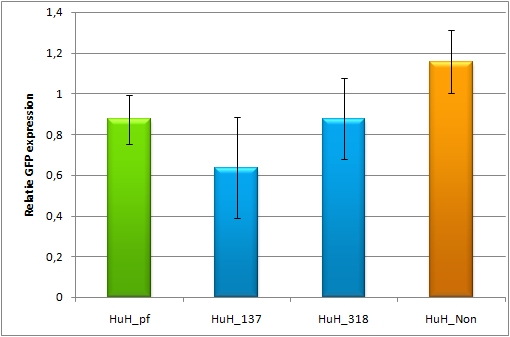
| - | [[Image: | + | [[Image:HuH7_miR122b.jpg|thumb|500px|center|'''Figure 8: different binding sites for miR122, Huh7 cells''']] |
| - | [[Image: | + | |
| + | |||
| + | |||
| + | The transfected HeLa are also imaged with the epifluorescent microscope. Large amount of cells in the negative control (miMeasure with perfect binding site cotransfected with miR-155, see a)are green, whereas most cells with the miMeasure construct containing the perfect binding sites (see b) are blue. | ||
| + | |||
| + | [[Image:BLUE+green.jpg|thumb|800px|center|'''Figure 9: epifluorescent microscopy image (10x) of Hela cells transfected with miMeasure''' miMeasure with a perfect binding site is a) cotransfected with miR-155, which has no specificity to miR-122, b) cotransfected with miR-122, which is complementary to the perfect binding site. EGFP is regulated by miR-122, EBFP2 is unregulated and serves as transfection control.]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | The Huh7 cells were also transfected with the 4 different constructs. Here a cotransfection with miR-122 is not necessary, since Huh7 cells express miR-122 themselves. The knock-down of the perfect binding sites are stronger than the knock-down in the Hela cells. Here the knock-down efficiency is 80% for the perfect binding site and the aligned constructs. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <!--Discussion This experiment shows, that the binding site pattern with 3 aligned perfect binding sites miM-1.3-7 gives the strongest knock-down. Whereby the binding site pattern with two imperfect binding sites 1-8 is weaker, but still stronger than the negative control. If one binding site is randomized from nucleotide 9-12 or 9-22 miM-r12, miM-r22, it loses its capability of protein down-regulation.--> | ||
===Analysis of endogenous miRNA=== | ===Analysis of endogenous miRNA=== | ||
| - | HepG2, another liver cell line, is also transfected with the constructs containing the perfect and the aligned constructs for miR-122. The cotransfection with miR-155 serves again as a negative control. For this cell line there was only a slight knock-down observed for all of the constructs, it is much less compared to the HuH7 cell line, where the knock-down ranges from 20-40%. | + | HepG2, another liver cell line, is also transfected with the constructs containing the perfect and the aligned constructs for miR-122. The cotransfection with miR-155 serves again as a negative control. For this cell line there was only a slight knock-down observed for all of the constructs, it is much less compared to the HuH7 cell line, where the knock-down ranges from 20-40%. |
| - | [[Image: | + | |
| + | [[Image:download-1.jpg|thumb|790px|center|'''Figure 10: relative EGFP expression of transfected HepG2 cells according to confocal microscopy analysis''' different binding sites for miR122 are cotransfected with miR-122. The negative control doesn't contain any binding sites.]] | ||
| Line 170: | Line 201: | ||
The EGFP/EBFP2 ratio from each transfected HuH7 cell is calculated for 52 cells. The cells were transfected with the construct carriying the three aligned perfect binding sites against miR-122. The EGFP/EBFP2 ratio in each cell is different and ranges from 0.1 to 6. | The EGFP/EBFP2 ratio from each transfected HuH7 cell is calculated for 52 cells. The cells were transfected with the construct carriying the three aligned perfect binding sites against miR-122. The EGFP/EBFP2 ratio in each cell is different and ranges from 0.1 to 6. | ||
| - | [[Image: | + | [[Image:Singlecellstuff.jpg|thumb|center|850px|'''Figure 11: Single cell analysis of relative EGFP expression of transfected HuH7 cells by confocal microscopy analysis''' The numbers in the barplot correspond to the cells on the right image.]] |
==Discussion== | ==Discussion== | ||
| - | + | Measurements of synthetic and endogenous miRNA expression level as well as the knock-down efficiency by different miRNAs can characterized via the miMeasure constructs. The fluorescence of EGFP and EBFP2, and therefore the efficiency of knockdown, can be estimated using different methods, for example [https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Flow_cytometry flow cytometry], manual and automated fluorescence [https://2010.igem.org/Team:Heidelberg/Notebook/Material_Methods#Microscopy microscopy] or potentially an automated fluorescence plate reader system. Flow cytometry and fluorescence microscopy offer the possibility to measure knockdown at the single cell level, the first one with the possibility of high throughput, the second with a good precision. Since flow cytometry has a smaller dynamic range, thus even a GFP knock-down of roughly 60% can cause the relative GFP fluorescence intensity values collapsing to zero. | |
| - | + | This quantitative approach offers many possibilities, two of them being expored here: estimating the knockdown capacity of binding sites that differs from a perfect one and determining the presence of endogenous miRNA directly in living cells.<br> | |
| - | + | The miMeasure construct can differ between a variation of knock-down efficiencies in the synthetic miRNA binding patterns, which was successfully generated by the [https://2010.igem.org/Team:Heidelberg/Notebook/BSDesign miRaPCR]. The perfect and aligned binding sites have the greatest knock-down efficiencies with roughly 50%. These experiemtal data corresponds to predictions from the modeling and hypothesis from the liturature. Hence the higher the complementarity from binding site to miRNA the more efficient the miRNA induced knock-down. Alignment of binding sites are shown to have additive effects, either. <br> | |
| - | The fluorescence of | + | miMeasure can also be used for the characterization of binding sites against synthetic miRNAs. The data only shows a difference between the perfect binding site and the construct without binding site. There were no more subtle differences detected from the slightly modified binding sites. The reason might be, that the synthtic miRNA is much less efficient concerning the knock-down than the natural binding site. So the modified binding sites, which would give an even weaker output, would be out of range. The possible low efficiency of the synthtic miRNA miRsAg could be explained by the fact, that it is assembled by taking the human miR-122 as a template, so that it can be driven by another promoter than Pol1, which is especially weak. So the synthetic miRNA might not expressed in the same amount like the natural miR-122. Real-time PCR could be done to validate the miRNA expression. <br> |
| + | The direct comparison between cell lines enables future miRNA profiling with miMeasure. Several cell lines can be compared directly with each other after cotransfection with the same miRNA. The analysis of the cell lines HeLa, HuH7 and HepG2 gave significant results in miRNA-expression level in those cell lines. This data also corresponds to liturature, which underlies the much higher miR-122 expression in HuH7 cells than in HepG2 cells {{HDref|Young et al., 2010}}. Besides the comparison between different cell lines, even single cells can be compared with each other, so that characterization of expression in single cells is possible, which will enable single cell studies as well. The great difference between single cell expression of miR-122 can be due to the carcinogenous property of HuH7. Accoring to {{HDref|Lin et al., 2008}} low levels of miR-122 are responsible for reduced apoptosis in liver cells, because miR-122 targets besides many other important genes, an anti-apoptotic gene, Bcl-w. | ||
==References== | ==References== | ||
| Line 183: | Line 215: | ||
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009 Jan 23;136(2):215-33. | Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009 Jan 23;136(2):215-33. | ||
| + | |||
| + | Coulouarn C, Factor VM, Andersen JB, Durkin ME, Thorgeirsson SS. Loss of miR-122 expression in liver cancer correlates with suppression of the hepatic phenotype and gain of metastatic propertiesmiR-122 repression is a marker of tumor progression in HCC. Oncogene. 2009 Oct 8;28(40):3526-36. | ||
Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem. 2010;79:351-79. | Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem. 2010;79:351-79. | ||
| + | |||
| + | Girard M, Jacquemin E, Munnich A, Lyonnet S, Henrion-Caude A. miR-122, a paradigm for the role of microRNAs in the liver. J Hepatol. 2008 Apr;48(4):648-56. | ||
| + | |||
| + | Lin CJ, Gong HY, Tseng HC, Wang WL, Wu JL. miR-122 targets an anti-apoptotic gene, Bcl-w, in human hepatocellular carcinoma cell lines. Biochem Biophys Res Commun. 2008 Oct 24;375(3):315-20. | ||
Latest revision as of 03:56, 28 October 2010

miMeasureThe miMeasure standard plasmid has been engineered to enable the easy input of synthetic microRNA binding sites behind one of two fluorescent proteins while the second is used for normalization. Expression of regulated reporter and control from a bidirectional CMV promter guarantee faithful and reproducible measurements in any kind of cell. The fluorescence readout can be used to quantify the regulatory efficiency of the binding site in knockdown percentage. Once the properties of a synthetic binding site are elucidated, they can be used to manipulate and accurately fine-tune gene expression in vitro and in vivo.IntroductionMicro RNAs mainly regulate the translation of their target genes by interacting with regions in the 3' untranslated region (UTR) of their target mRNA. Base-pairing with the miRNA binding site (BS) causes formation of diverse miRNA-mRNA duplexes [http://2010.igem.org/Team:Heidelberg/Project/miMeasure#References (reviewed by Fabian et al., 2010)]. The BS consists of a seven nucleotide long seed region that is perfectly matched to the miRNA, and surrounding regions that matched partially. The seed region is defined as being the minimal required base-pairing that can regulate the mRNA. Apart from the seed region, binding can be unspecific, creating mismatches and bulges. The position and properties of the bulges seem to play a central role in miRNA binding and therefore knockdown efficiency [http://2010.igem.org/Team:Heidelberg/Project/miMeasure#References (reviewed by Bartel et al., 2009)]. Since we were going to use synthetic miRNA BS in our genetherapeutic approach, we had to find a way to study their effects in a standardized manner that would be comparable and reproducible. One goal of the iGEM Team Heidelberg 2010 was to test the effects of changes in BS sequences on expression control. Thereby miRNA BS should be characterized. To standardize our measurements of knockdown according to BS specificity, we had to come up with a new standard that is independent from the endogenous cell machinery. We decided to bring synthetic miRNAs into play, hence we engineered BS for them creating an artificial regulatory circuit. Of course there are also differences that arise through the availability of the enzymes involved in the miRNA pathway that may differ slightly from cell to cell. Therefore, constructs containing changes in BS sequences has to be compared to construct containing no binding sites and containing perfect binding sites. Ideally, the miRNA would be stably expressed in the cell line, but a uniform co-transfection also leads to an even distribution of synthetic shRNA-like miRNAs (shRNA miRs). Additionally, miRNA levels can be adjusted by differing transfection ratios. The main purpose of our measurement standard, miMeasure, is to express two nearly identical but discernible proteins: one of them tagged with a BS, the other one unregulated (even though the possibility exists to clone in a reference binding site). The two reporters are expressed by a bidirectional CMV promoter to make sure their transcription rate is comparable. We used a destabilized version of GFP, dsEGFP and a dsEBFP2 that was derived from the same sequence (Ai et al., 2007). Thus, we could make sure that both proteins exhibit the same synthesis and degradation properties, making them directly comparable. Hereby, we can also neglect the difference between mRNA and protein knockdown and can take the fluorescence of the marker protein as a direct, linear output of mRNA down-regulation. We included a BBb standard site into our plasmid, which allows to clone BS behind the GFP. If co-transfected with the corresponding shRNA miR, GFP will be down-regulated, while BFP expression is maintained. The ratio of GFP to BFP expression can be used to conclude the knockdown efficiency of the BS. Having destabilized marker proteins with a turnover time of two hours enables us not only to avoid accumulation of marker proteins, which would make the knockdown harder to observe, but also to conduct time-lapse experiments. In the future, this could be for example a way to observe dynamic activity patterns of endogenous miRNAs. ResultsAnalysis of Randomized Binding Sites Against Synthetic miRNAWe modified binding sites for the synthetic miRNA by introducing random basepairs surrounding the seed region as described above, thereby changing the knockdown efficiency. In figure 1 we plotted the knockdown percentage of our constructs. The miMeasure construct and negative control were co-transfected with either shRNA miRsAg expressed from a CMV promoter on pcDNA5 or an inert synthetic RNA as control in 1:4 ratio, respectively.
Flow cytometry measurementsHela cells transfected with the constructs (described above) are taken for flow cytometry. Each measurement contains around 10000 cells. The cells are plotted on a logarithmic scale in relation to EGFP and EBFP2 intensity. Each dot intensity represents the count of fluorescent cells for each EGFP/EBFP2 intensity pair. The dots are colour coded, so that the orange dots represent cells cotransfected with different miMeasure constructs and the miRsAg and the blue one represent cells cotransfected with non-specific miRNA (miR-155), respectively. So the blue set of measurements represent the negative control. Before the real measurements, cells transfected with EGFP or EBFP2 alone were measured to establish the gain of the detectors and compensate for fluorescent bleedthrough, especially of EBFP2 into the EGFP channel . This ensured that EGFP and EBFP2 alone shows a perfectly horizontal and vertical distribution respectively (data not shown). For the miMeasure constructs, both population of dots make up a oblique line on the logarithmic scale, which shows the correlation of EGFP and EBFP2 very well. If the two different coloured dots overlap, they become white. Thus both lines overlap almost completely in the negative control containing no binding site, whereas the orange line shifts to the left for the miMeasure construct with the perfect binding site. All the other constructs are like the negative control. As the difference is subtle on the logarithmic scale, we also observed the cell distribution on a linear scale. All coloured distributions appeared more scattered on the linear scale, but the shifting of the orange dots was more visible for the construct containing the perfect binding site. again all the other constructs have the same range of scattering as the negative control. 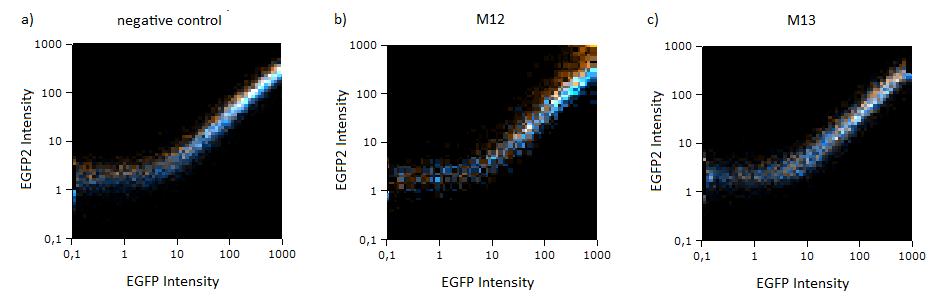 Figure 1: GFP/BFP correlation of single transfected Hela cells according to flow cytometry analysis on a logarithmic scale different binding sites for miRsAg cotransfected with miRsAg or with mi-R155, respectively. The orange dots represent the cotransfected cells with miRsAg and the blue dots the cotransfected cells with miR-155. Hela cells were used. 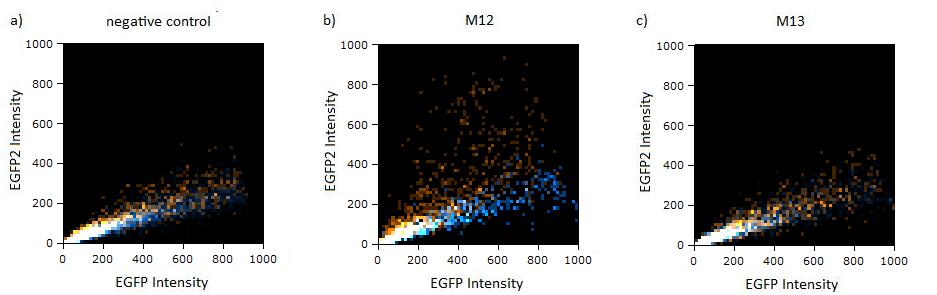 Figure 2: GFP/BFP correlation of single transfected Hela cells according to flow cytometry analysis on a linear scale different binding sites for miRsAg cotransfected with miRsAg or with mi-R155, respectively. The orange dots represent the cotransfected cells with miRsAg and the blue dots the cotransfected cells with miR-155. Hela cells were used. Flow cytometry offers an efficient way to estimate the efficiency of expression regulation. Here we could observe that while the perfect binding site generate a partial knockdown, all modified binding site constructs looked similar to the negative control. To characterize potential differences that cannot be identified by flow cytometry, we next imaged the same cells and precisely quantified their fluorescence intensity using laser scanning confocal microscopy at high resolution.
Confocal microscopy measurementsWe used microscopy analysis to determine the EGFP expression in relation to EBFP2. EBFP2 serves as a normalization for transfection efficiency. Nine miMeasure constructs with different binding sites were designed. Those were cloned downstream of EGFP behind the miMeasure construct, whereas the EBFP2 expression stays unaffected. The GFP/BFP-ratio stand for the level of GFP-expression normalized to one copy per cell.
Analysis of miRaPCR Generated Binding Sites Against a Natural miRNAThe miRaPCR generates binding sites out of rationally designed fragments. These are aligned with each other by chance, whereby different spacer regions are inserted randomly in between. It has been suggested that having more than one binding site of for the same miRNA behind each other can lead to stronger down-regulation than a single one. If imperfect binding sites are aligned, it is also supposed to be stronger than a single one. This is what we tested using MiRaPCR for effortless assembly of binding site fragments.
For our experiments, we took advantage of the high abundance of miRNA 122 in liver cells and tested different combinations of binding sites created by miRaPCR. We transfected HeLa and HuH7 cells with the constructs described in table 2. Since HuH7 cells express miR-122, the constructs will be affected in the HuH7 cells without cotransfecting any miRNAs, whereas miR-122 were cotransfected for the HeLa cells in 2:1 ratio. The result shows the ratio between GFP and BFP normalized to the negative control (miMeasure constructs without binding sites). The miMeasure constructs were also compared to the expression of miMeasure containing one perfect binding site for miRNA 122.
Flow cytometryThe same constructs in Hela cells were analyzed by flow cytometry, too. Here the orange dots also represent the miMeasure construct transfected with the specific miRNA and the blue dots make up the negative control. The orange dots from the construct containing the perfect and the aligned binding sites have lower EGFP expression compared to EBFP2 expression, since the EGFP trend doesn't correspond to the EBFP2 trend, but shifts to the laft. The orange dots rise with the fluorescence intensity and collapses to zero, when the EBFP2 intensity is high. For the other constructs the EGFP range fully corresponds with the EBFP2 range. The linear plot shows the orange and blue dots in more distinct lines. The orange dots from the construct containing the perfect and aligned binding sites assemble along the y-axis, where the EGFP fluorescence intensity is zero. For the other construct the EGFP line fully overlap with the EBFP2.
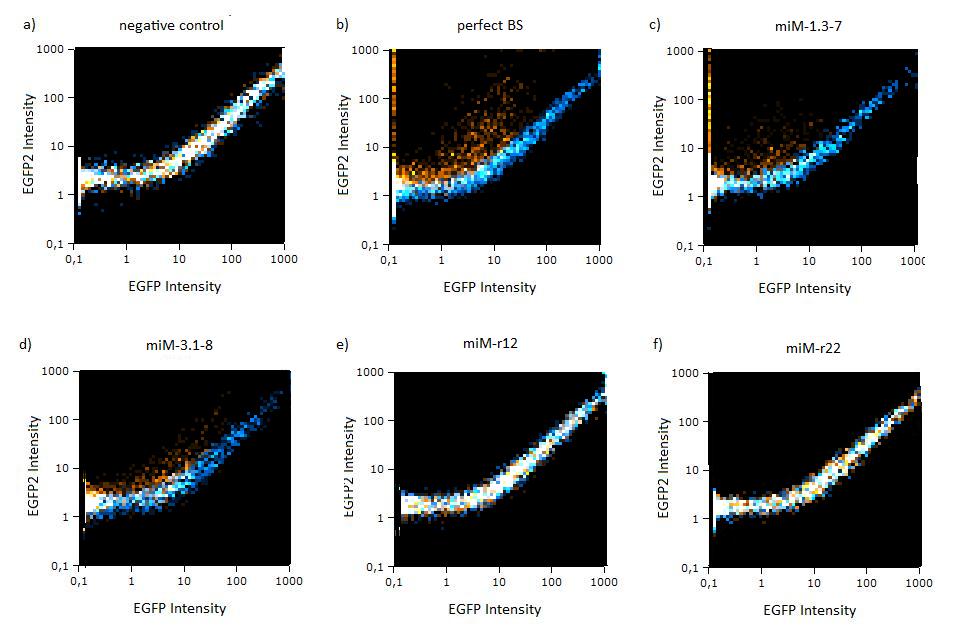 Figure 4:EGFP2/EBFP correlation of single transfected Hela cells according to flow cytometry analysis different binding sites for miR122 cotransfected with miR-122 or with miR-155, respectively. The orange dots represent the cotransfected cells with miR122 and the blue dots the cotransfected cells with miR-155. Hela cells were used. 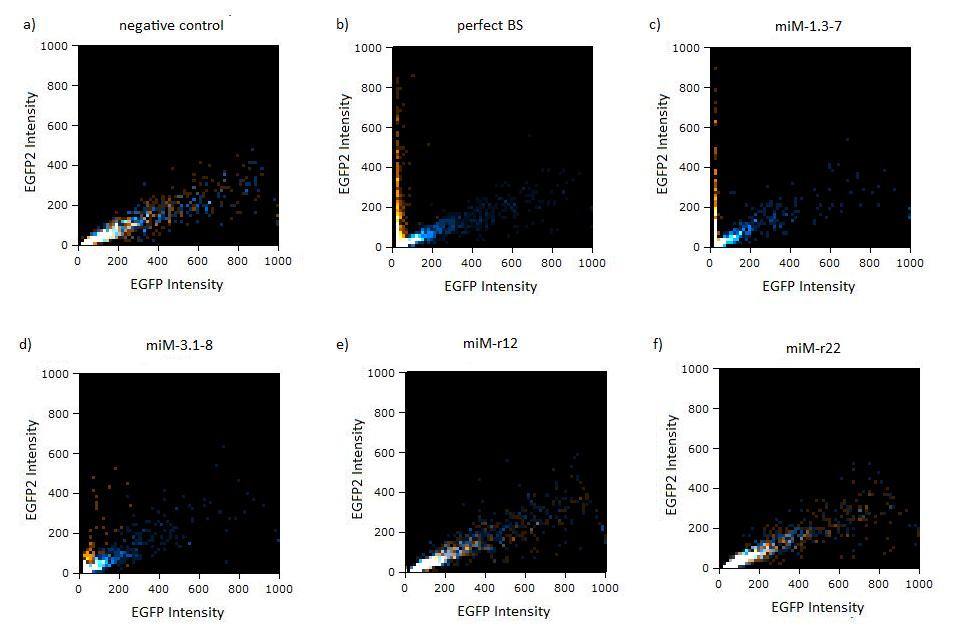 Figure 5: EGFP/EBFP2 correlation of single transfected Hela cells according to flow cytometry analysis different binding sites for miR122 cotransfected with miR-122 or with miR-155, respectively. The orange dots represent the cotransfected cells with miR122 and the blue dots the cotransfected cells with miR-155. Hela cells were used.
Confocal microscopy
 Figure 9: epifluorescent microscopy image (10x) of Hela cells transfected with miMeasure miMeasure with a perfect binding site is a) cotransfected with miR-155, which has no specificity to miR-122, b) cotransfected with miR-122, which is complementary to the perfect binding site. EGFP is regulated by miR-122, EBFP2 is unregulated and serves as transfection control.
Analysis of endogenous miRNAHepG2, another liver cell line, is also transfected with the constructs containing the perfect and the aligned constructs for miR-122. The cotransfection with miR-155 serves again as a negative control. For this cell line there was only a slight knock-down observed for all of the constructs, it is much less compared to the HuH7 cell line, where the knock-down ranges from 20-40%.
The EGFP/EBFP2 ratio from each transfected HuH7 cell is calculated for 52 cells. The cells were transfected with the construct carriying the three aligned perfect binding sites against miR-122. The EGFP/EBFP2 ratio in each cell is different and ranges from 0.1 to 6. DiscussionMeasurements of synthetic and endogenous miRNA expression level as well as the knock-down efficiency by different miRNAs can characterized via the miMeasure constructs. The fluorescence of EGFP and EBFP2, and therefore the efficiency of knockdown, can be estimated using different methods, for example flow cytometry, manual and automated fluorescence microscopy or potentially an automated fluorescence plate reader system. Flow cytometry and fluorescence microscopy offer the possibility to measure knockdown at the single cell level, the first one with the possibility of high throughput, the second with a good precision. Since flow cytometry has a smaller dynamic range, thus even a GFP knock-down of roughly 60% can cause the relative GFP fluorescence intensity values collapsing to zero.
This quantitative approach offers many possibilities, two of them being expored here: estimating the knockdown capacity of binding sites that differs from a perfect one and determining the presence of endogenous miRNA directly in living cells. ReferencesAi HW, Shaner NC, Cheng Z, Tsien RY, Campbell RE. Exploration of new chromophore structures leads to the identification of improved blue fluorescent proteins. Biochemistry. 2007 May 22;46(20):5904-10. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009 Jan 23;136(2):215-33. Coulouarn C, Factor VM, Andersen JB, Durkin ME, Thorgeirsson SS. Loss of miR-122 expression in liver cancer correlates with suppression of the hepatic phenotype and gain of metastatic propertiesmiR-122 repression is a marker of tumor progression in HCC. Oncogene. 2009 Oct 8;28(40):3526-36. Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem. 2010;79:351-79. Girard M, Jacquemin E, Munnich A, Lyonnet S, Henrion-Caude A. miR-122, a paradigm for the role of microRNAs in the liver. J Hepatol. 2008 Apr;48(4):648-56. Lin CJ, Gong HY, Tseng HC, Wang WL, Wu JL. miR-122 targets an anti-apoptotic gene, Bcl-w, in human hepatocellular carcinoma cell lines. Biochem Biophys Res Commun. 2008 Oct 24;375(3):315-20.
|
||||||||||||||||||||||||||||||||||||||||
 "
"